- Materials
-
- Flow Cell Wash Kit XL (EXP-WSH004-XL)
- Sequencing Auxiliary Vials V14 (EXP-AUX003)
- Sequencing Buffer (SB)
- Library Solution (LIS)
- Library Beads (LIB)
- Flow Cell Flush (FCF)
- Flow Cell Tether (FCT)
- Equipment
-
- P1000 pipette and tips
- P20 pipette and tips
- Ice bucket with ice
-
Day 6: Pore-C experiment flow cell washing and reloading
We recommend washing and reloading your PromethION Flow Cell with a new library to maintain high data acquisition.
For the Pore-C experiment, up to four libraries prepared using the Pore-C protocol and Ligation Sequencing Kit XL V14 (SQK-LSK114-XL) can be loaded on the PromethION Flow Cell during a sequencing run. We recommend washing your flow cell when ~20-25% of active pores are remaining, which typical occurs after ~18 hours of sequencing. Washing removes most of the initial library as well as unblocking pores to prepare a flow cell for loading a fresh library for further sequencing. Pore-C DNA extracts are prone to blocking and may require closer monitoring and washing earlier into the sequencing run than the duplex experiment to optimise sequencing output.
Navigate to the Pore Activity or the Pore Scan Results plot to see pore availability. Below is an example of pore states observed on a flow cell before and after wash steps are performed. The red asterisks indicates the reloads.
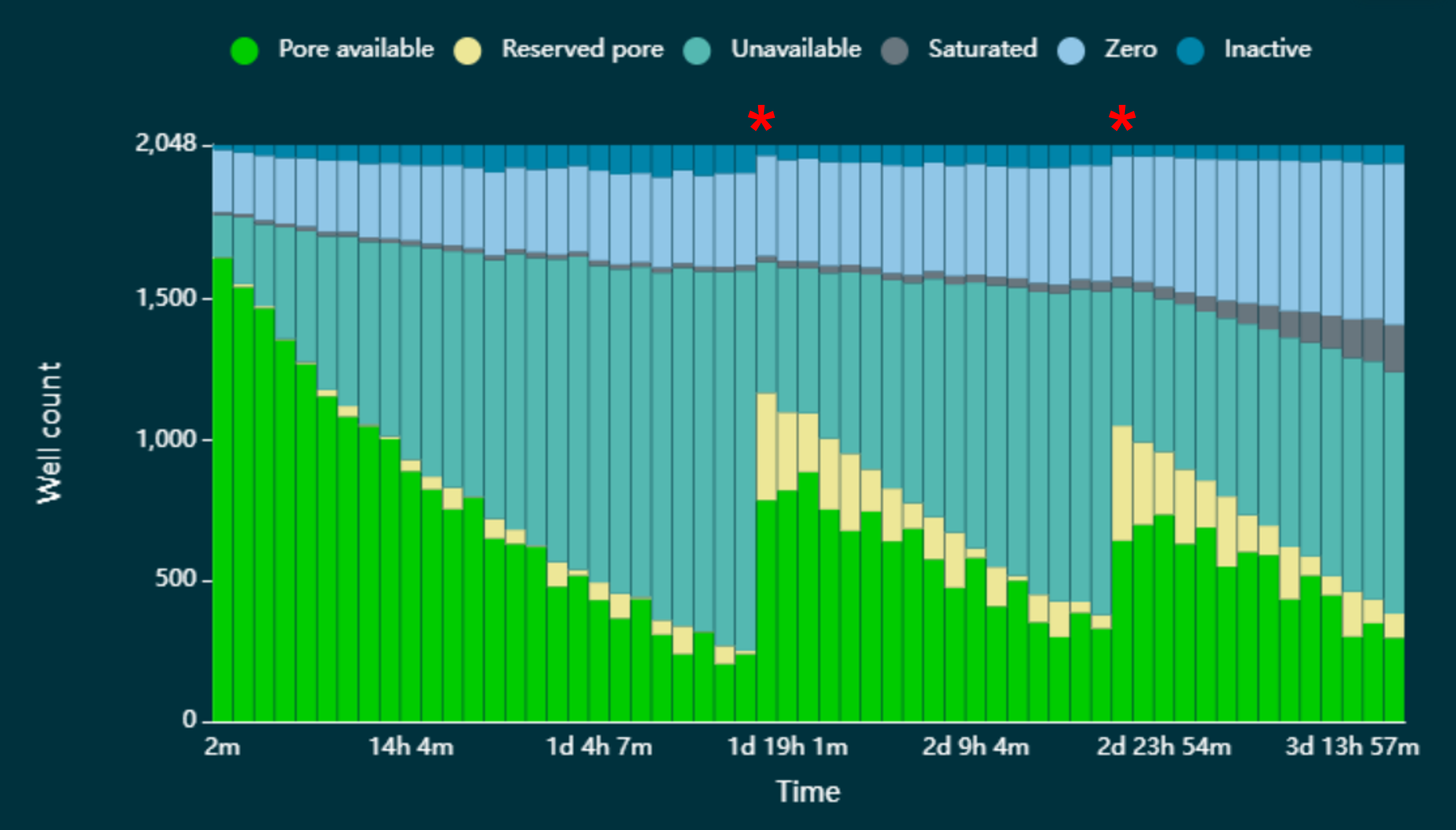
We recommend reloading your Pore-C experiment on days 5, 6 and 7.
-
Place the tube of Wash Mix (WMX) on ice. Do not vortex the tube.
-
Thaw one tube of Wash Diluent (DIL) at room temperature.
-
Mix the contents of Wash Diluent (DIL) thoroughly by vortexing, then spin down briefly and place on ice.
-
In a fresh 1.5 ml Eppendorf DNA LoBind tube, prepare the following Flow Cell Wash Mix:
Reagent Volume per flow cell Wash Mix (WMX) 2 μl Wash Diluent (DIL) 398 μl Total 400 μl -
Mix well by pipetting, and place on ice. Do not vortex the tube.
-
Pause the sequencing experiment in MinKNOW, and leave the flow cell in the device.
-
Remove waste buffer, as follows:
- Close the inlet port.
- Insert a P1000 pipette into a waste port and remove the waste buffer.
Note: As both the inlet port is closed, no fluid should leave the sensor array area.
-
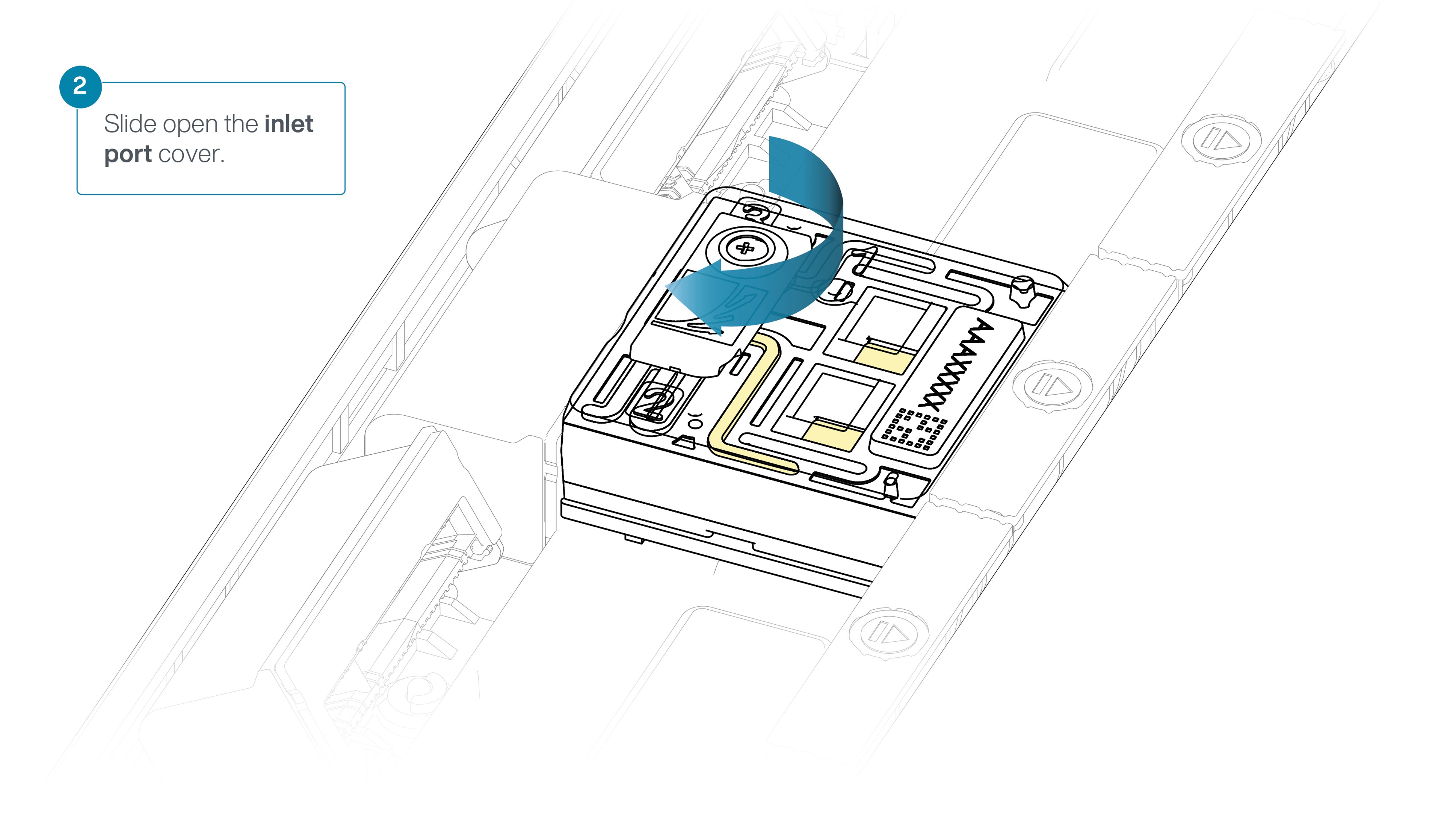
Slide the inlet port cover clockwise to open the inlet port.
