- Materials
-
- Ligation Sequencing Kit (SQK-LSK109)
- Flow Cell Priming Kit (EXP-FLP002)
- Native Barcoding Expansion 1-12 (EXP-NBD104) and 13-24 (EXP-NBD114) if multiplexing more than 12 samples
- ASFV-positive blood samples
- Consumables
-
- ASFV primers
- Agencourt AMPure XP beads (Beckman Coulter, A63881)
- Lysis buffer (10 mM ammonium chloride, 150 mM sodium EDTA, 10 mM sodium bicarbonate)
- 5x TEN buffer (0.05 M EDTA, 0.5 M NaCl, 20 mg/ml Proteinase K, 20% SDS, in 0.05 M Tris-HCl, pH 8.0)
- Isopropanol, 100% (Fisher, 10723124)
- NEBNext® Companion Module for Oxford Nanopore Technologies® Ligation Sequencing (NEB, E7180S or E7180L).
Alternatively, you can use the NEBNext® products below:
- NEBNext FFPE Repair Mix (NEB, M6630)
- NEBNext Ultra II End repair/dA-tailing Module (NEB, E7546)
- NEBNext Quick Ligation Module (NEB, E6056)
- 1.5 ml Eppendorf DNA LoBind tubes
- 0.2 ml thin-walled PCR tubes
- Nuclease-free water (e.g. ThermoFisher, AM9937)
- Freshly prepared 70% ethanol in nuclease-free water
- Equipment
-
- Heat block or water bath set to 95°C
- Hula mixer (gentle rotator mixer)
- Magnetic rack, suitable for 1.5 ml Eppendorf tubes
- Microfuge
- Vortex mixer
- Thermal cycler
- P1000 pipette and tips
- P200 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- P10 pipette and tips
- P2 pipette and tips
- Ice bucket with ice
- Timer
- Optional equipment
-
- Agilent Bioanalyzer (or equivalent)
- Qubit fluorometer (or equivalent for QC check)
- Eppendorf 5424 centrifuge (or equivalent)
-
NEBNext® Companion Module for Oxford Nanopore Technologies® Ligation Sequencing
For customers new to nanopore sequencing, we recommend buying the NEBNext® Companion Module for Oxford Nanopore Technologies® Ligation Sequencing (catalogue number E7180S or E7180L), which contains all the NEB reagents needed for use with the Ligation Sequencing Kit.
Please note, for our amplicon protocols, NEBNext FFPE DNA Repair Mix and NEBNext FFPE DNA Repair Buffer are not required.
-
Ligation Sequencing Kit (SQK-LSK109) contents
Name Acronym Cap colour No. of vials Fill volume per vial (µl) DNA CS DCS Yellow 1 50 Adapter Mix AMX Green 1 40 Ligation Buffer LNB Clear 1 200 L Fragment Buffer LFB White cap, orange stripe on label 2 1,800 S Fragment Buffer SFB Grey 2 1,800 Sequencing Buffer SQB Red 2 300 Elution Buffer EB Black 1 200 Loading Beads LB Pink 1 360 -
Flow Cell Priming Kit (EXP-FLP002) contents
