-
Run reports overview
Run reports contain information about the sequencing run and include performance graphs. These graphs are automatically generated at the end of a run but can be manually generated by clicking Export run report and selecting which experiment to export to html.
The run report includes the run summary and configuration, sequence output, run health, and run log. This html report is interactive, allowing users to jump to the information they need and hide other details they do not require. Troubleshooting suggestions are also available throughout the report, with links to further information available on the Community.
-
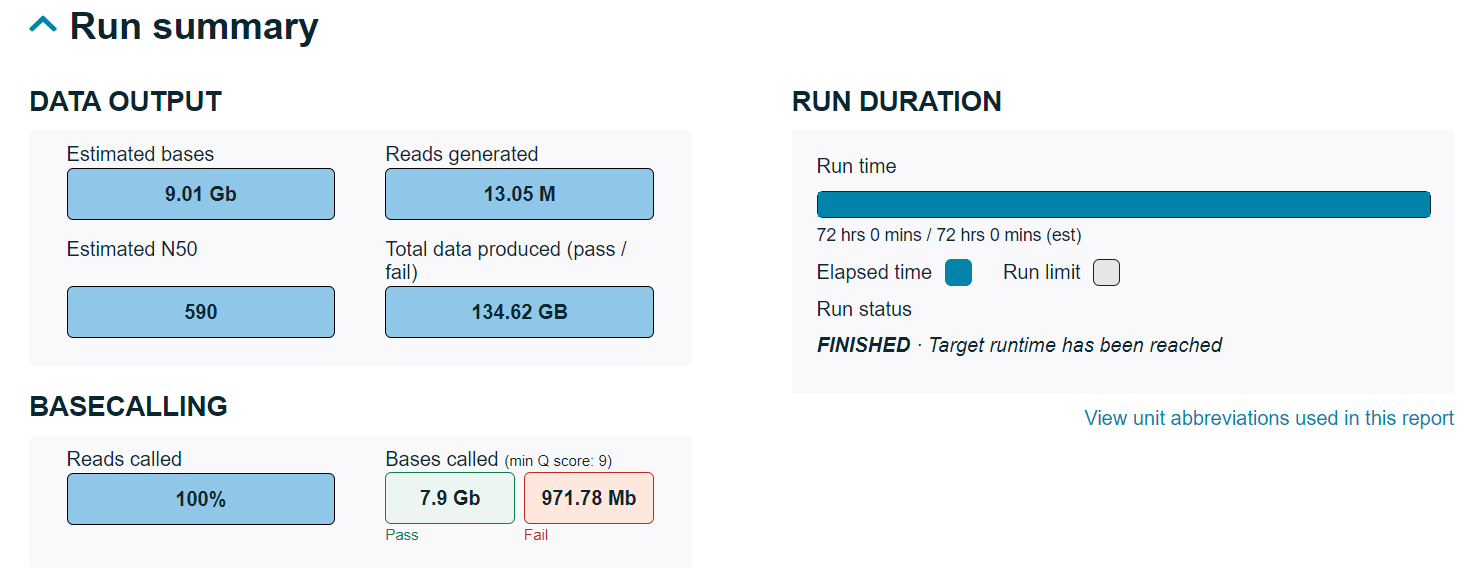
Run summary
This is an overview of the sequencing run, including output and baseacalling results along with the duration of the run.
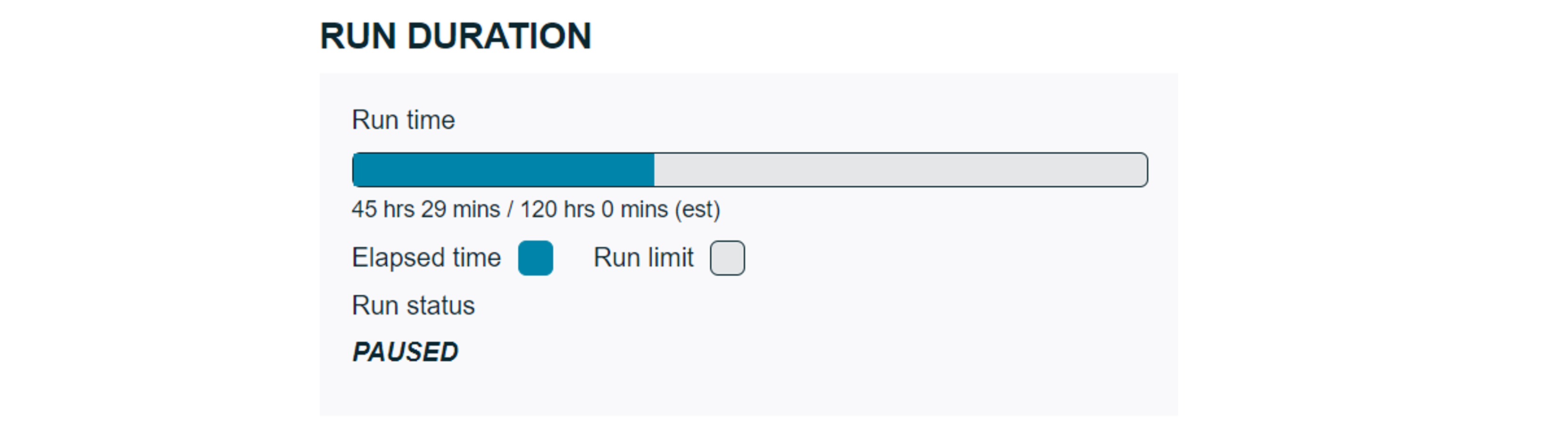
During a run:
When a run report has been generated during a run, the elapsed time will show in the progress bar, along with:- The estimated time until flow cell end of life (if selected)
- The estimated time until the data target condition will be met and if the flow cell will reach the target during the run time or flow cell life (if selected). A warning will appear under the progress bar if the data target will not be met.
