-
Pore activity plot introduction
The pore activity plot feature in the MinKNOW software can be used to judge the quality of your experiment. The pore activity plot shows the distribution of channel states over time, grouped by time chunks, or 'buckets'. The basic view shows the five main channel states: Sequencing, Pore, Recovering, Inactive, and Unclassified. Clicking the "More" button shows a more detailed breakdown of channel states.
It is recommended to observe the pore activity plot populating over the first 30 min-1 hr of the sequencing run. By this time, the channel state distribution will give an indication whether the DNA/RNA library is of a good quality, and whether the flow cell is performing well.
If Active Channel Selection is enabled during the run, the software instantly switches to a new channel in the group if a channel is in the “Saturated” or “Multiple” state, or after ~5 minutes if a channel is “Recovering”. This feature maximises the number of channels sequencing at the start of the experiment, however this may also result in an artificially high number of "Sequencing" or "Pore" channels in the pore activity plot. For this reason, we recommend referring to the pore scan plot, which shows the true distribution of channel states at the point of the most recent pore scan.
-
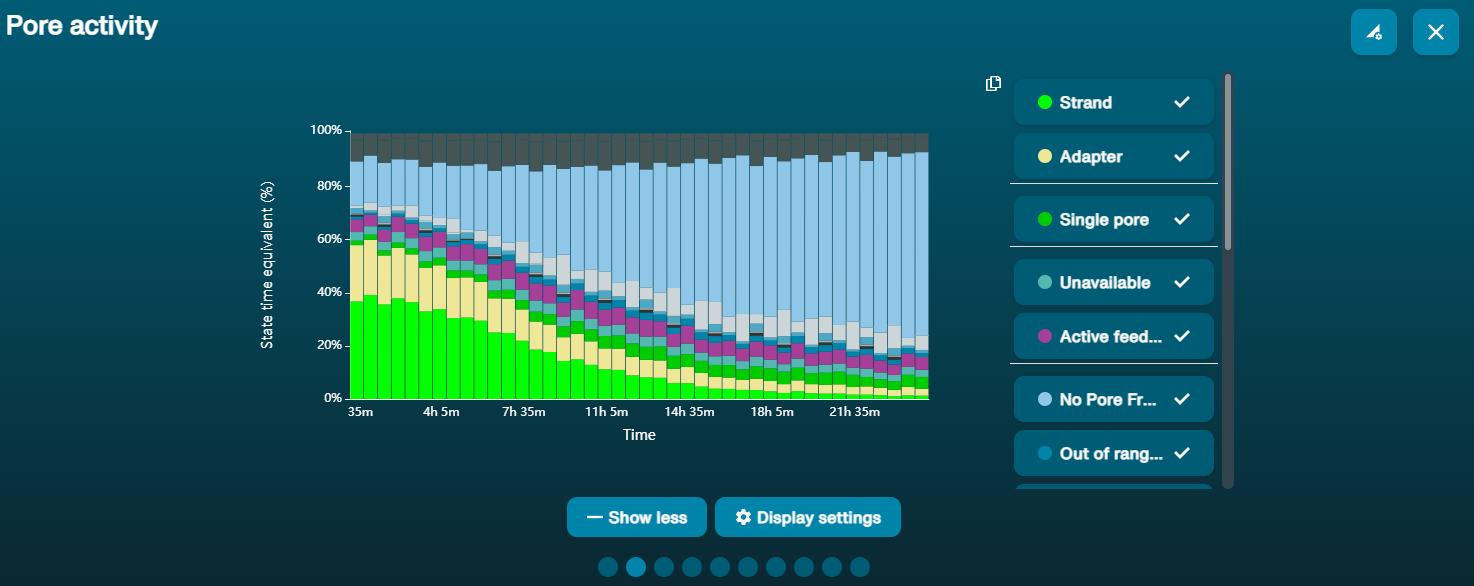
Good library
A good quality library will result in most of the pores being in the "Sequencing" state, and very few in "Pore", "Recovering" or "Inactive". A library that looks like this is likely to give a good sequencing output.