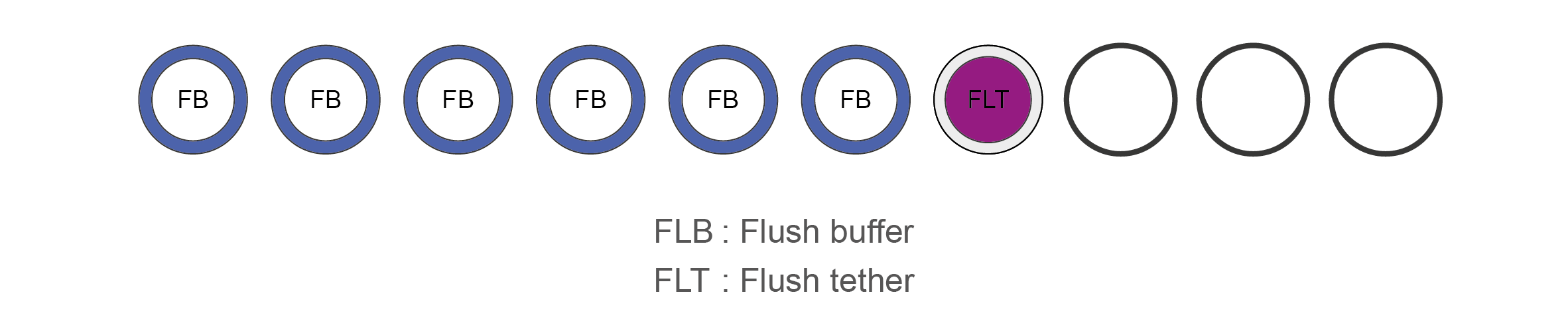
Name Acronym Cap colour No. of vial Fill volume per vial (μl) Flush Buffer FB Blue 6 1,170 Flush Tether FLT Purple 1 200 -
Native Barcoding Expansion 1-12 (EXP-NBD104) and 13-24 (EXP-NBD114) contents
EXP-NBD104 kit contents
Name Acronym Cap colour No. of vials Fill volume per vial (μl) Native Barcode 01-12 NB01-12 White 12 20 Adapter Mix II AMII Green 1 40
EXP-NBD114 kit contents
Name Acronym Cap colour No. of vials Fill volume per vial (μl) Native Barcode 13-24 NB13-24 White 12 20 Adapter Mix II AMII Green 1 40 -
Native barcode sequences
The native barcode sequences are the reverse complement of the corresponding barcode sequence in other kits. 24 unique barcodes are available in the Native Barcoding Expansion 1-12 and 13-24 (EXP-NBD104 and EXP-NBD114).
Native Barcoding Expansion 1-12 and 13-24 (EXP-NBD104 and EXP-NBD114)
Component Forward sequence Reverse sequence NB01 CACAAAGACACCGACAACTTTCTT AAGAAAGTTGTCGGTGTCTTTGTG NB02 ACAGACGACTACAAACGGAATCGA TCGATTCCGTTTGTAGTCGTCTGT NB03 CCTGGTAACTGGGACACAAGACTC GAGTCTTGTGTCCCAGTTACCAGG NB04 TAGGGAAACACGATAGAATCCGAA TTCGGATTCTATCGTGTTTCCCTA NB05 AAGGTTACACAAACCCTGGACAAG CTTGTCCAGGGTTTGTGTAACCTT NB06 GACTACTTTCTGCCTTTGCGAGAA TTCTCGCAAAGGCAGAAAGTAGTC NB07 AAGGATTCATTCCCACGGTAACAC GTGTTACCGTGGGAATGAATCCTT NB08 ACGTAACTTGGTTTGTTCCCTGAA TTCAGGGAACAAACCAAGTTACGT NB09 AACCAAGACTCGCTGTGCCTAGTT AACTAGGCACAGCGAGTCTTGGTT NB10 GAGAGGACAAAGGTTTCAACGCTT AAGCGTTGAAACCTTTGTCCTCTC NB11 TCCATTCCCTCCGATAGATGAAAC GTTTCATCTATCGGAGGGAATGGA NB12 TCCGATTCTGCTTCTTTCTACCTG CAGGTAGAAAGAAGCAGAATCGGA NB13 AGAACGACTTCCATACTCGTGTGA TCACACGAGTATGGAAGTCGTTCT NB14 AACGAGTCTCTTGGGACCCATAGA TCTATGGGTCCCAAGAGACTCGTT NB15 AGGTCTACCTCGCTAACACCACTG CAGTGGTGTTAGCGAGGTAGACCT NB16 CGTCAACTGACAGTGGTTCGTACT AGTACGAACCACTGTCAGTTGACG NB17 ACCCTCCAGGAAAGTACCTCTGAT ATCAGAGGTACTTTCCTGGAGGGT NB18 CCAAACCCAACAACCTAGATAGGC GCCTATCTAGGTTGTTGGGTTTGG NB19 GTTCCTCGTGCAGTGTCAAGAGAT ATCTCTTGACACTGCACGAGGAAC NB20 TTGCGTCCTGTTACGAGAACTCAT ATGAGTTCTCGTAACAGGACGCAA NB21 GAGCCTCTCATTGTCCGTTCTCTA TAGAGAACGGACAATGAGAGGCTC NB22 ACCACTGCCATGTATCAAAGTACG CGTACTTTGATACATGGCAGTGGT NB23 CTTACTACCCAGTGAACCTCCTCG CGAGGAGGTTCACTGGGTAGTAAG NB24 GCATAGTTCTGCATGATGGGTTAG CTAACCCATCATGCAGAACTATGC -
ASFV primer sequences
ASFV primer sequences described in the protocol are subject to change. Any updates can be found at ASFV Lilo GitHub page.