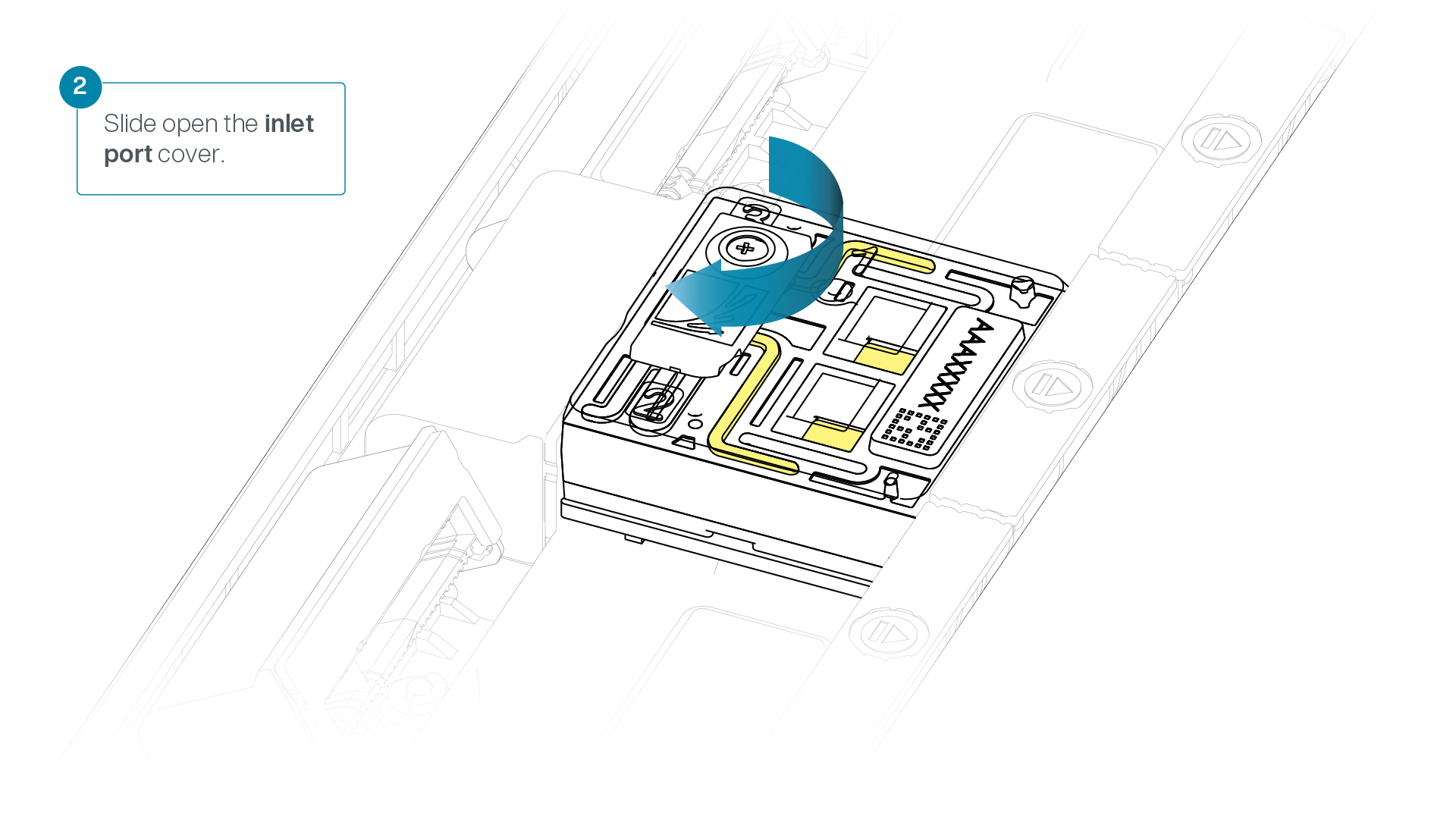
-
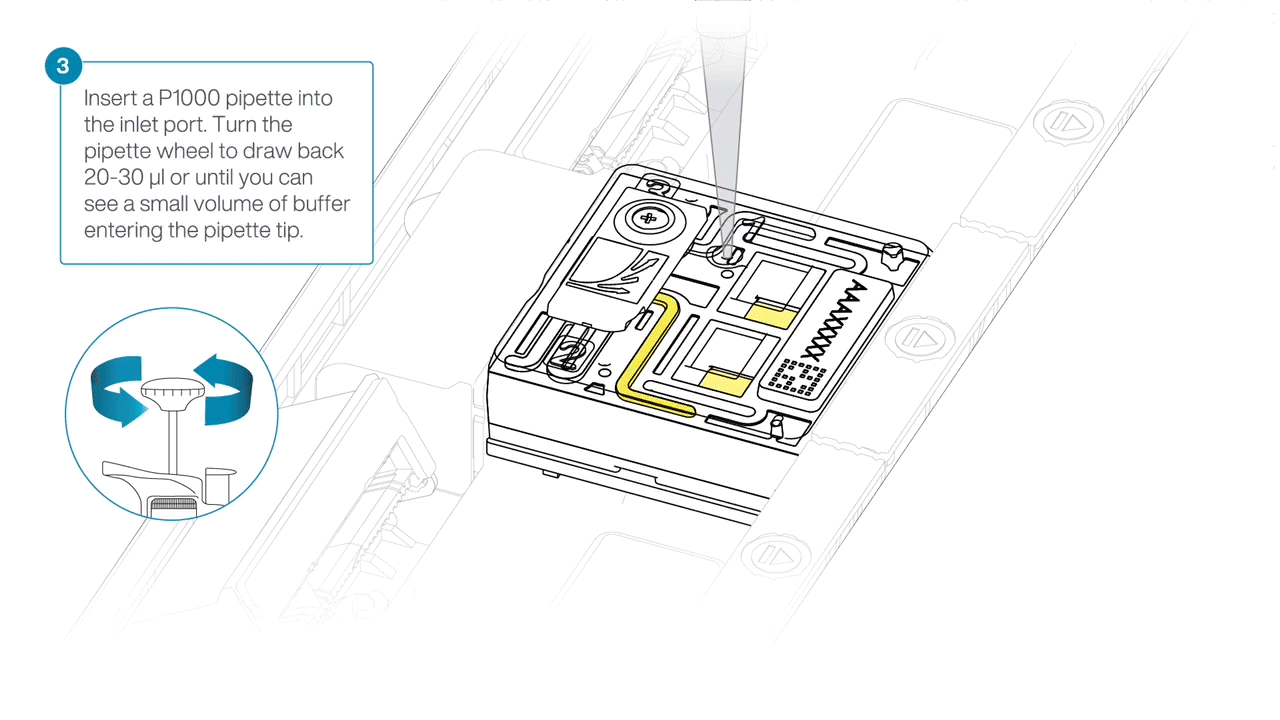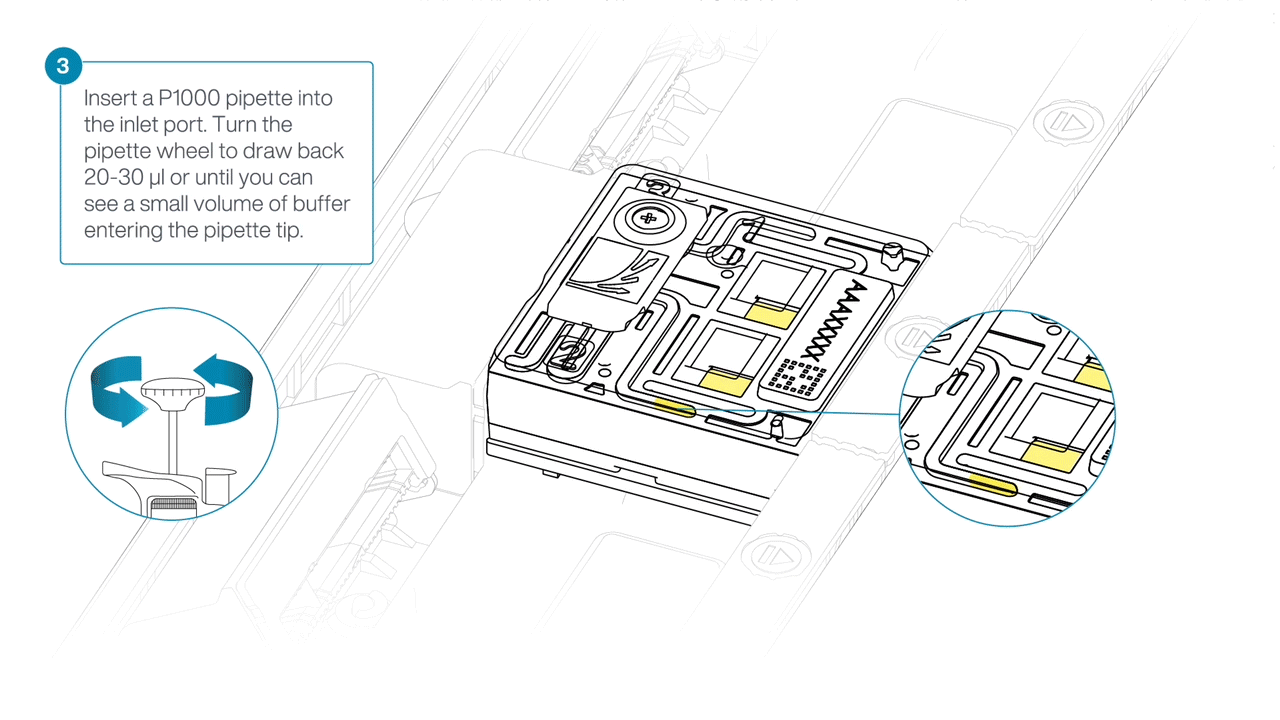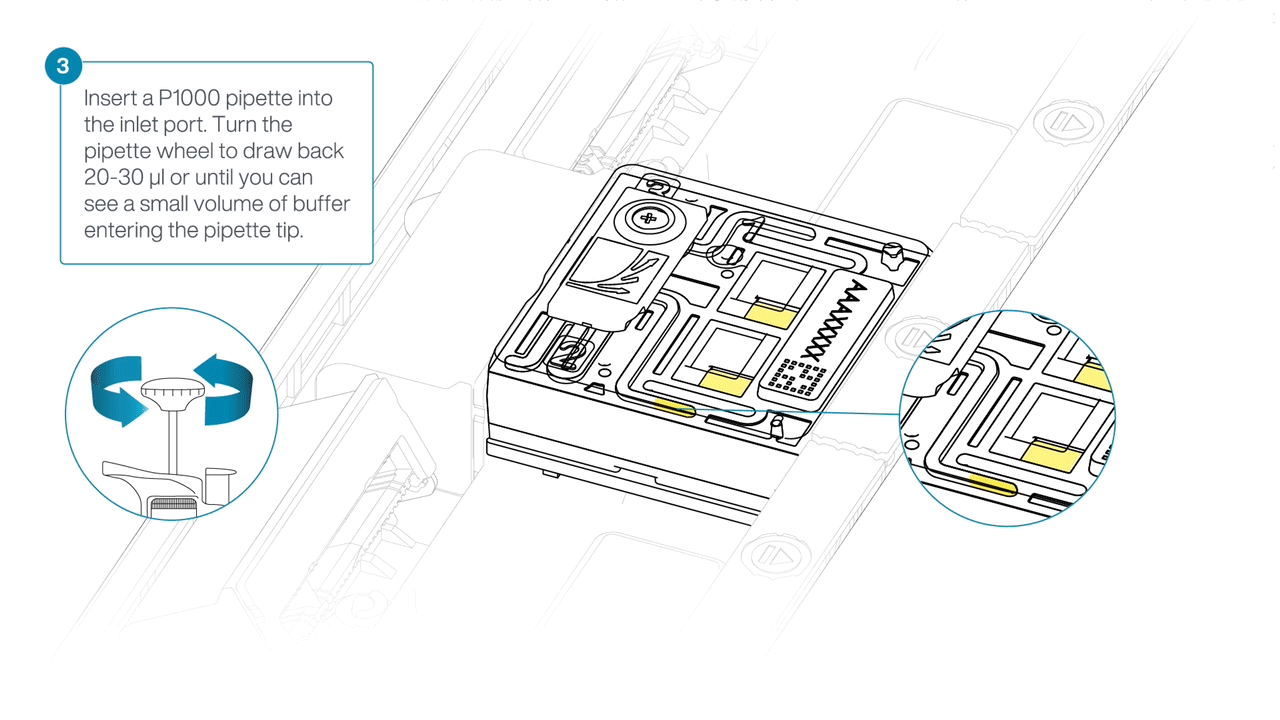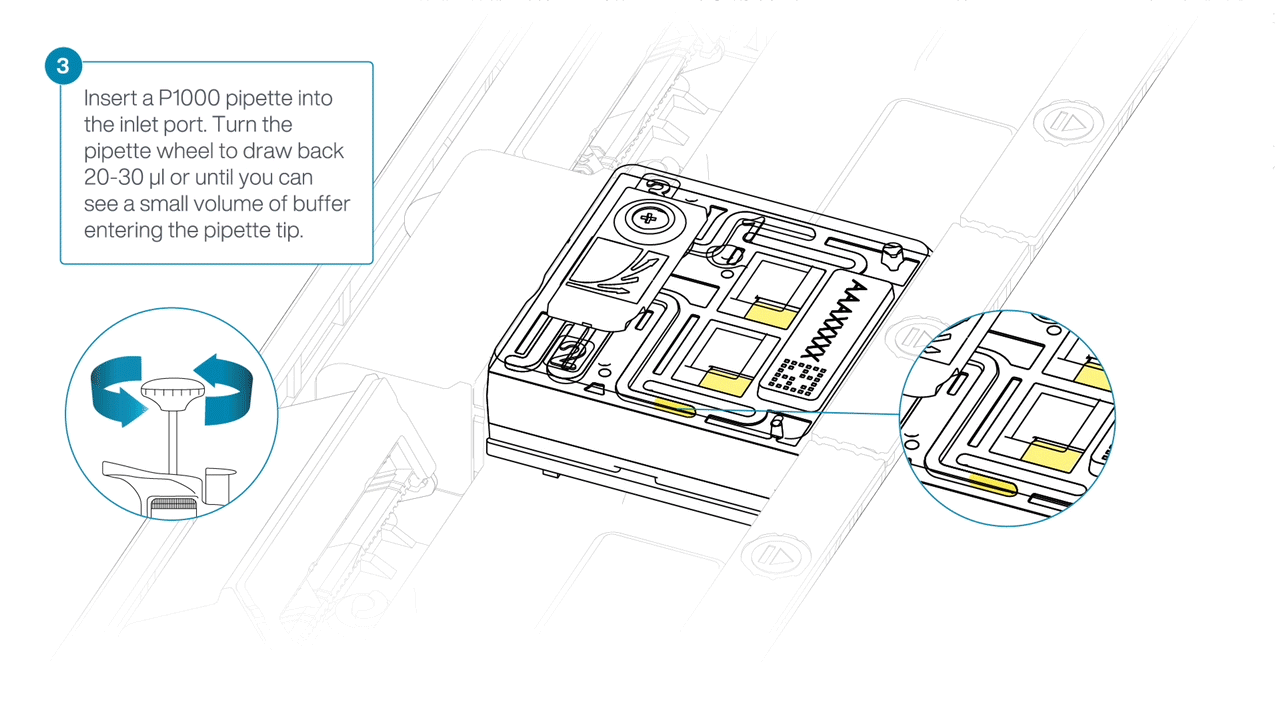
After opening the inlet port, check for a small air bubble under the cover. Draw back a small volume to remove any bubbles:
- Set a P1000 pipette to 200 µl
- Insert the tip into the inlet port
- Turn the wheel until the dial shows 220-230 µl, or until you can see a small volume of buffer entering the pipette tip.
-
Slowly load 200 µl of the prepared flow cell wash mix into the inlet port, as follows:
- Using a P1000 pipette, take 200 µl of the flow cell wash mix
- Insert the pipette tip into the inlet port, ensuring there are no bubbles in the tip
- Slowly twist the pipette wheel down to load the flow cell (if possible with your pipette) or push down the plunger very slowly, leaving a small volume of buffer in the pipette tip.
- Set a timer for a 5 minute incubation.
-
Once the 5 minute incubation time is complete, carefully load the remaining 200 µl of the prepared flow cell wash mix into the inlet port, as follows:
- Using a P1000 pipette, take 200 µl of the flow cell wash mix
- Insert the pipette tip into the inlet port, ensuring there are no bubbles in the tip
- Slowly twist the pipette wheel down to load the flow cell (if possible with your pipette) or push down the plunger very slowly, leaving a small volume of buffer in the pipette tip.
-
Close the inlet port and wait for 1 hour.
-
Remove the waste buffer, as follows:
- Ensure the inlet port is closed.
- Insert a P1000 pipette into a waste port and remove the waste buffer
Note: As the inlet port is closed, no fluid should leave the sensor array area.