-
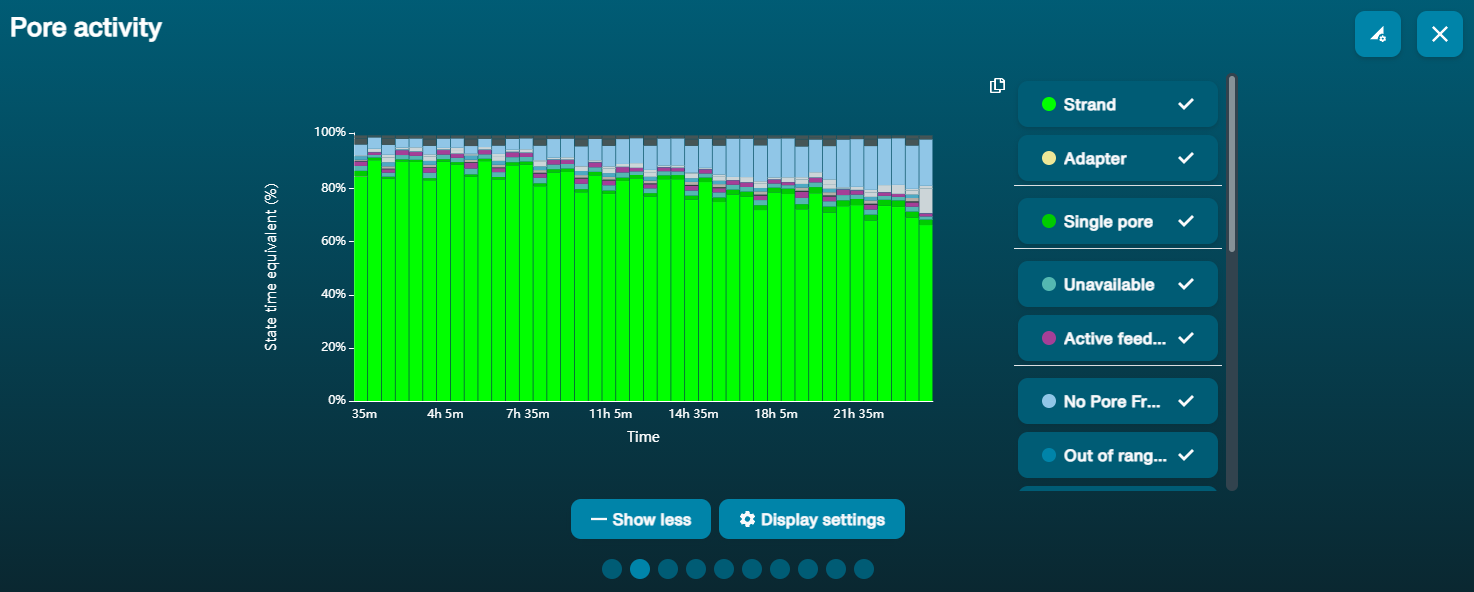
Channel blocking
Under certain conditions (usually the presence of contaminants in the library), pores may become blocked and therefore unable to sequence. This manifests itself as a build-up of "Unavailable" pores over time.
Recommendation:
- If, despite the channel blocking, the library is still producing a sufficient number reads to answer your biological question, you can carry on with the sequencing experiment.
- Otherwise, stop the sequencing run in MinKNOW. Then wash out the library from the flow cell using the instructions for the Flow Cell Wash Kit. Then prepare another library and load it on the flow cell.