Amplicon - primer pair Forward sequence Reverse sequence ASFV1 GGCGTTCATTTCACAAGATGC ACGGCATCTAAGCAGCTCAATG ASFV2 CAGGCCGATATATCATTTCATCAATATTCA ACCCAAAGCCCTGGAATCCTTA ASFV3 GCAAACCAAGTGACTCACCCTC ATTGTATGACGTCGGGGCAGAT ASFV4 ACCTAGTAAAAGTCCTAGAAAAACCTTCA CGCCATTGTTTTACACAGTCGC ASFV5 TCGAGATTTTATTATTTGGATGCATCATCA GGACTGATGAAAGCCGTGAA ASFV6 TCCACGCGGTACTTGGCTCC AGGCCTCGTTGGTGGAAAGGA ASFV7 AGGGCTGATGCAAATCTCTTTTTCA TCTCCGATTTTCGCATGCCAAA ASFV8 AGTTTGCAAAGAGCCTAAAGATAGACT AGCGTGGAACTGTAGATGACGA ASFV9 TGCAGAAACCGCAGATGAATGT ATAGGATTAGATGCGACGCCCA ASFV10 GCATGTAGAGAGGTTTTGGTAGTCA GGAAACAGCTGGAGAGTTGTGG ASFV11 TGGTTTTGAAATAAAATGCCTTCTACGG GGAATGCATGGACGAAGAAGCA ASFV11 (alt) CTATGGGATGGGAAGAGTGGTCAA CGTCAACCGCCGCATTAGC ASFV12 TCCTTGGGAGTTACAGCGAAGA AATGAAATCATTCGCGGCGAGT ASFV13 CAGACATTGGCAGTGATGGCTA GAAATGCCGGGCCTTCTACAAA ASFV14 GCTACTCCCCCAAATATCACATATAATTGT TTTTTCGTGTTGCTGTTCGGGA ASFV15 GGATGGCACCCTTCTCACAATC TGCGTATGACCCGATGTTGTTG ASFV16 GTCGACTTCACAGGAACAACGG ACCCGCTTTACACAAAACACGT ASFV17 TGGAATTTCCTGACGTGGCAAA GCAACCGCTATTCCAAACAGGA ASFV18 AGTTGTTGTCCTAGACCGTGGCA TGAAAAGGAGGGCACGATCC ASFV19 CCCGTATGCGGGCGTACTTT TGGCCTCTTCTTTCCCCCGA ASFV20 GGCCGCAACATTTGTGTCAAAG GCTCGCGAACAAATTACTCCCA ASFV21 GAATGGCAGCGATGATCTCAGG TGCAGGGCAAGGGTATACTGAA ASFV22 TGGCGTCGTTTAACAGCTTGAT GCTGGATGGCAAATCGGTTGTA ASFV23 AGGCGTGAAAATTCTTCTTCAAACA AGACGTTTTAAGCTGCATGGCA ASFV24 GGCAGCAGGATCTTAAAACCGG TGCATAATGCCCAGCTTTTCGT ASFV25 GCTGTTTAAGCGTTTCAAGCTGA CTCCGCGGGGAACATTGTTTTA ASFV26 CCCTGGGAGGAGTCATCATGAA GGTCATTGACTTTGGAAGCGCT ASFV27 ACTGTCTGCTAGACTCCCAGGA CCCAAGAGGAGGAATGGTTTGC ASFV28 GCCCCCTAGCGTCACCGAAT CCAAGCCTGCTGCGAAGCTC ASFV29 GACGCAATTTCGGCTGTTTTTAAAA GACTTGGTCTCCGGCTCAAAAG ASFV30 GTTGGGGTGTTGGAGCGAATAA TTCTGCTTACGGACGATGCAAC ASFV31 TAGTTGTGAAGCGTTCTCGGGT GAGCACATGTTACTCGCCACTC ASFV32 GGACTTCTTATGCTCAGATGGGC ACTGCTGCAGGCGTTAAACATT