-
Thaw the Sequencing Buffer (SB), Library Beads (LIB) or Library Solution (LIS, if using), Flow Cell Tether (FCT) and Flow Cell Flush (FCF) at room temperature before mixing by vortexing. Then spin down and store on ice.
-
Prepare the flow cell priming mix in a suitable tube for the number of flow cells to flush. Once combined, mix well by briefly vortexing.
Reagents Volume per flow cell Flow Cell Flush (FCF) 1,170 µl Flow Cell Tether (FCT) 30 µl Total volume 1,200 µl -
Slide the inlet port cover clockwise to open.
-
After opening the inlet port, draw back a small volume to remove any air bubbles:
- Set a P1000 pipette tip to 200 µl.
- Insert the tip into the inlet port.
- Turn the wheel until the dial shows 220-230 µl, or until you see a small volume of buffer entering the pipette tip.

-
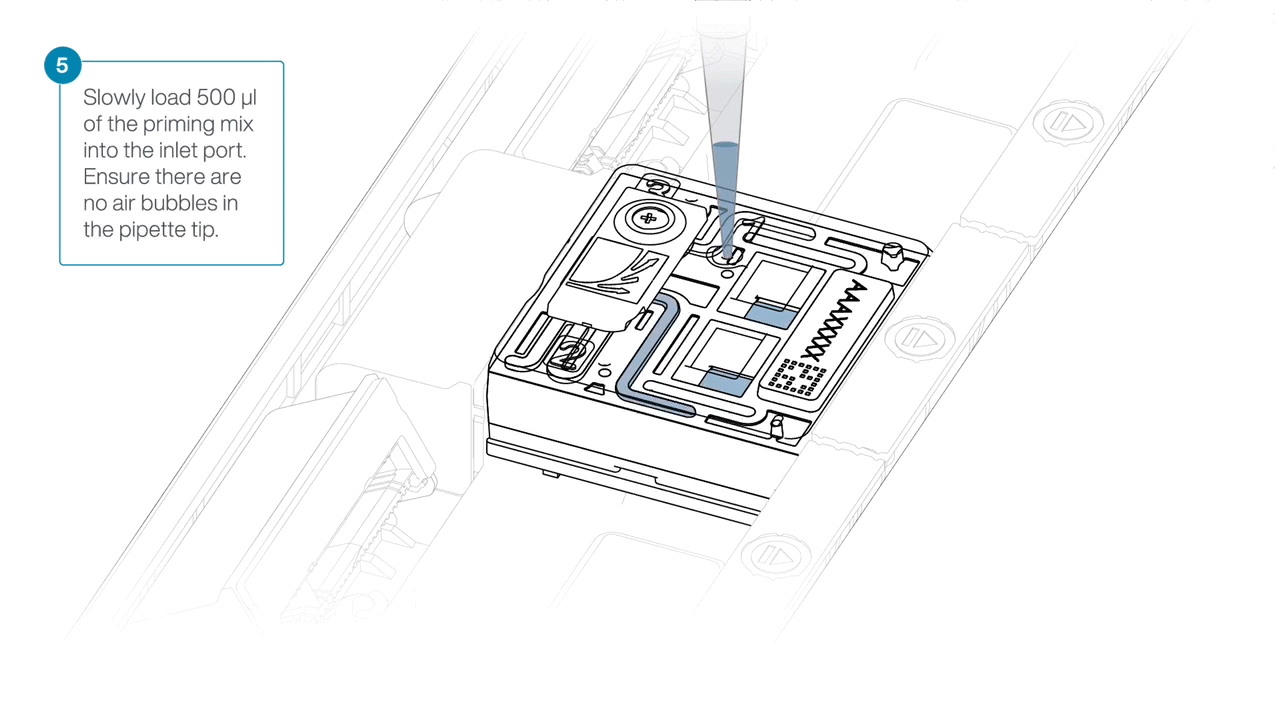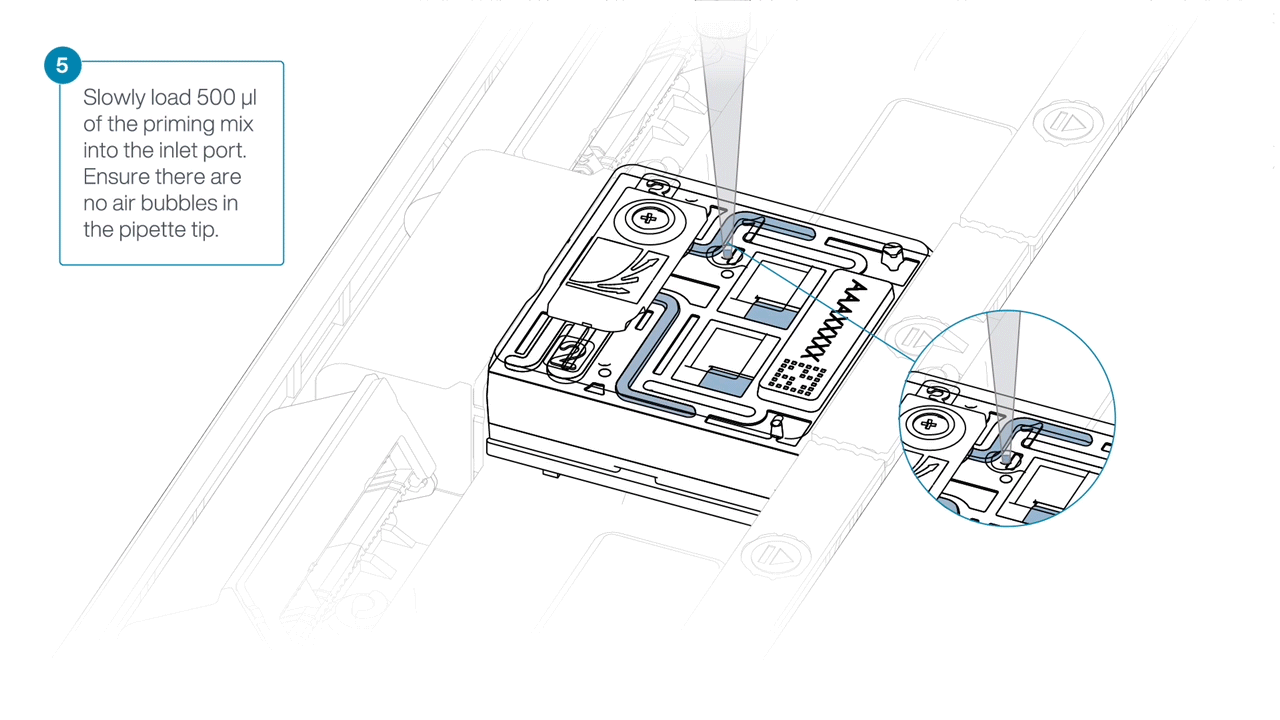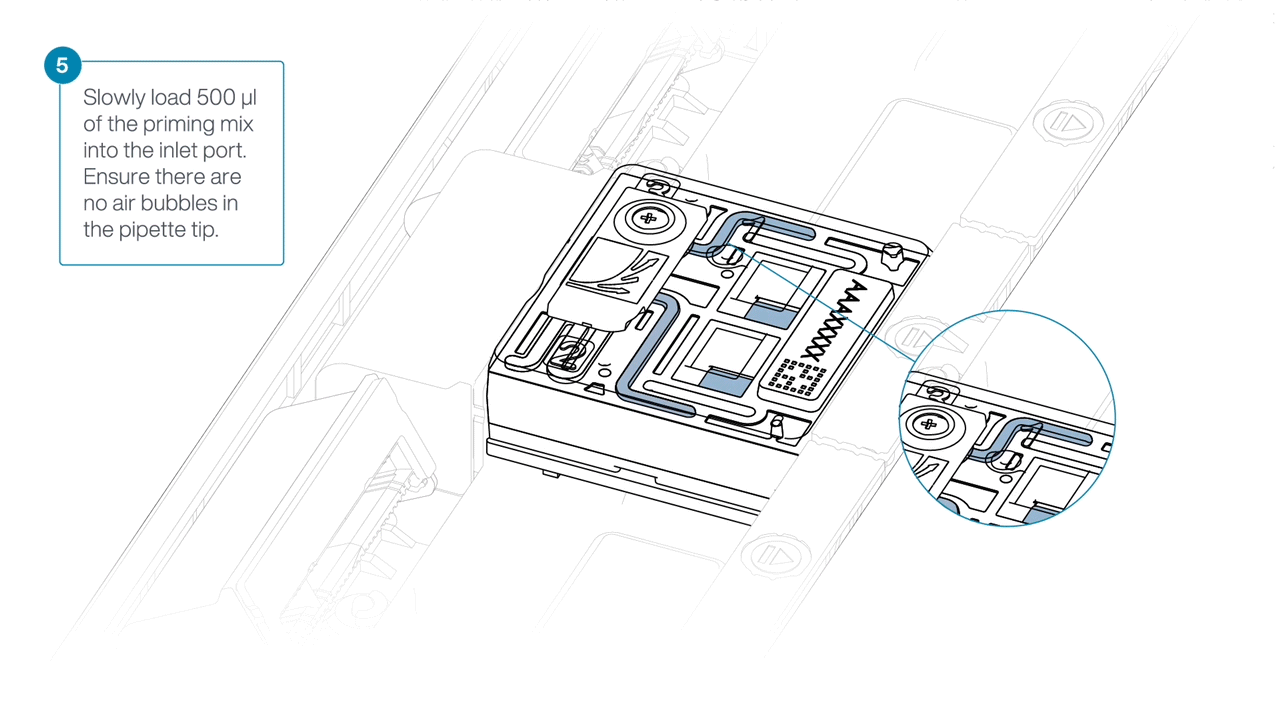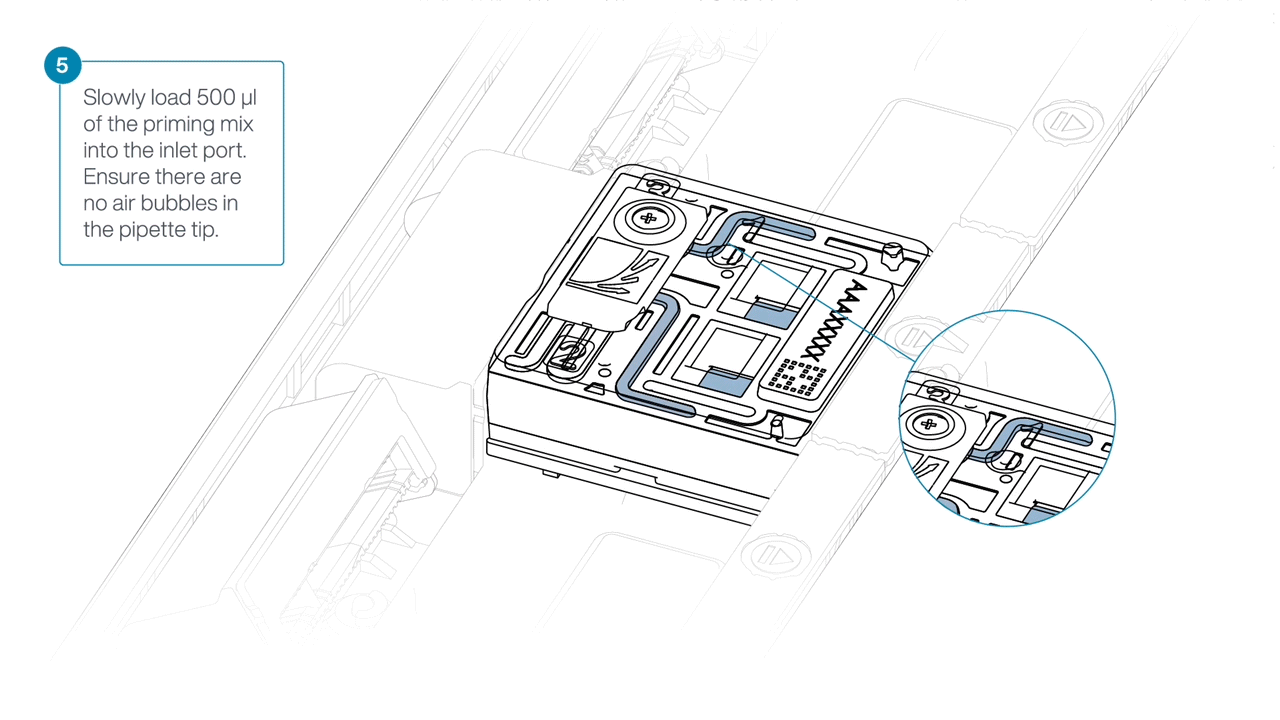
Slowly load 500 µl of the priming mix into the inlet port, as follows:
- Using a P1000 pipette, take 500 µl of the priming mix
- Insert the pipette tip into the priming port, ensuring there are no bubbles in the tip
- Slowly twist the pipette wheel down to load the flow cell (if possible with your pipette) or push down the plunger very slowly, leaving a small volume of buffer in the pipette tip.
-
Close the inlet port and wait five minutes.
During this time, prepare the library for loading using the next steps in the protocol.
-
Thoroughly mix the contents of the Library Beads (LIB) by pipetting.
-
In a new 1.5 ml Eppendorf DNA LoBind tube, prepare the library for loading as follows:
Reagent Volume per flow cell Sequencing Buffer (SB) 100 µl Library Beads (LIB) thoroughly mixed before use, or Library Solution (LIS) 68 µl DNA library 32 µl Total 200 µl Note: Library loading volume has been increased to improve array coverage.
-
Remove the waste buffer, as follows:
- Ensure the inlet port is closed.
- Insert a P1000 pipette into a waste port and remove the waste buffer
Note: As the inlet port is closed, no fluid should leave the sensor array area.
-
Slide the inlet port cover clockwise to open.

-
After opening the inlet port, draw back a small volume to remove any air bubbles:
- Set a P1000 pipette tip to 200 µl.
- Insert the tip into the inlet port.
- Turn the wheel until the dial shows 220-230 µl, or until you see a small volume of buffer entering the pipette tip.

-
Slowly load 500 µl of the priming mix into the inlet port, as follows:
- Using a P1000 pipette, take 500 µl of the priming mix
- Insert the pipette tip into the priming port, ensuring there are no bubbles in the tip
- Slowly twist the pipette wheel down to load the flow cell (if possible with your pipette) or push down the plunger very slowly, leaving a small volume of buffer in the pipette tip.

-
Remove waste buffer, as follows:
- Close the inlet port.
- Insert a P1000 pipette into a waste port and remove the waste buffer.
Note: As both the inlet port is closed, no fluid should leave the sensor array area.
-
Slide the inlet port cover clockwise to open.

-
After opening the inlet port, draw back a small volume to remove any air bubbles:
- Set a P1000 pipette tip to 200 µl.
- Insert the tip into the inlet port.
- Turn the wheel until the dial shows 220-230 µl, or until you see a small volume of buffer entering the pipette tip.

