-
Basecalling overview
A user can basecall and demultiplex their data directly in MinKNOW after a sequencing experiment has finished to avoid having to use command-line tools, or to re-analyse old data using the latest basecalling models. Reads can also be aligned against a reference post-sequencing.
Note: Both barcoding and alignment can be run on FASTQ, POD5 or FAST5 reads when coupled with basecalling.
-
Click 'Analysis' on the start page.
-
Click 'Basecalling' to open the post-run basecalling set-up options.
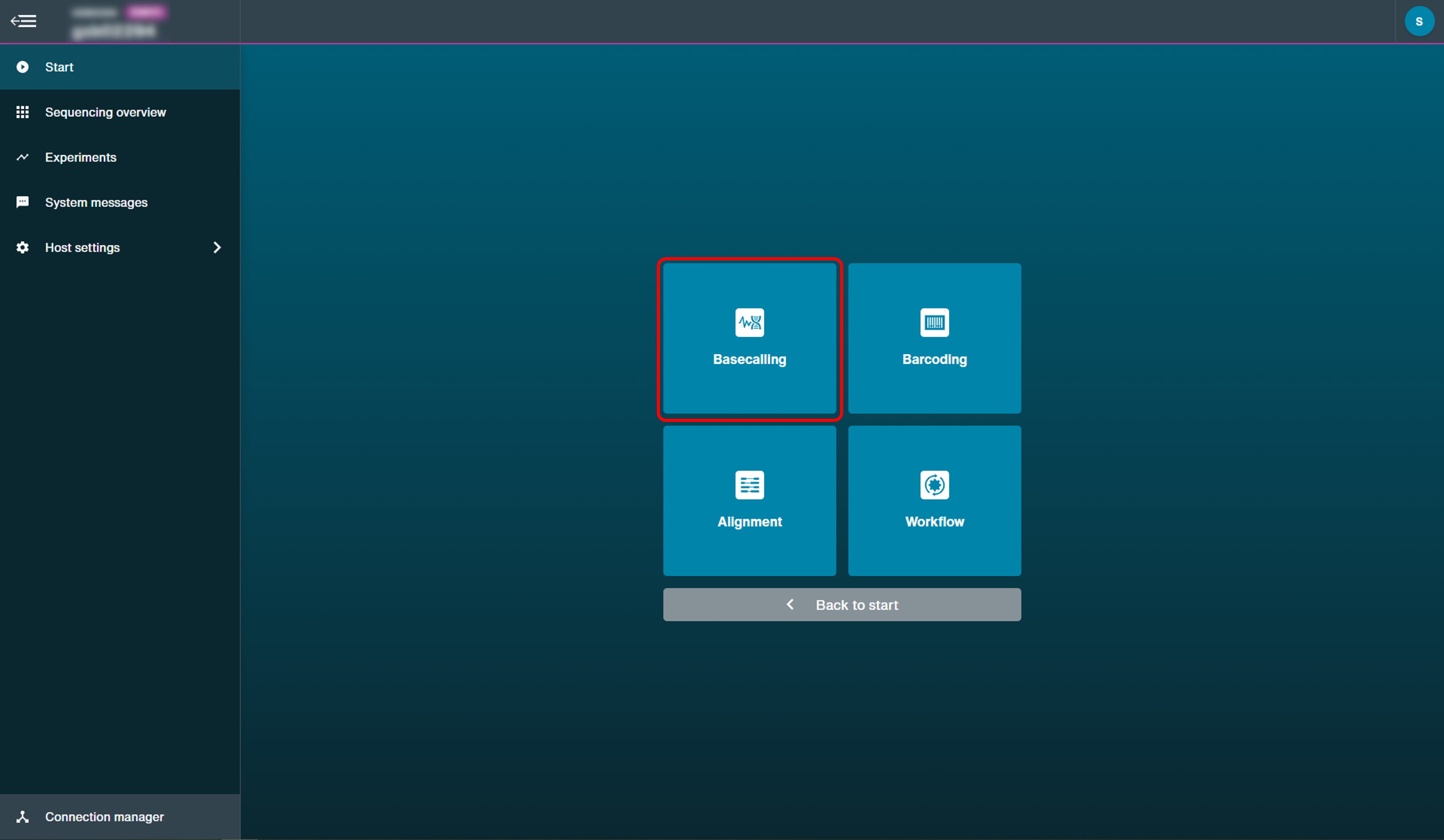
-
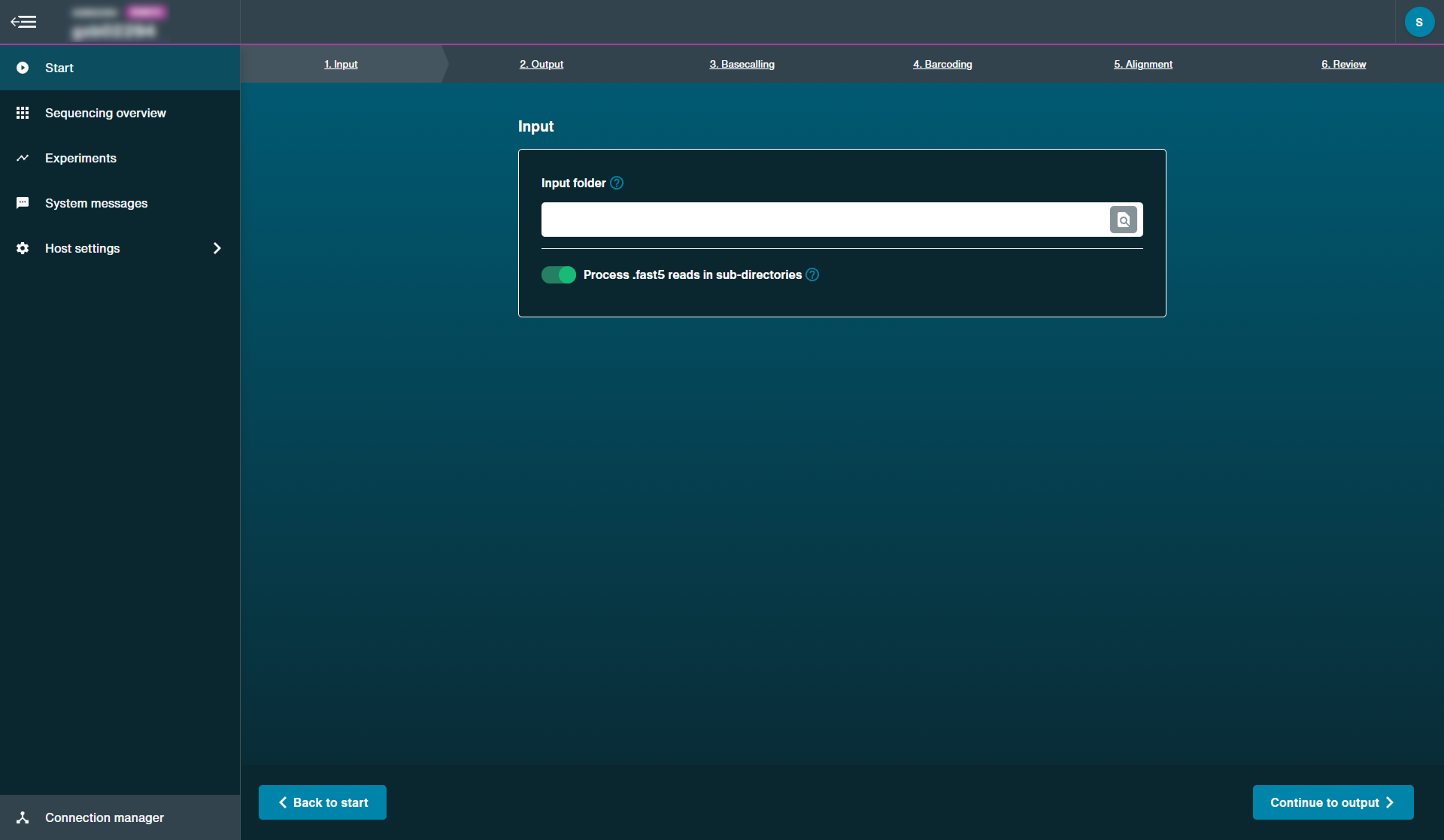
Select input folder containing the FAST5 files to basecall from a previous run.
Select whether you want to process FAST5 reads in sub-directories.
If your chosen read input folder contains sub-directories with FAST5 files, you can choose whether to basecall FAST5 files in these sub-directories by enabling this option.
-
Select output folder and file type.
By default, MinKNOW will create a
/basecalledfolder in/data. You can set a different folder in which to save the basecalled reads.
Note: This must be a sub-folder of/datadirectory.Select whether to output FAST5 files. If selected, these files will be written to a folder within
/basecalledcalled/workspace, if selected.Users are also able to enable read splitting options post-run.
-
Optional actionSelect to use alignment and import a reference sequence in a FASTA file.
We currently only recommend uploading an alignment reference locally for bacterial-sized genomes. Upload can take a few minutes and is compute dependent.
A reference file must by uploaded as a FASTA file that can contain multiple entries in the same file (e.g. multiple chromosomes).
However, a .bed file may also be uploaded alongside the reference FASTA file. The BED file option can be used when the user is interested in a particular region of the reference (e.g. specific gene in the chromosome).
Alignment generates FASTQ output files when performed with basecalling.
-
Click 'Start' to begin post-run basecalling.