-
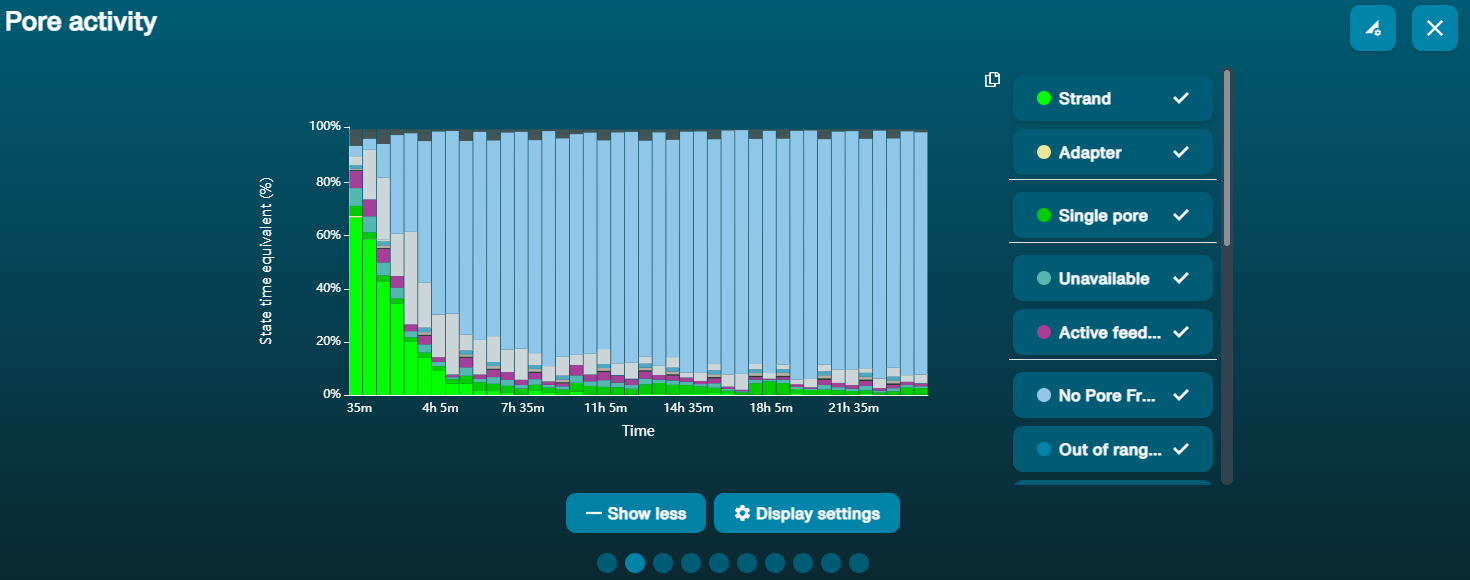
Osmotic imbalance
If the plot shows a high number of "Inactive" channels building up over time, this could indicate that the channels or membranes have been damaged, for example by air bubbles, osmotic imbalance, or the presence of detergents or surfactants in the library.
Pore activity:
Pore scan:
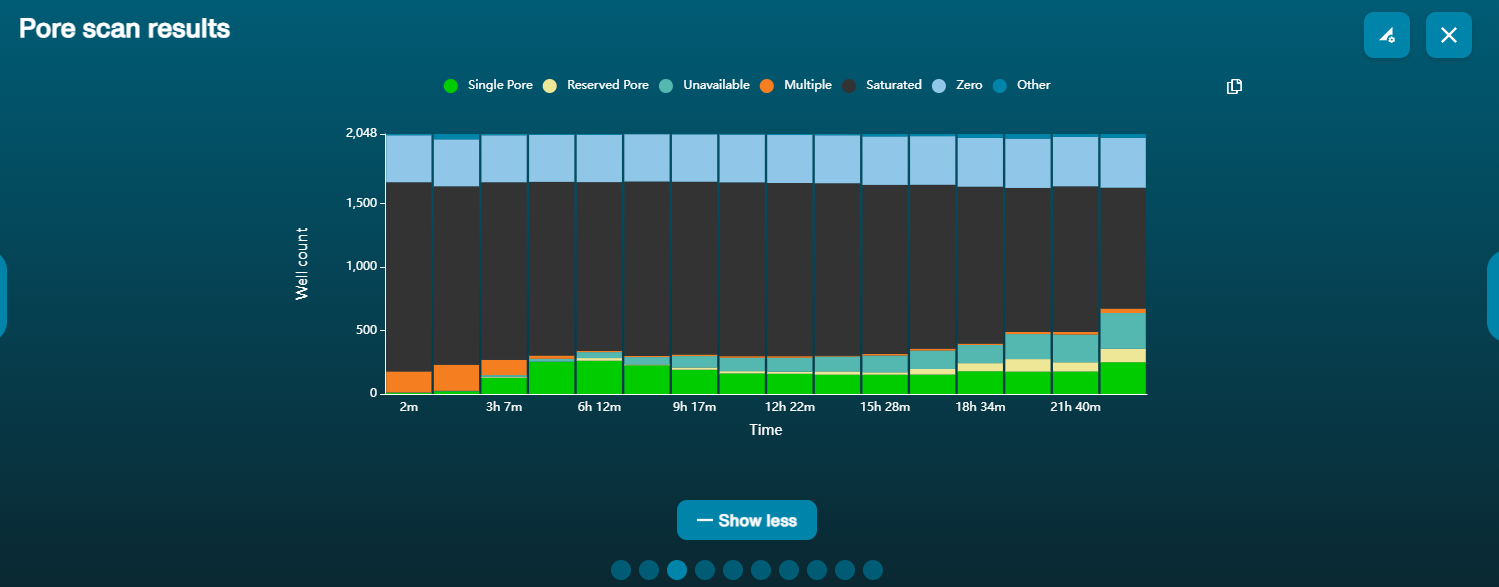
Recommendation:
- Check the channel panel: if the Inactive channels are all grouped in one part of the flow cell, this could indicate an air bubble that has been introduced during flow cell flushing or library loading. If the remaining channels are still