-
Catch-up notifications
Catch-up basecalling is carried out either after the sequencing run completes (e.g. after 72 hours), or if the run is manually terminated. To manually terminate a run, select sequencing flow cells and click Stop.
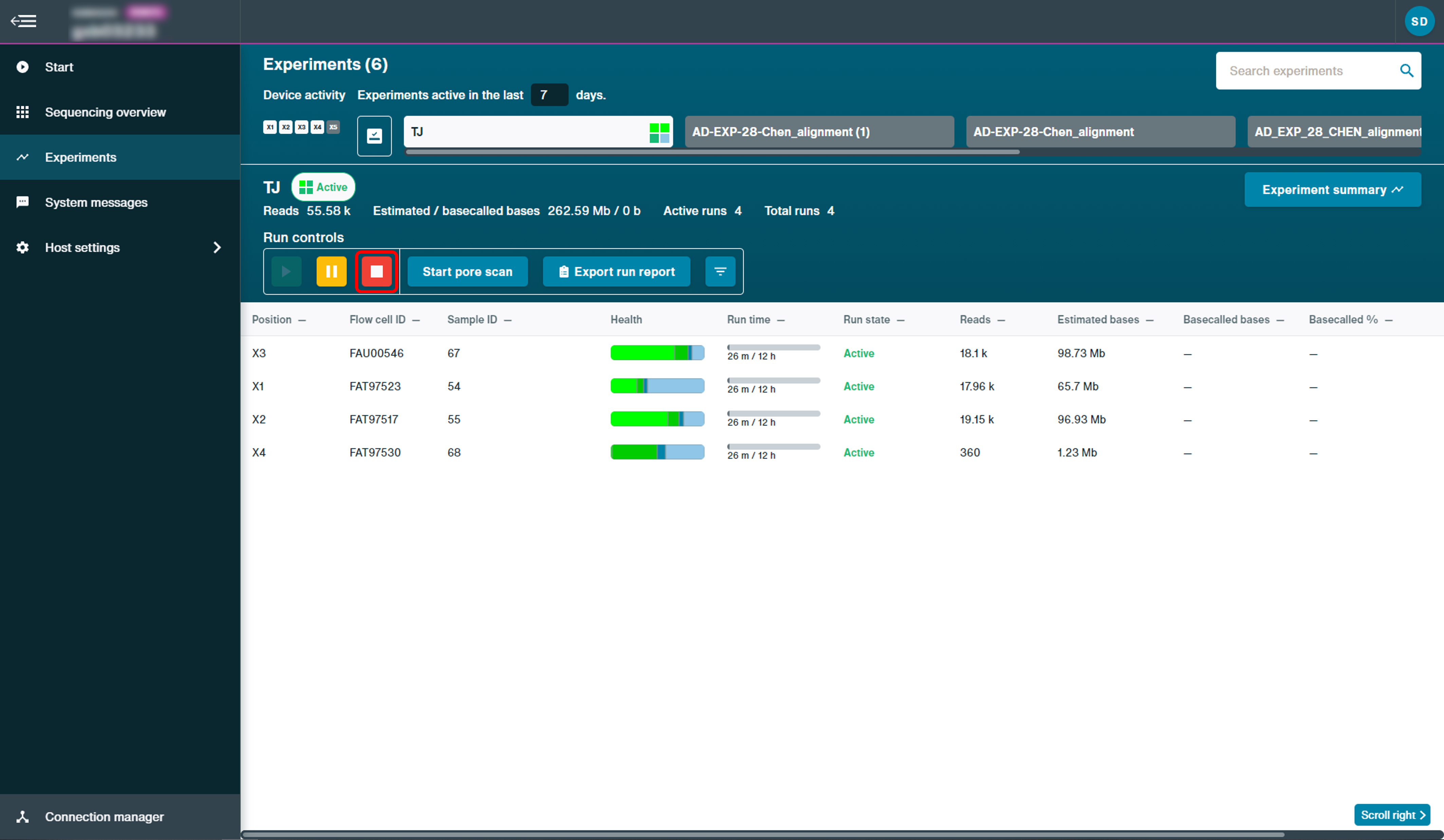
A pop-up window will appear:
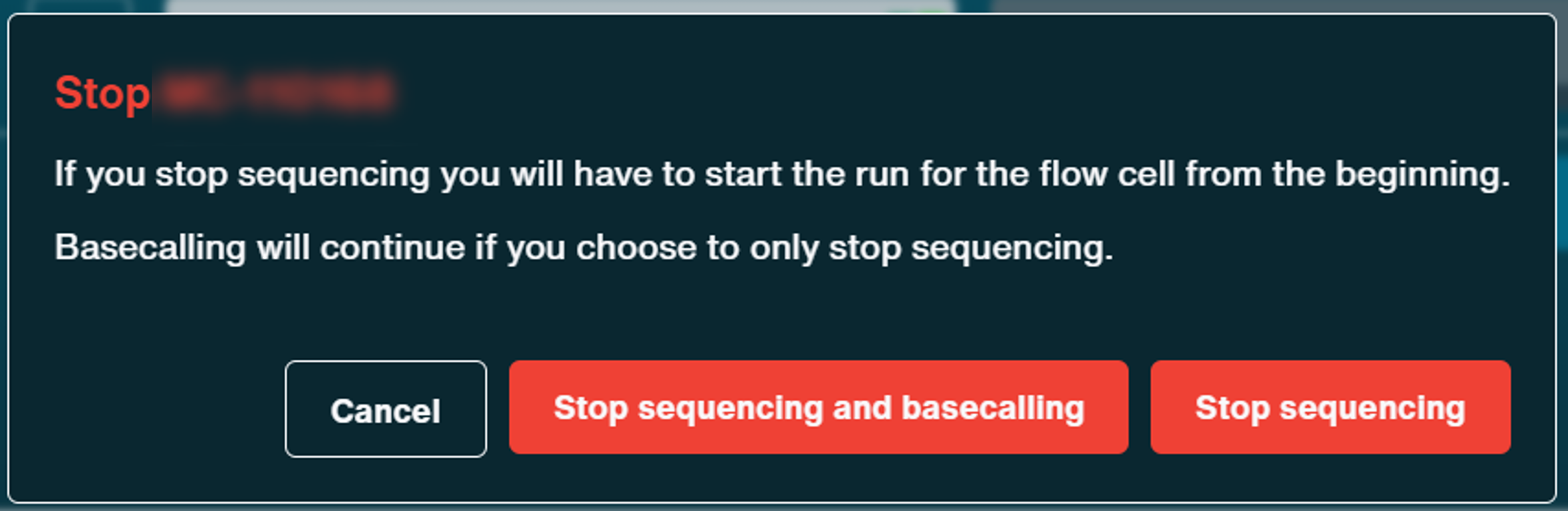
A progress bar is displayed during catch-up basecalling on the Sequencing Overview and Experiments pages.