-
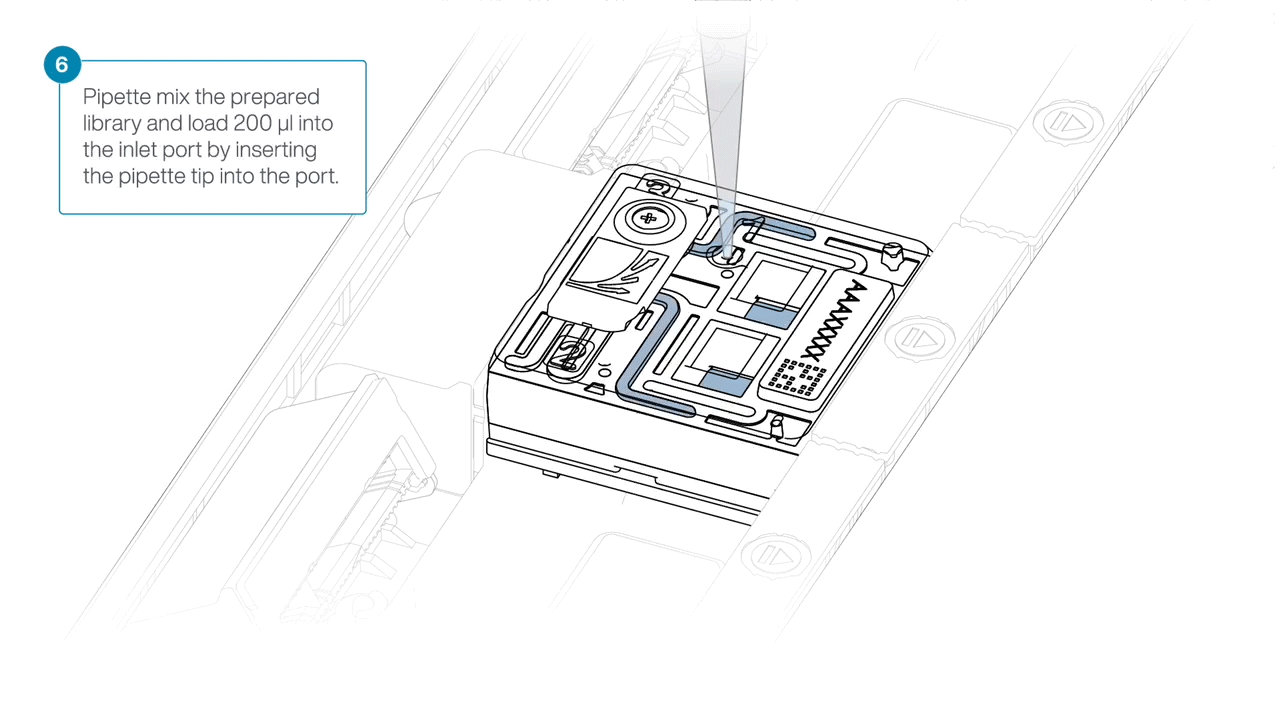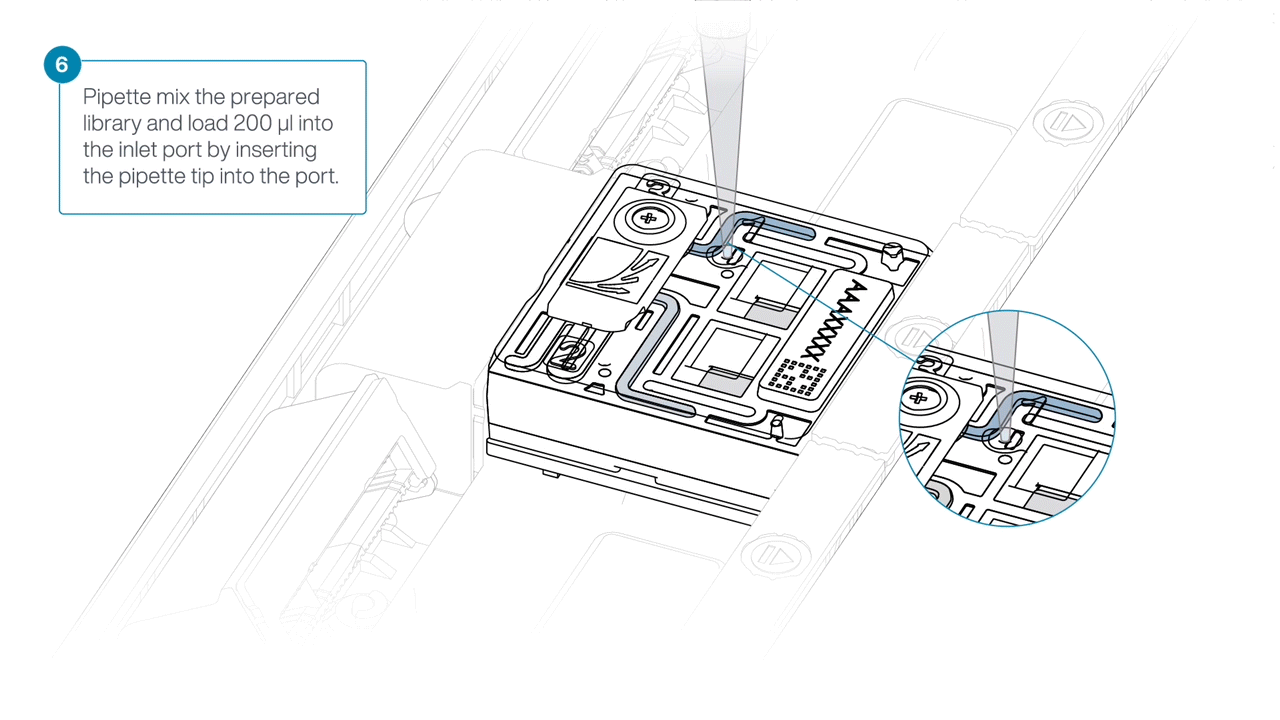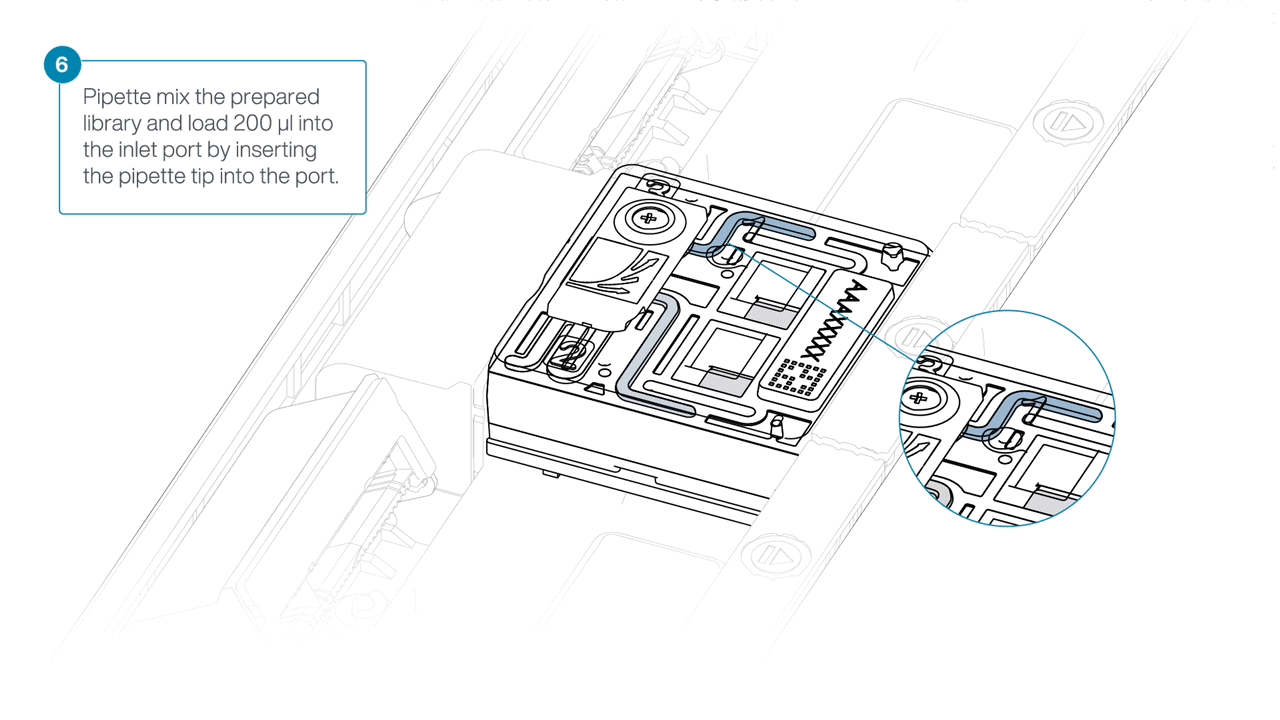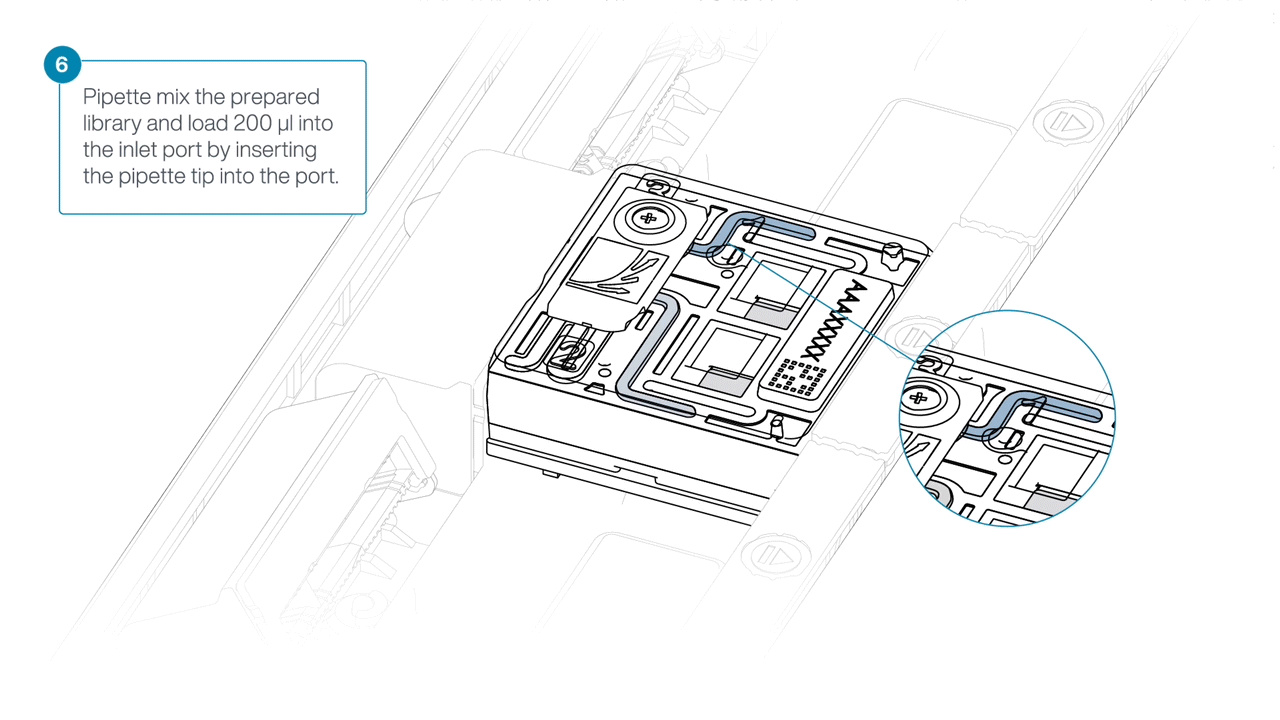
Mix the prepared library gently by pipetting up and down just prior to loading.