sequencing, it is possible to carry on with the run. Do not try to move the air bubble, as this can damage even more channels.
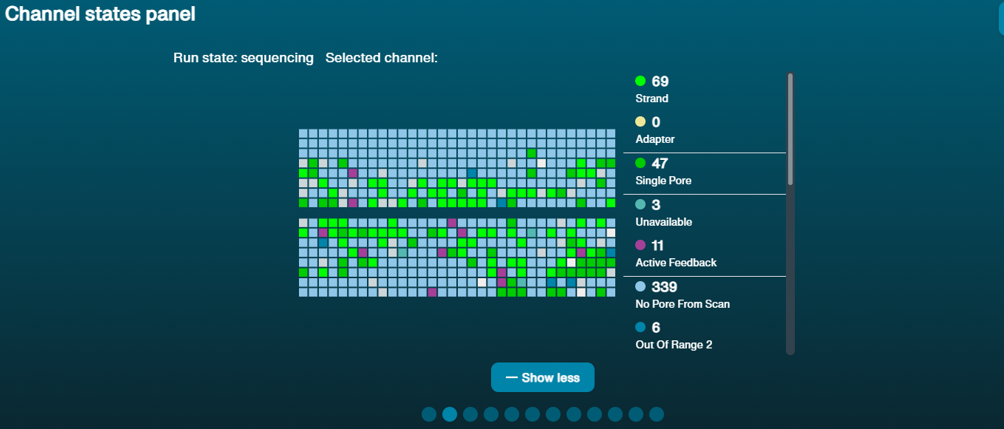
If the Inactive channels are distributed throughout the flow cell:
- Check that the heat tape on the underside of the flow cell is intact.
- Make sure that the input DNA is in either TE buffer or nuclease-free water, and that the buffer contains no detergents or surfactants.
- Make a new batch of flow cell priming buffer (a mixture of Flush Buffer/Flow Cell Flush and Flush Tether/Flow Cell Tether). Flush the flow cell with the mixture and load the library again.