If a run report was generated when a sequencing run is paused, the run status will only display "PAUSED" with no progress bar or estimations.
After a run:
When a run report is generated after a sequencing run has completed, the duration of the run will be displayed, along with:- The run status confirming the flow cell end of life condition was met and completed progress bar.
- The run status confirming the data target was met and a partially filled progress bar reaching the data target. If the data target was not met, a warning will appear under the completed progress bar.
Note: The relevant run until parameters must be defined during the sequencing run set up for these status reports to appear on the run report.
-
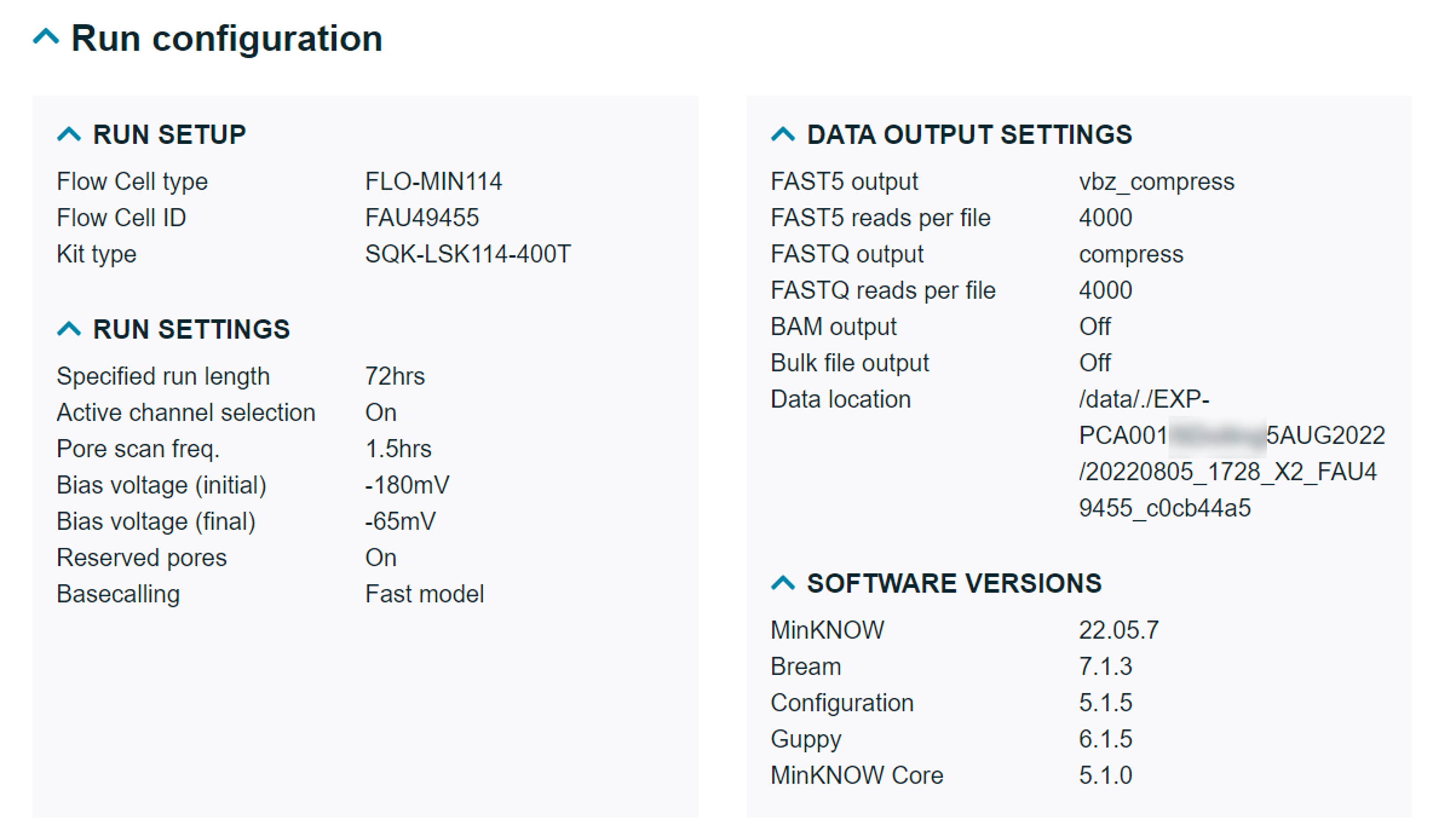
Run configuration
A detailed overview of the settings selected for the sequencing run and data output, including the software versions used to sequence the data. We have also included further information about the modified bases and basecalling models chosen, barcode balancing settings, expansion kits used and read splitting preferences selected.