-
CSV files output by MinKNOW
The data used by MinKNOW to generate the pore activity and Cumulative Output plots are output as CSV files. These files are in the run-specific folder in the reports directory (e.g. \data\reports on Windows).
Using the data in the CSV files, users can recreate pore activity and Cumulative Output graphs themselves in Excel by adding the various pore states to the 'Legend Entries (series)' panel:
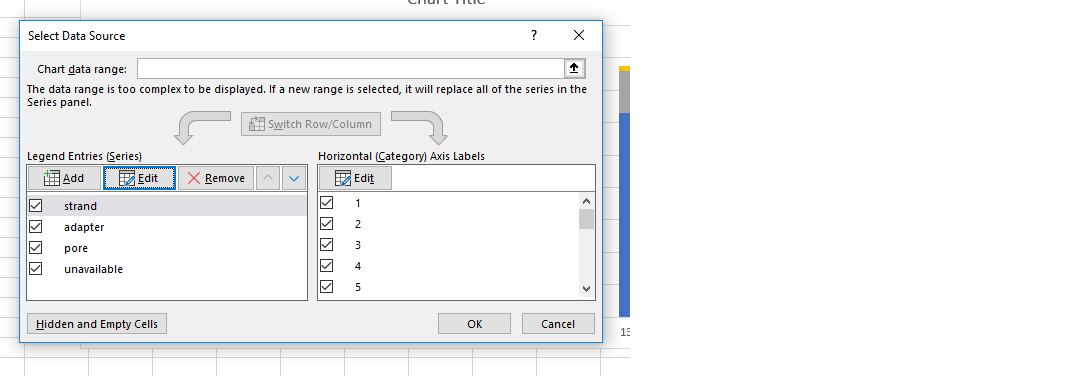
Below are definitions of the columns in the output CSV file from MinKNOW:
Column title Description Experiment time (minutes) Time in minutes from the beginning of the sequencing run. Reads Cumulative number of reads basecalled since the beginning of the sequencing run. Basecalled reads passed Cumulative number of reads basecalled since the beginning of the sequencing run with a Qscore above the filter threshold. Basecalled reads failed Cumulative number of reads basecalled since the beginning of the sequencing run with a Qscore below the filter threshold. Basecalled reads skipped Cumulative number of reads that have not been basecalled since the beginning of the sequencing run. Selected raw samples Raw analog-to-digital converter (ADC) samples that have been identified by MinKNOW as part of a read. Selected events Measure MinKNOW uses to estimate bases and distinct from events used in basecalling. Estimated bases Number of (MinKNOW) events multiplied by some fixed factor defined per chemistry. Basecalled bases Number of estimated bases that have been in all selected reads above. Basecalled samples Number of basecalled bases that have been in all selected reads above. -
Acquisition termination file for LIMS
Customers using LIMS will receive a summary file once data acquisition and file writing has completed. The file is named final_summary.txt, and is written out to the output directory.
An example of the file contents is shown below:
instrument=PCT0009 position=1-A5-D5 flow_cell_id=PAD15520 sample_id=Chinese_NA24631 protocol_group_id=JB_47x_P48005_270319 protocol=4a37d56a62b6425a2af4d01d4970c7b5cc7b7c80 protocol_run_id=2a4ccaa1-8aaf-4872-a1f3-773a686d64de acquisition_run_id=7eeaea330d1fe2ec43583072bbe21288254bef1b started=2019-03-27T17:23:55.542284+00:00 acquisition_stopped=2019-03-28T09:58:37.505691+00:00 processing_stopped=2019-03-28T09:58:37.505788+00:00 basecalling_enabled=1 sequencing_summary_file=sequencing_summary/PCT0009_20190327_172337_PAD15520_promethion_sequencing_run_Chinese_NA24631_sequencing_summary.txt fast5_files_in_final_dest=1606 fast5_files_in_fallback=0 fastq_files_in_final_dest=1197 fastq_files_in_fallback=0