-
Load 200 µl of library into the inlet port using a P1000 pipette.
-
Close the valve to seal the inlet port.
-
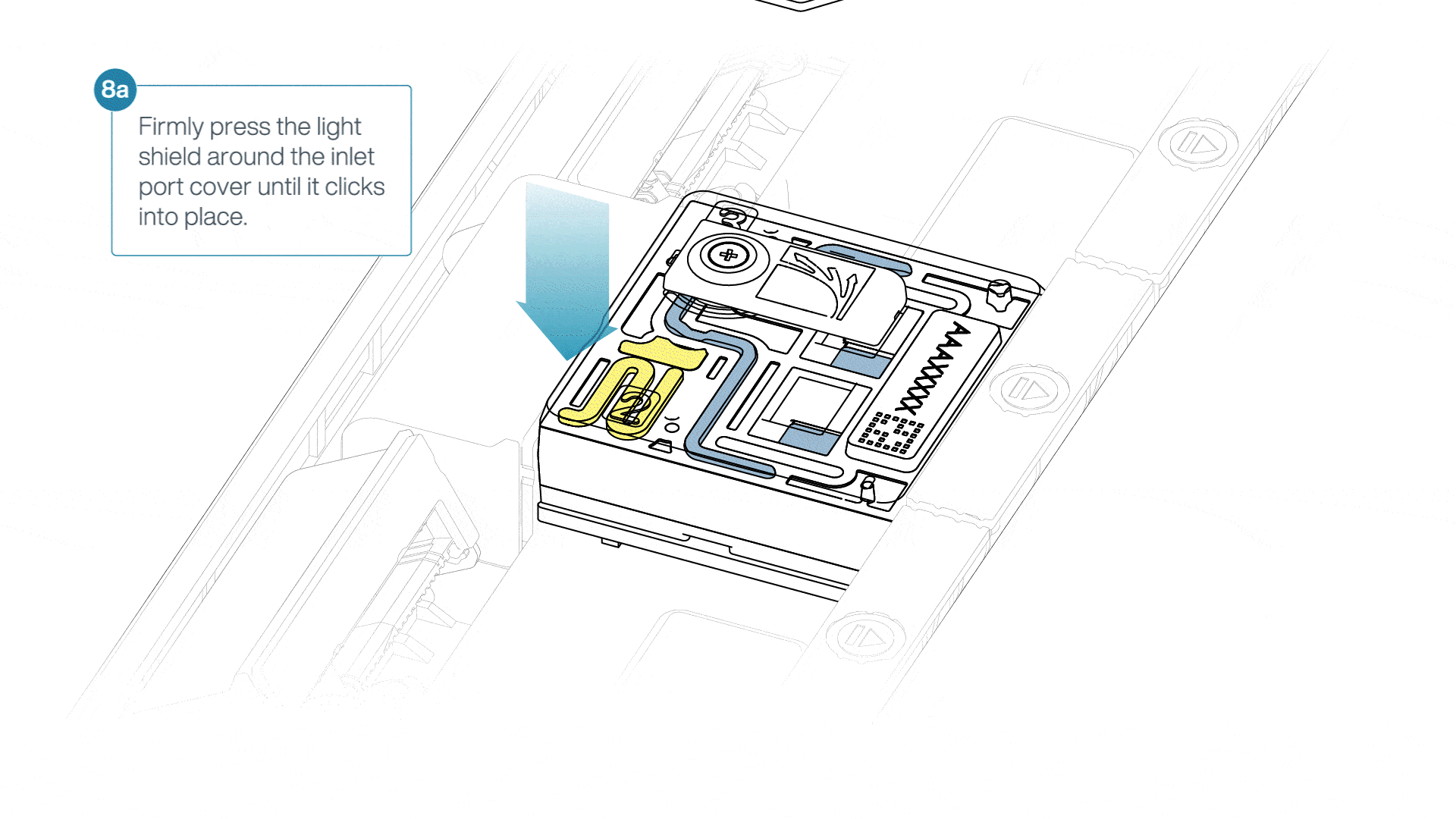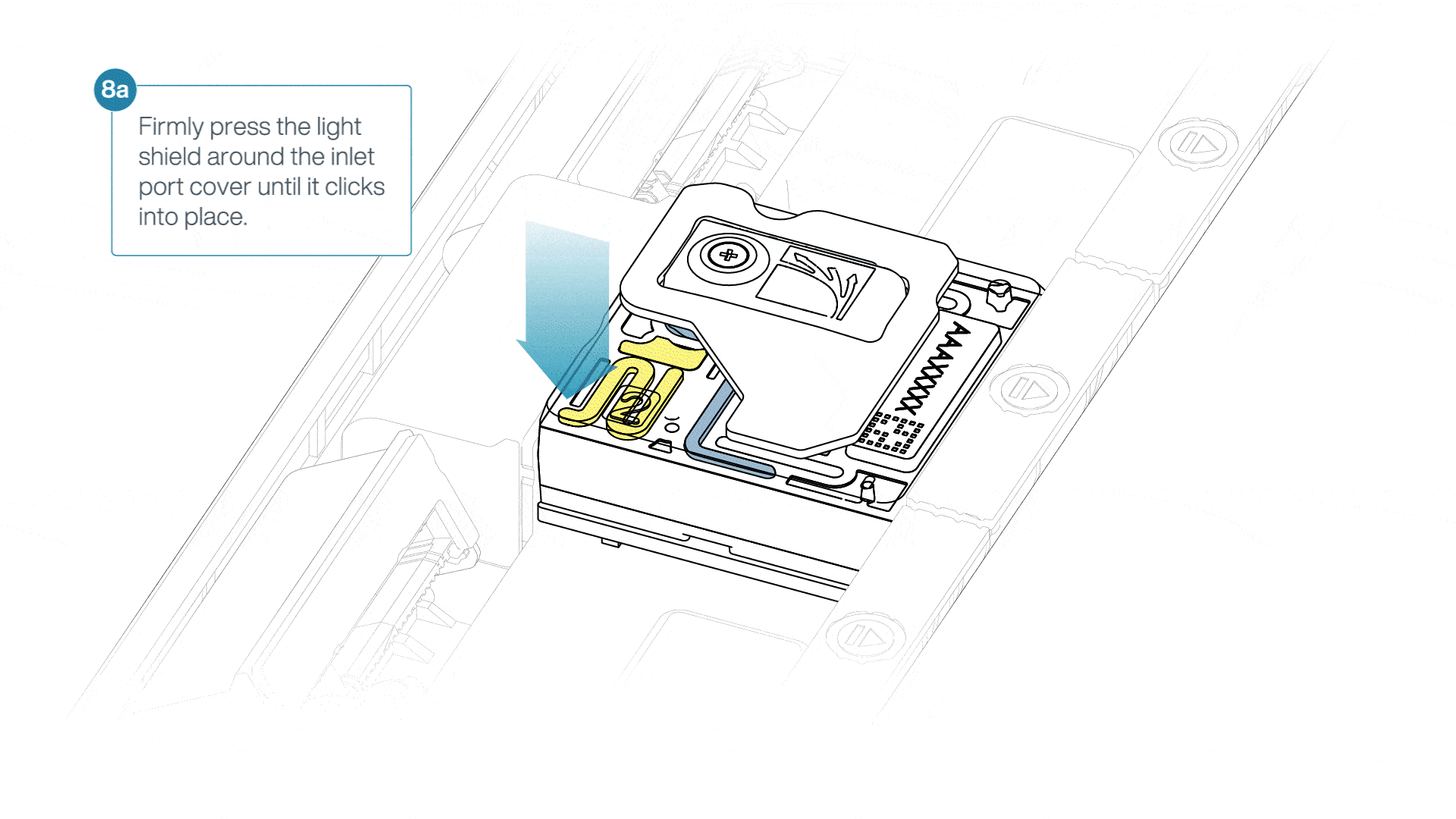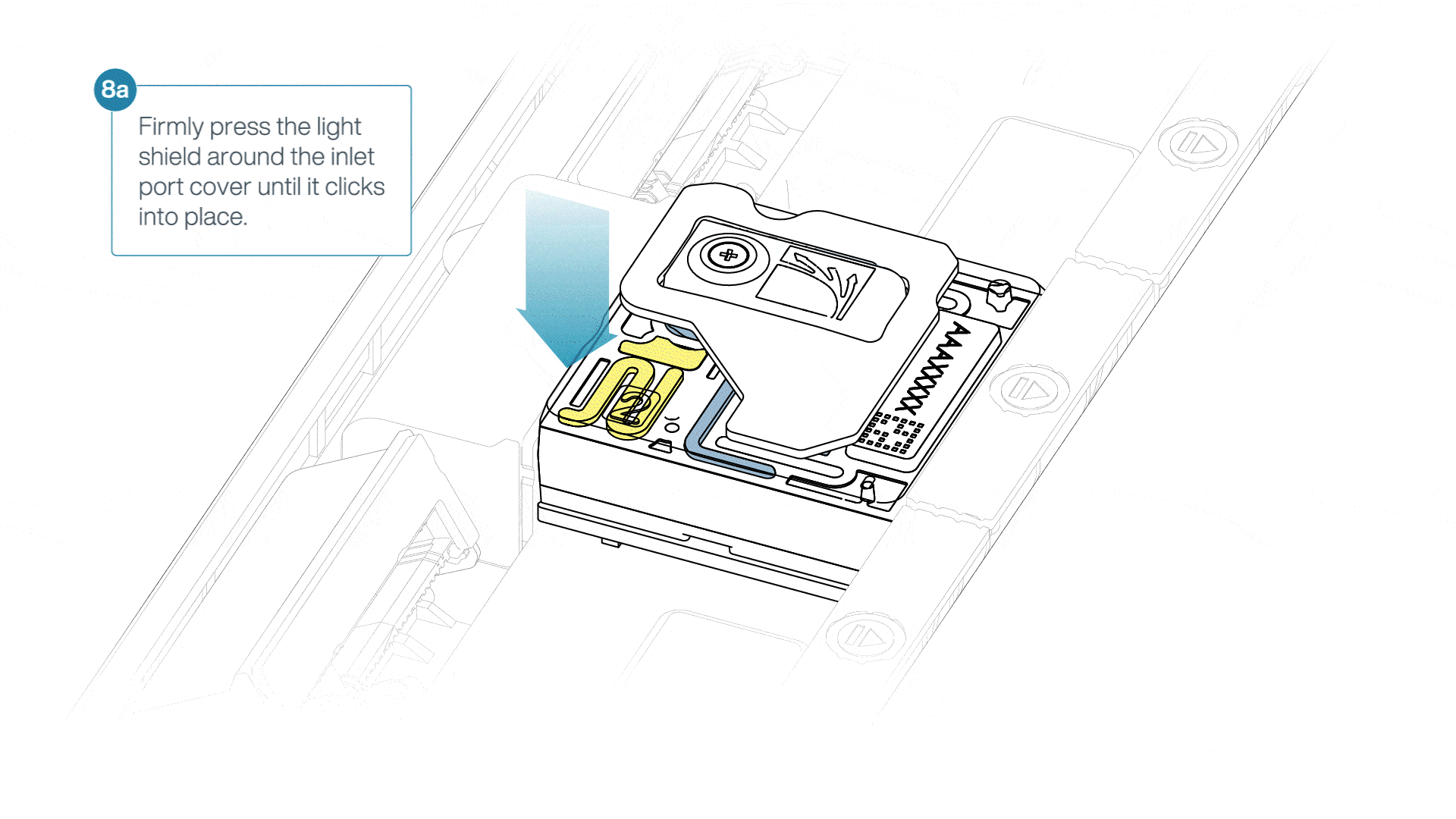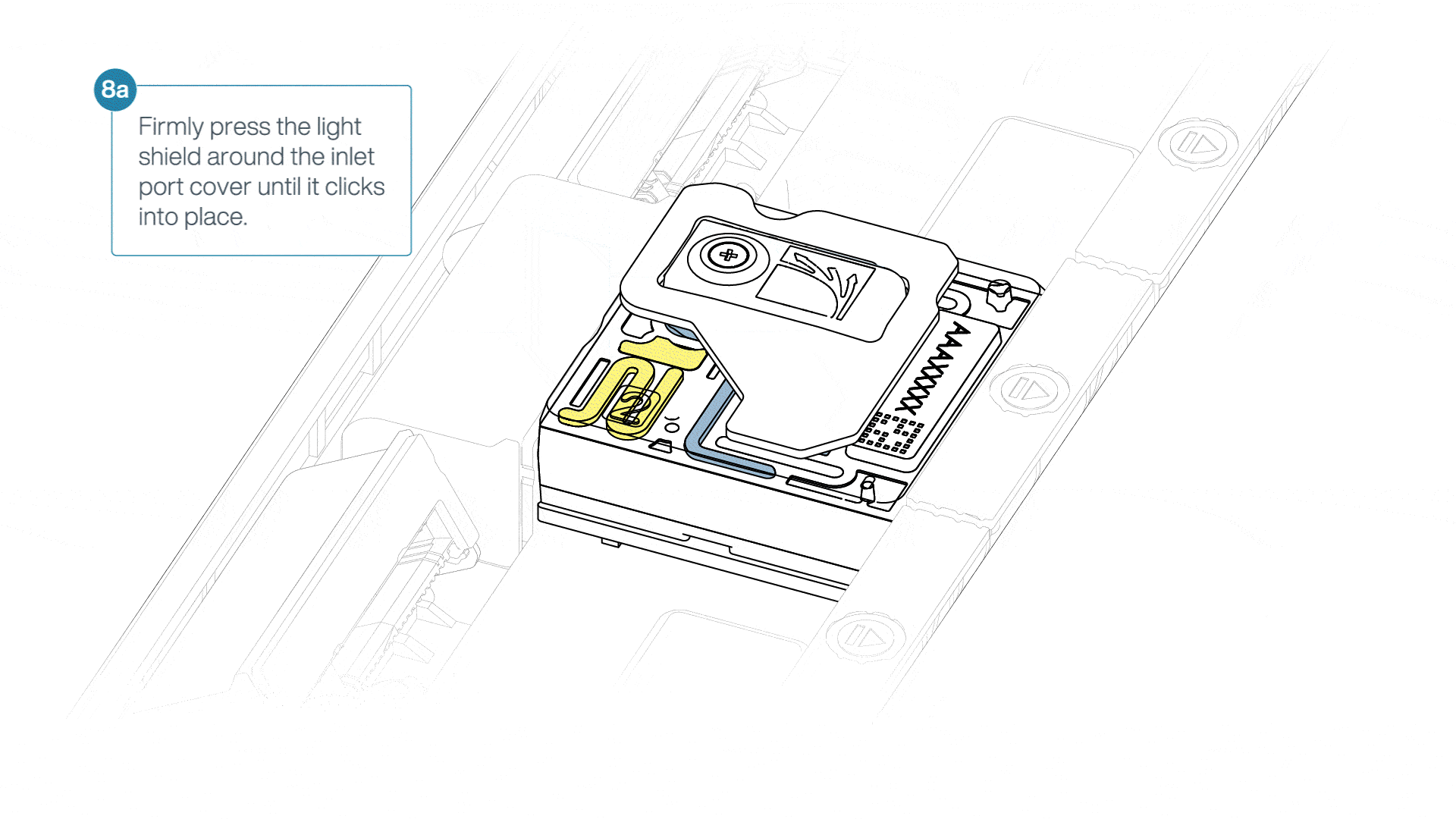
If the light shield has been removed from the flow cell, install the light shield as follows:
- Align the inlet port cut out of the light shield with the inlet port cover on the flow cell. The leading edge of the light shield should sit above the flow cell ID.
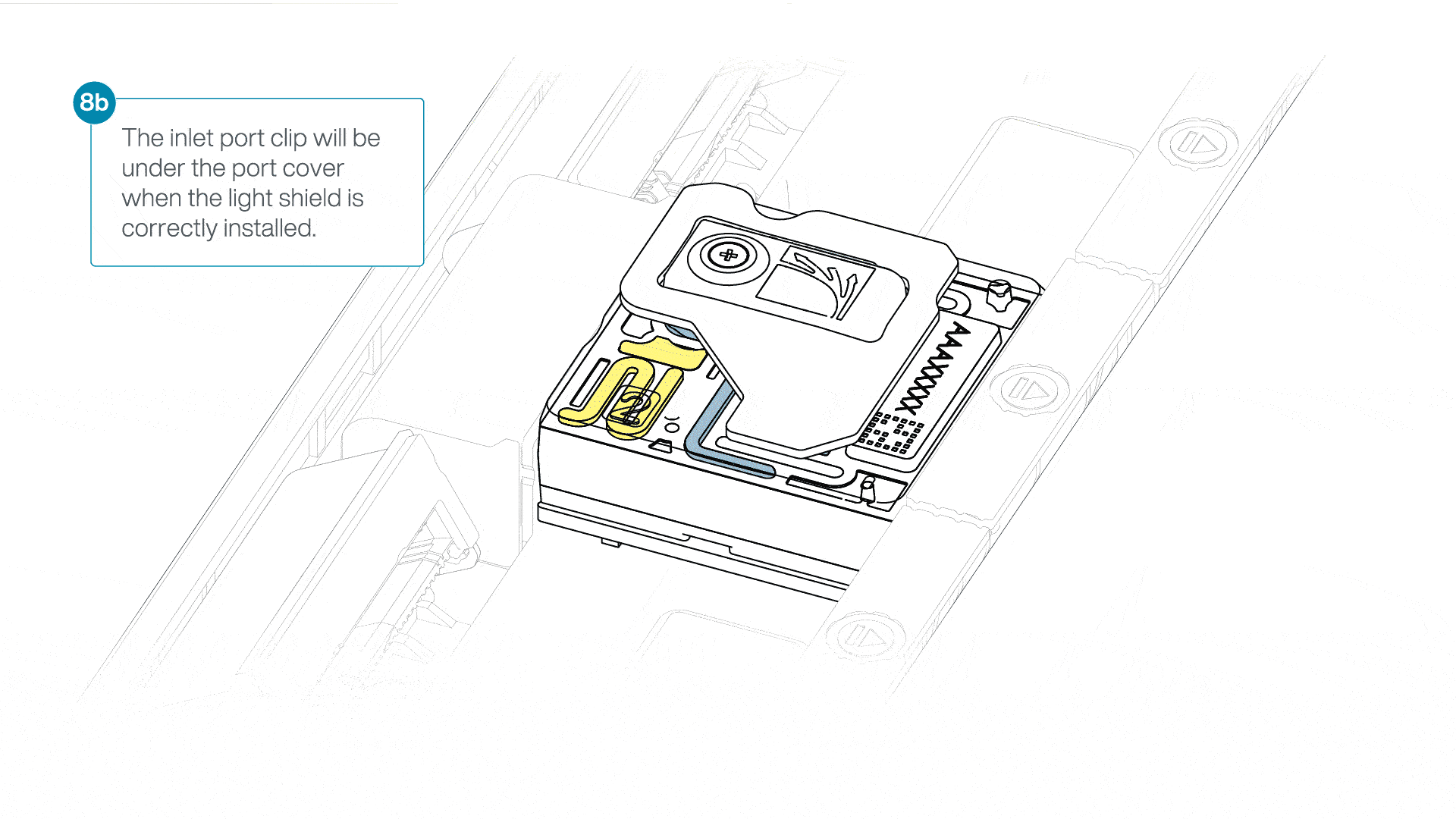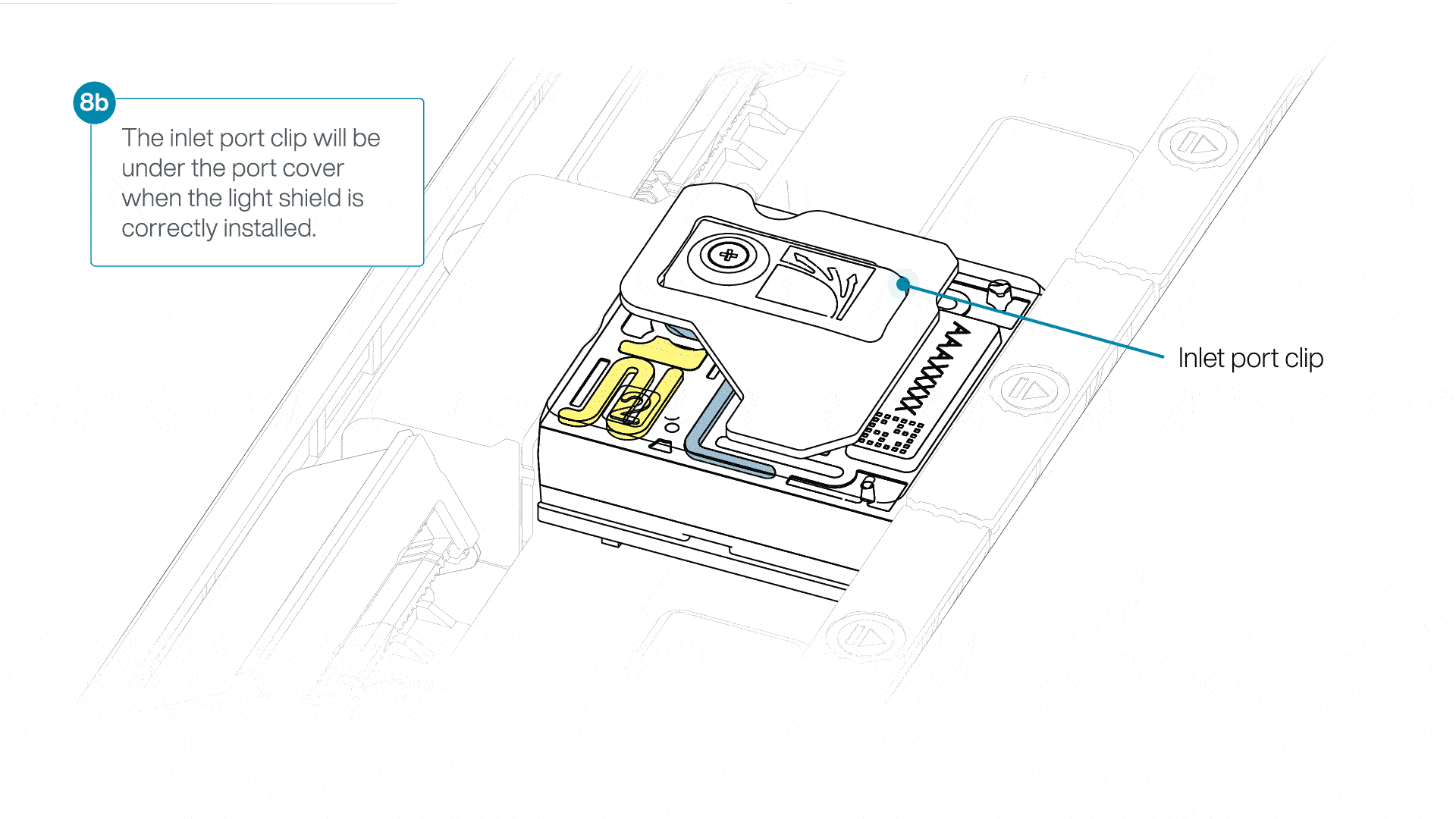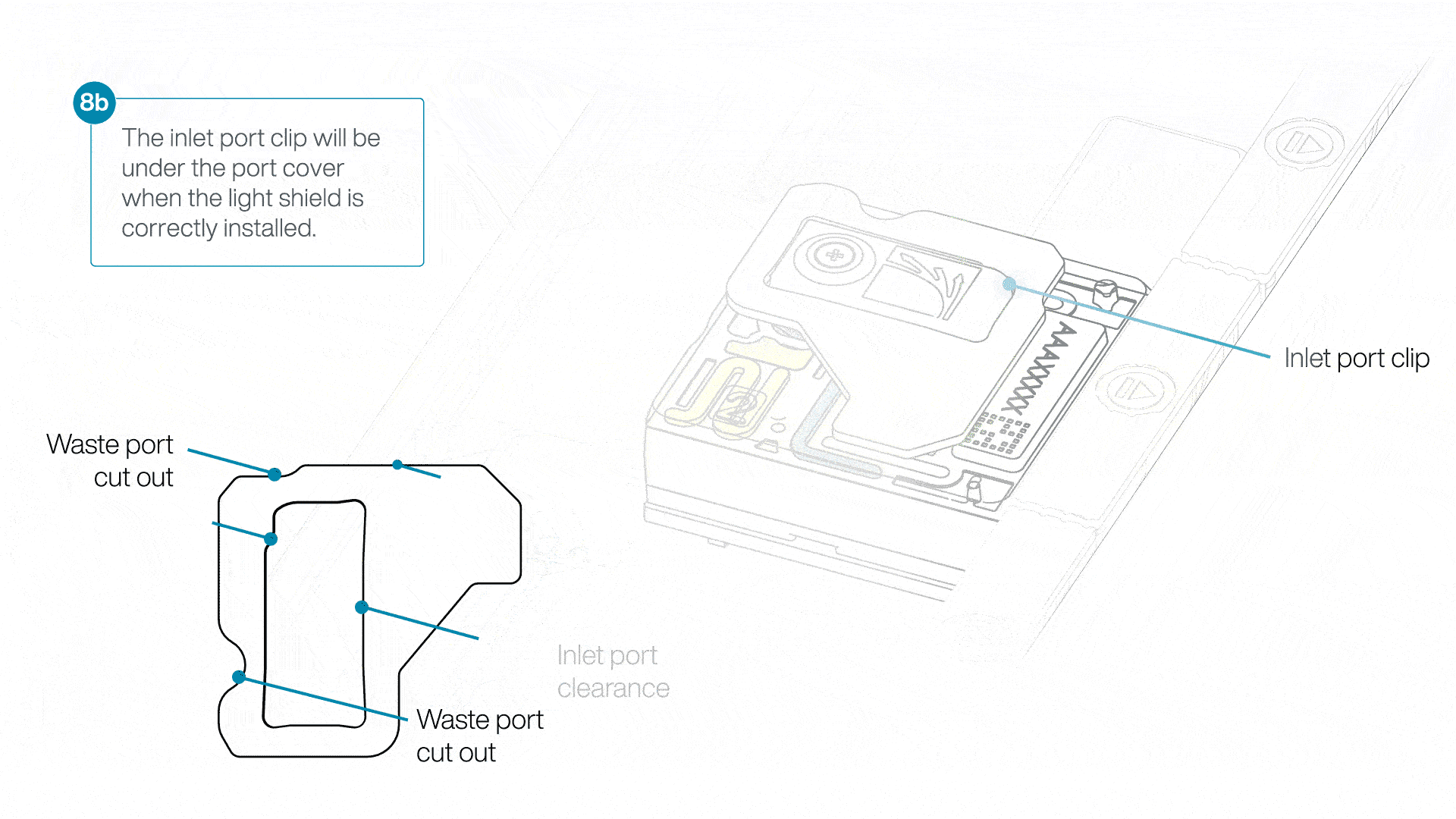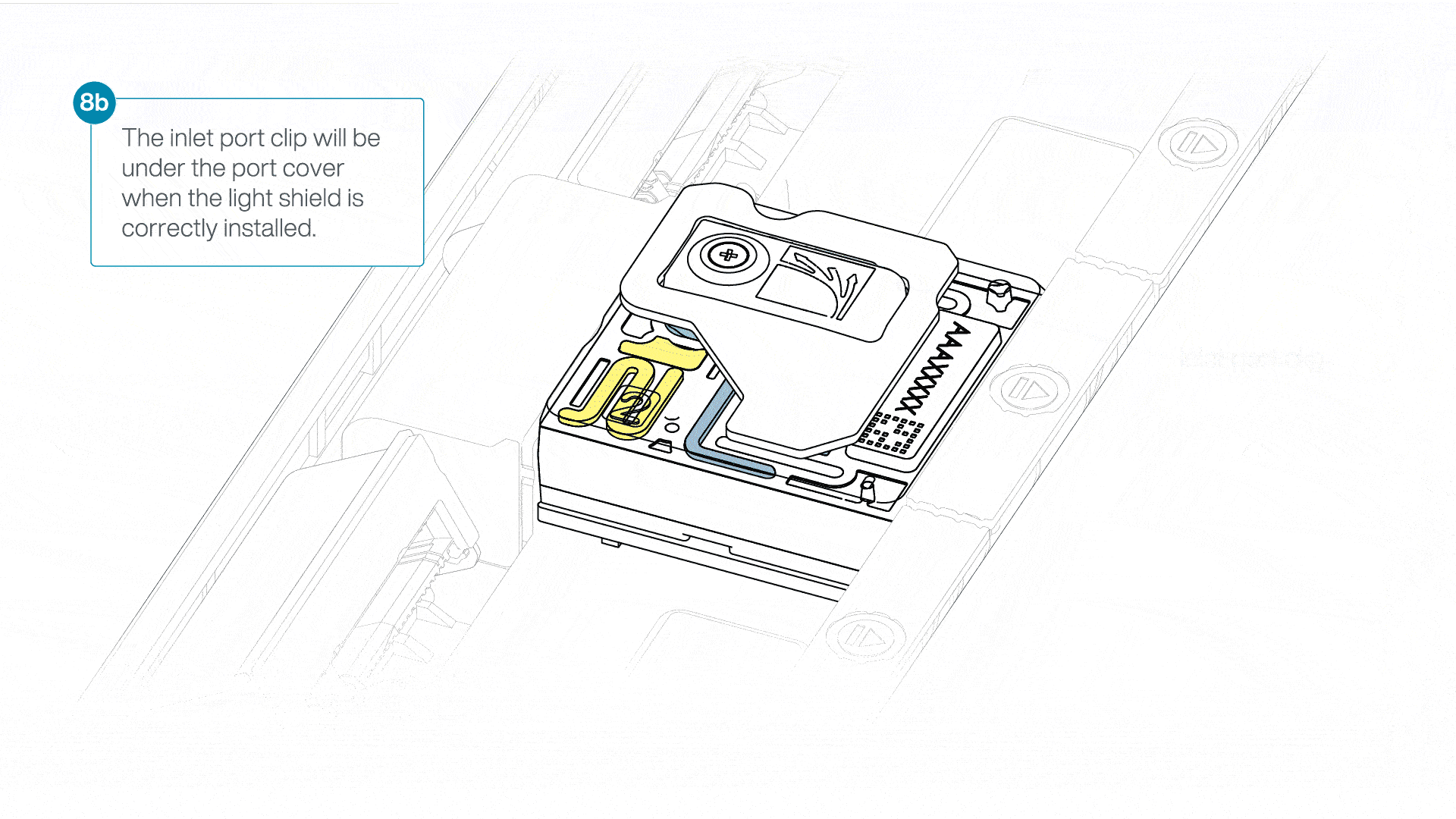
- Firmly press the light shield around the inlet port cover. The inlet port clip will click into place underneath the inlet port cover.
-
Close the PromethION lid when ready to start a sequencing run on MinKNOW.
Wait a minimum of 10 minutes after loading the flow cells onto the PromethION before initiating any experiments. This will help to increase the sequencing output.