-
Sequence output
This section includes a more detailed view of the sequencing output, such as read lengths, barcodes detected and quality score.
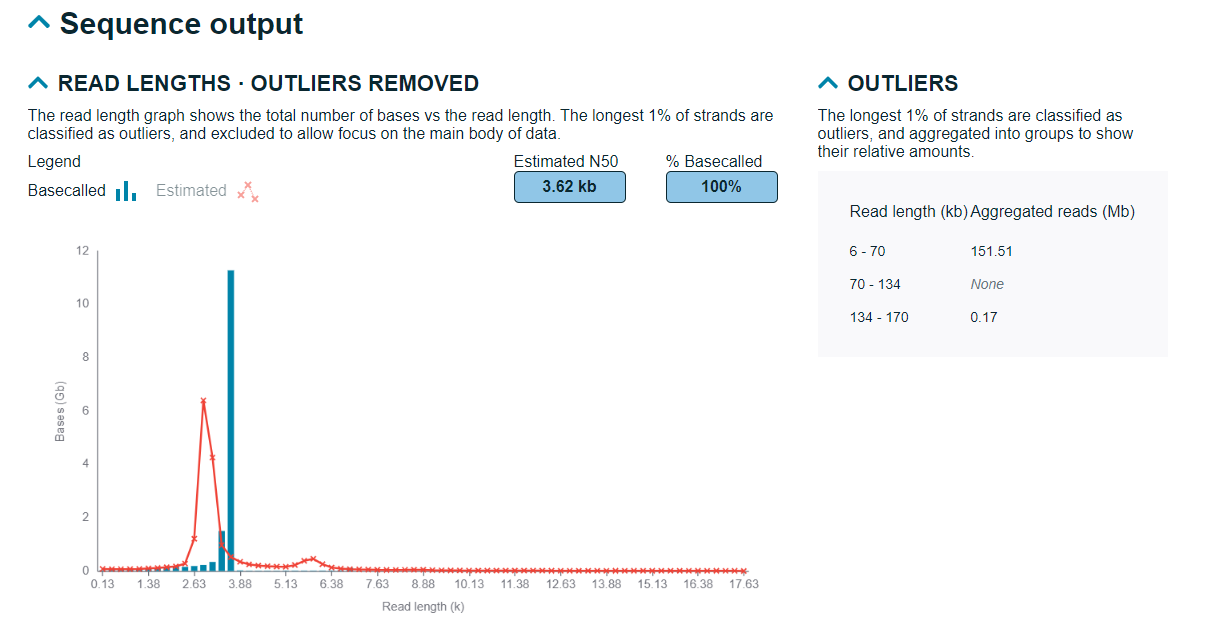
Read lengths
The total number of bases against the read length are displayed in this graph. Any outliers displayed in a table