-
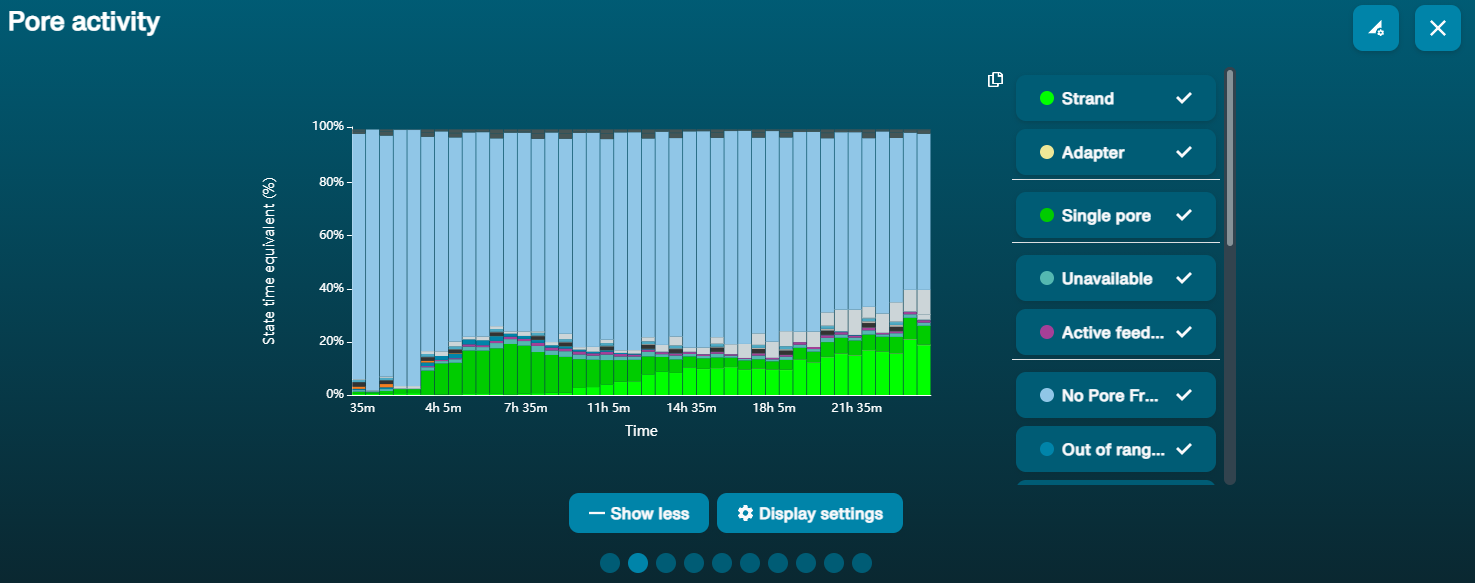
Low pore occupancy
If there was insufficient starting material, or some sample has been lost during library prep, or the sequencing adapters did not ligate well to the strand ends, the plot will show a high ratio of "Pore available" to "Sequencing" states, meaning that only a limited number of pores are sequencing at any one time.
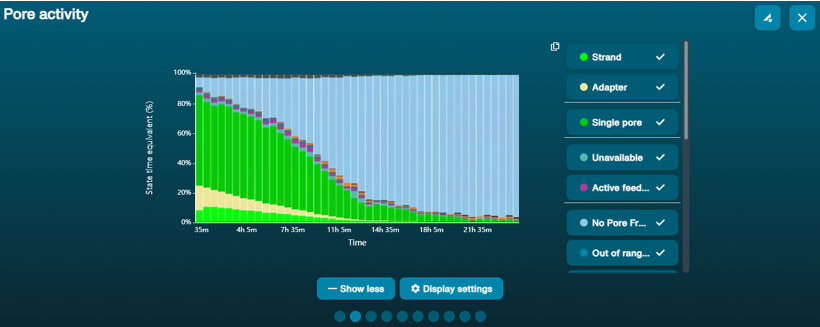
Recommendation:
- Check the amount of DNA/RNA in your prepared library, for example by using the Qubit fluorometer. We recommend preparing a fresh library and reloading your flow cell with the recommended loading input from the relevant protocol
- If your library is at a low concentration, prepare the library again using a higher amount of starting material.
- Ensure you are adding the flush tether to your priming mix during flow cell priming and loading before adding your library.
- For ligation-based protocols, ensure you are following the protocol to ligate the sequencing adapter correctly and performing any clean-up steps after adapter ligation with short or long fragment buffer rather than ethanol.
-
RNA pore activity
For RNA in particular, the expanded channels view may show a large proportion of pores sequencing Adapter. This happens because RNA strands are usually shorter than DNA, and the adapter takes up a larger proportion of the strand. Additionally, the RNA sequencing chemistry is optimised for sequencing RNA, whereas the adapter is DNA, and is processed slower. As long as the 'basic' pore activity plot view shows the majority of pores in "Sequencing", a high proportion of Adapter should not be a problem.