below, displaying read length and aggregated reads.
Barcodes
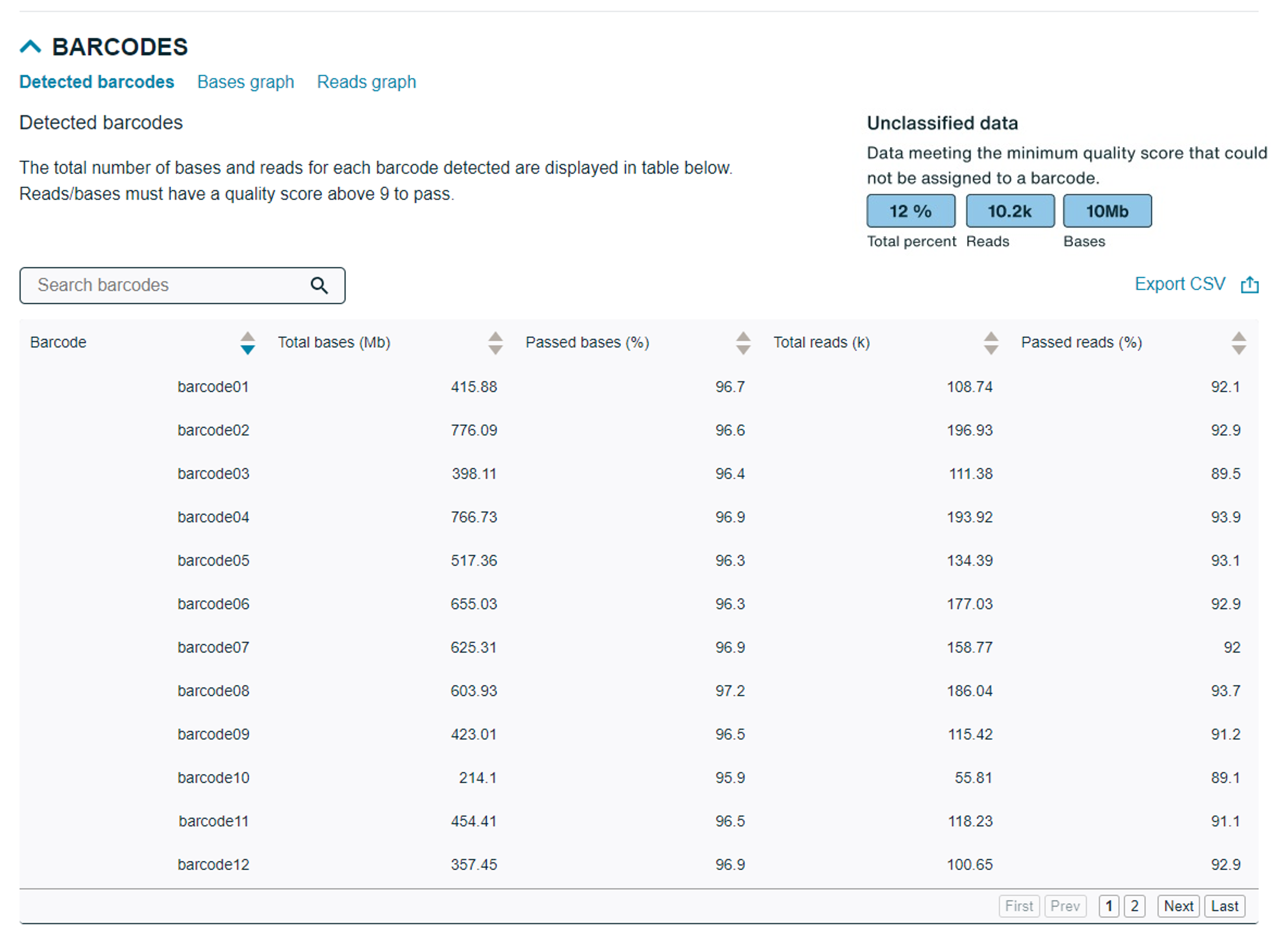
Barcodes detected are displayed in a searchable table which can be exported as a .CSV file.
Two graphs for bases and reads are available to view and displays total number of reads/bases above a Qscore of 9. Passed and failed data is also displayed and can be sorted by barcode.Detected barcodes:
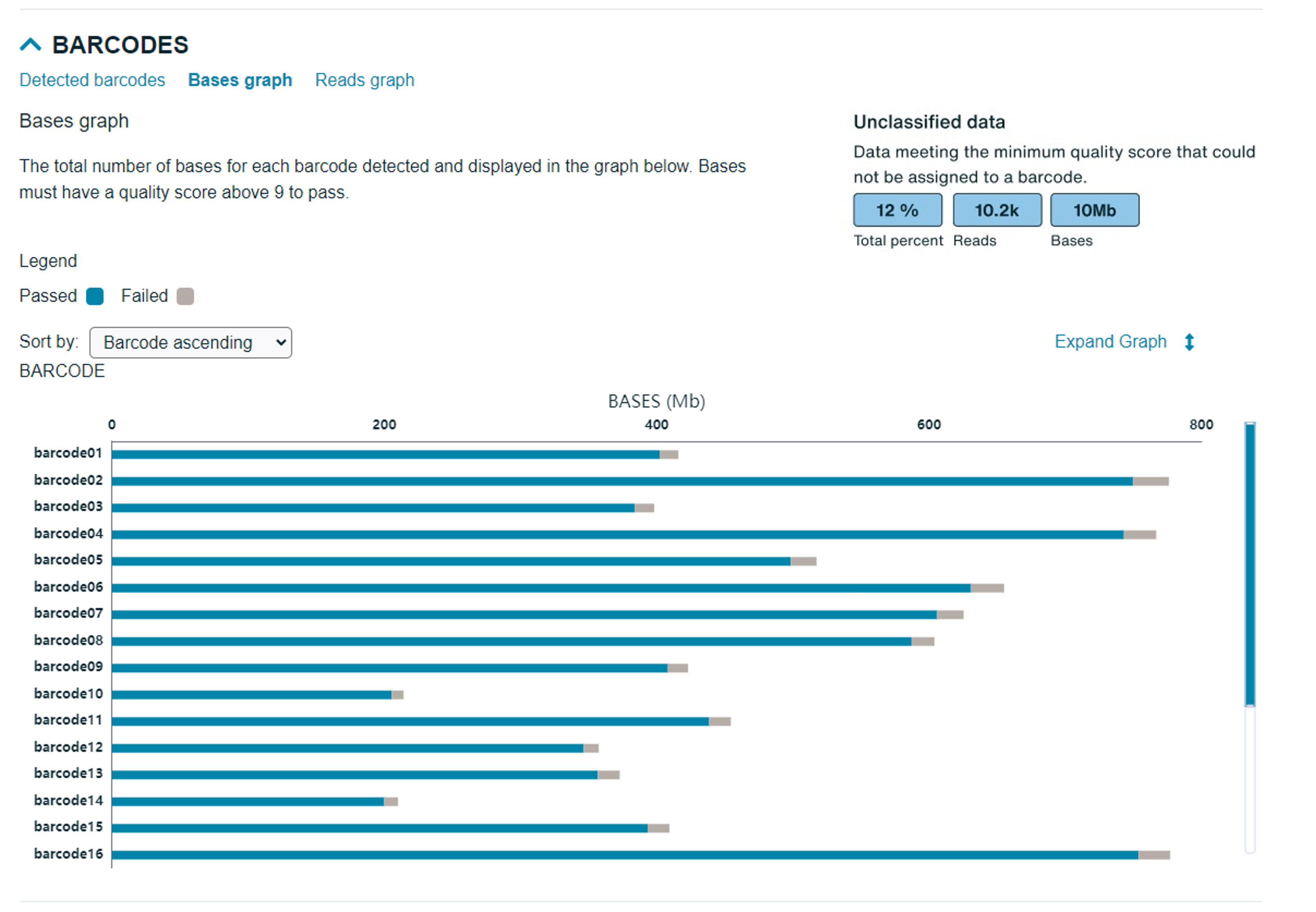
Bases graph:
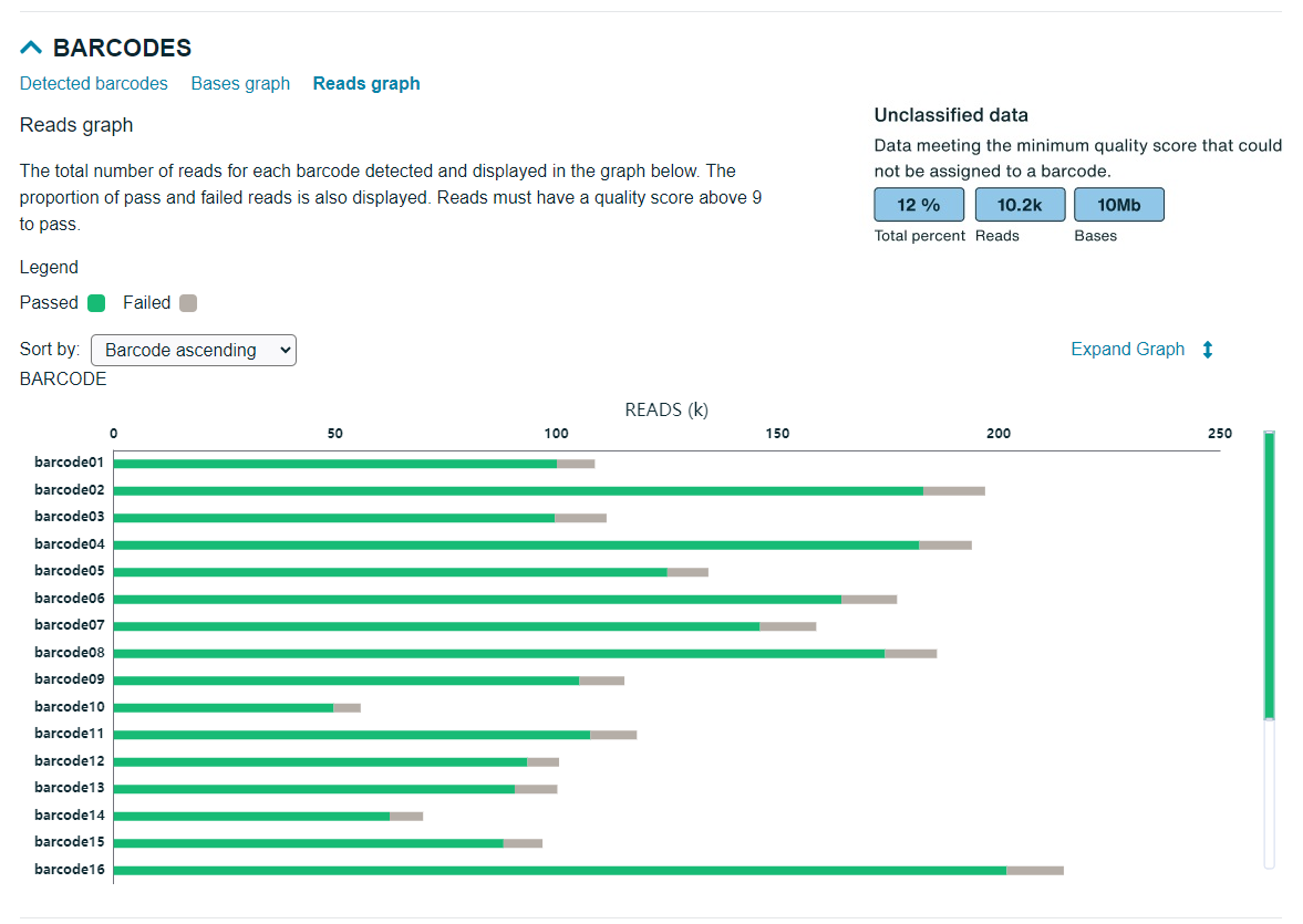
Reads graph:
Cumulative output
Total output of the bases and reads sequenced during the experiment are outlined in the graphs here.
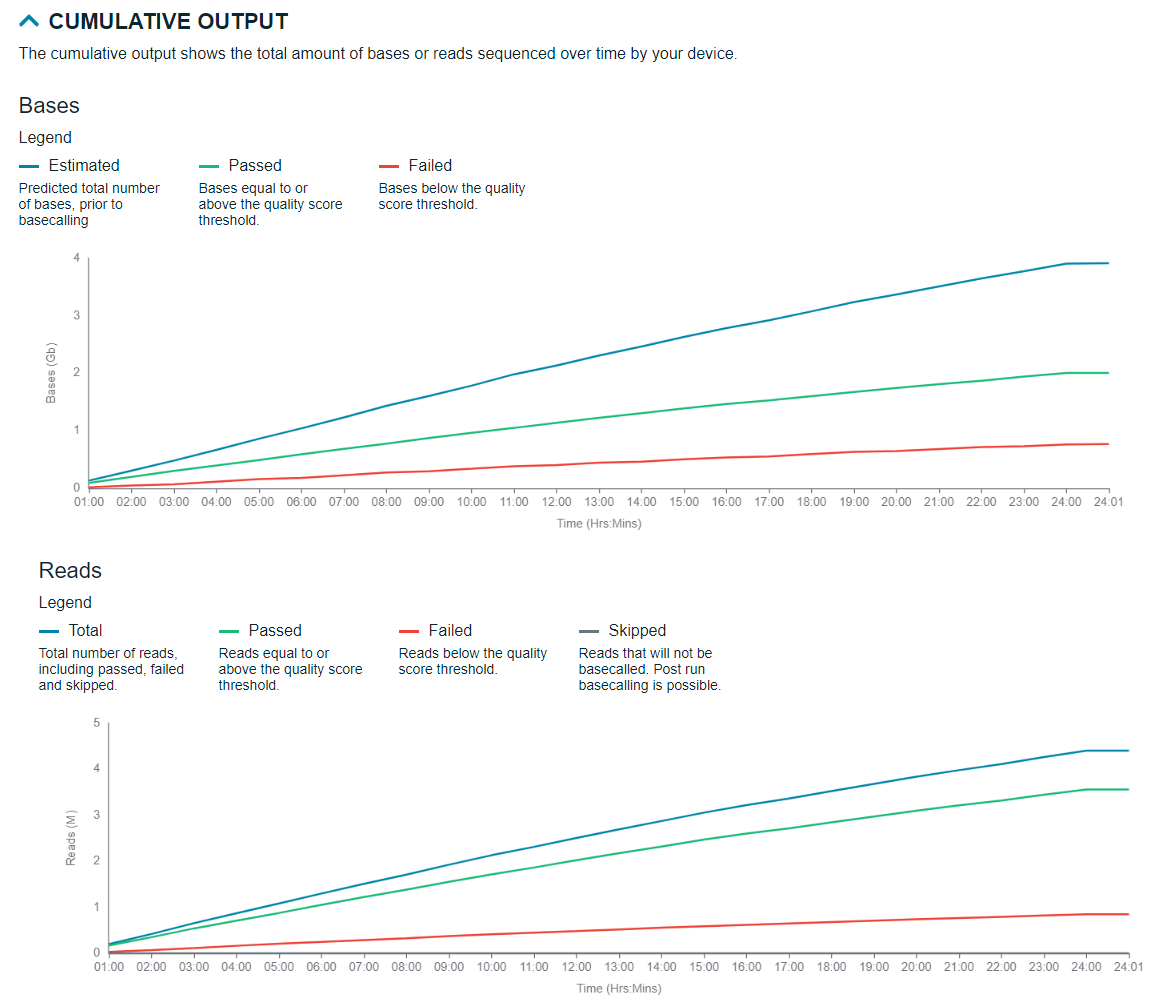
Quality score
Quality score (Qscore) of your sequencing run is displayed here, with the minimum value set on MinKNOW during run set up.
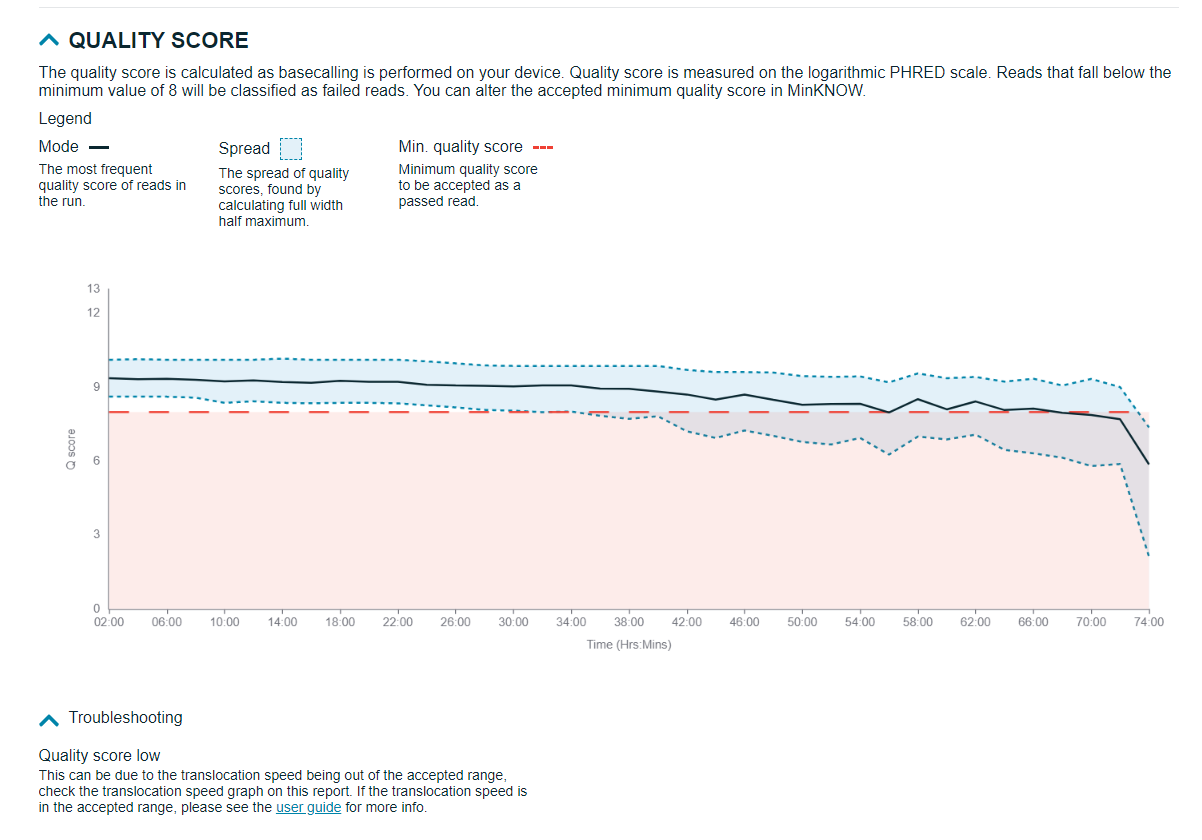
Alignment
If an alignment file was supplied and alignment turned on during run set-up, the Reference alignment and Bed regions sections will be displayed with tables that can be exported as CSV files.Reference alignment:
This table will display the specified reference sequence and the number of reads, bases and coverage percentage.Bed region:
This table will display the specified reference sequence with the coordinates of the specific target within the reference sequence as well as the number of reads, bases and coverage percentage.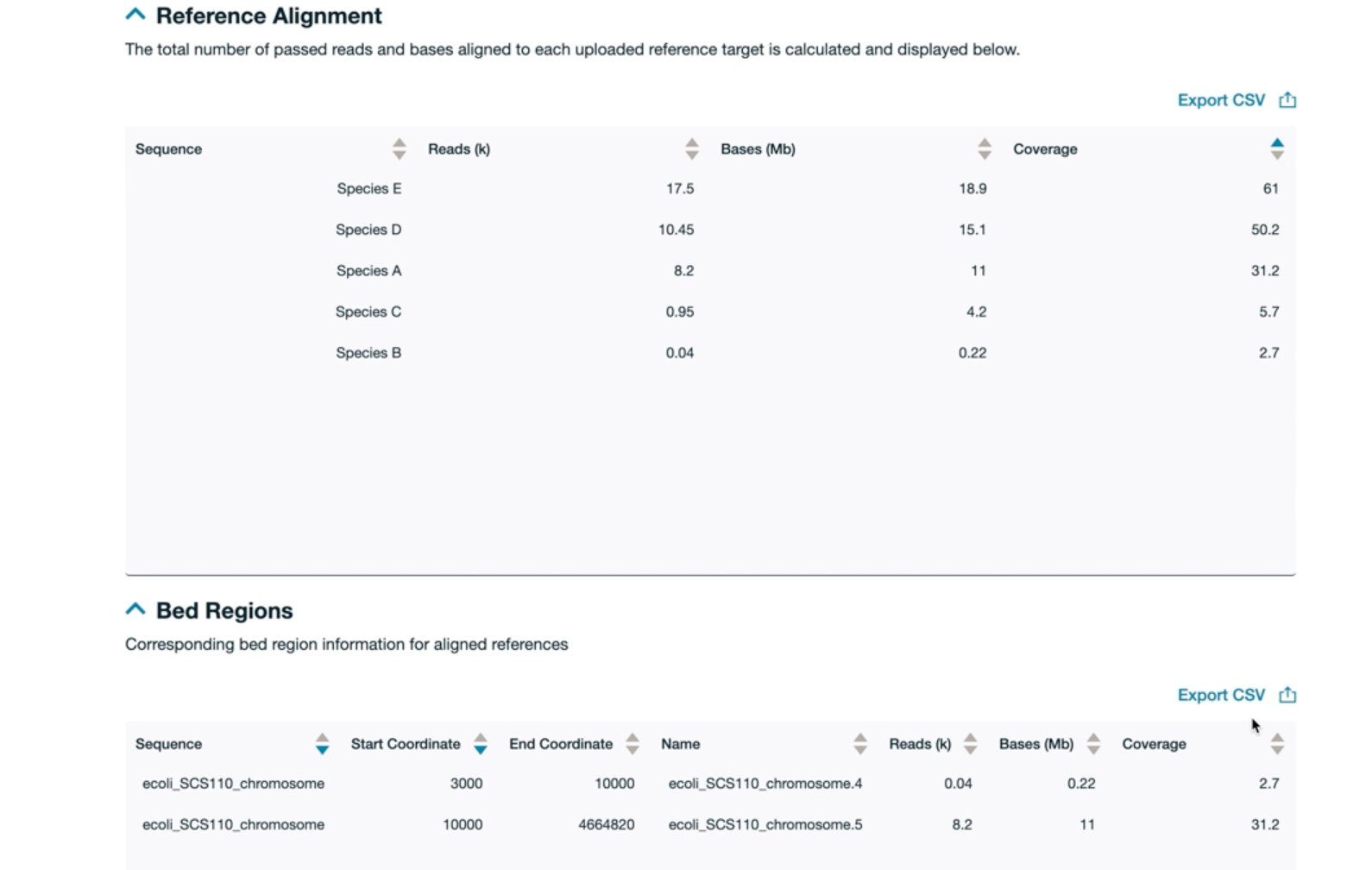
-
Run health
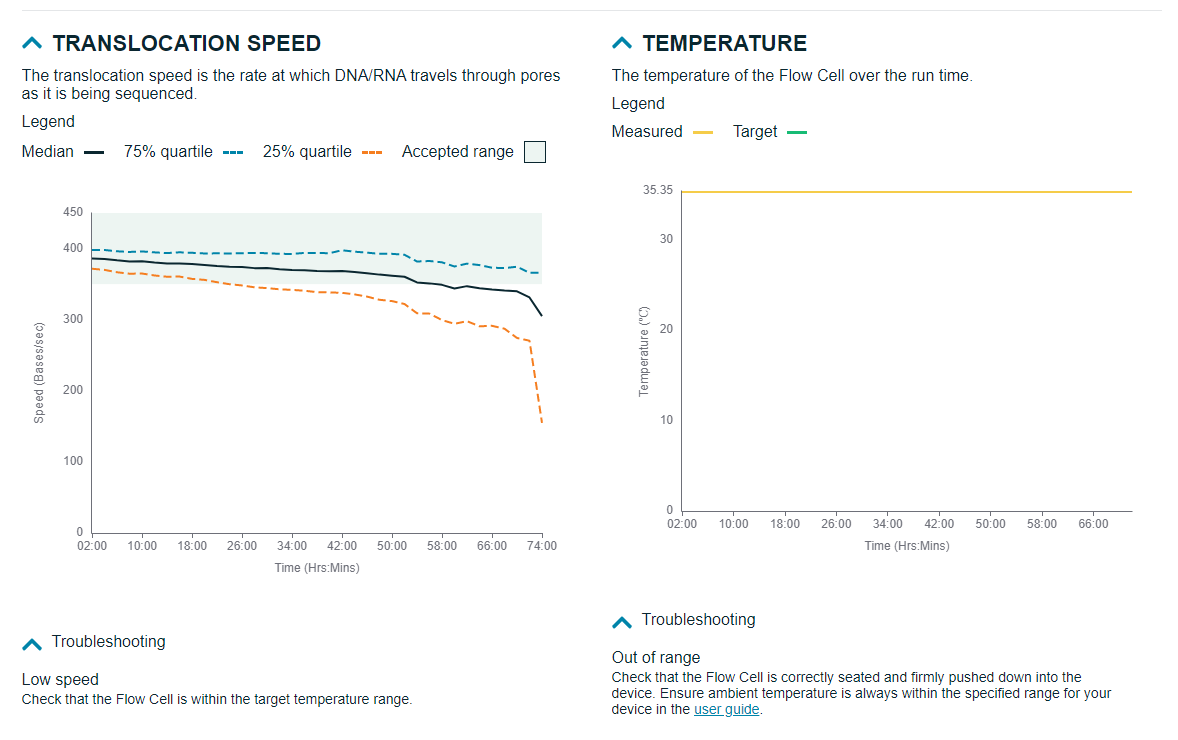
This provides a detailed view of the pore activity and pore scan results throughout the run, along with graphs displaying the translocation speed and temperature.
Pore activity states:
- No pore from scan: No pore detected in channel
- Sequencing: Pore currently sequencing
- Adapter: Pore currently sequencing adapter
- Pore available: Pore available for sequencing
- Unavailable: Pore unavailable for sequencing
- Active feedback: Channel ejecting analyte
- Out of range-high: Current is positive but unavailable for sequencing
- Multiple: Multiple pores detected. Unavailable for sequencing
- Saturated: The channel has switched off as current levels exceed hardware limitations
- Out of range-low: Current is negative but unavailable for sequencing
- Zero: Pore currently unavailable for sequencing
- Channel disabled: Channel is disabled and awaiting another pore scan
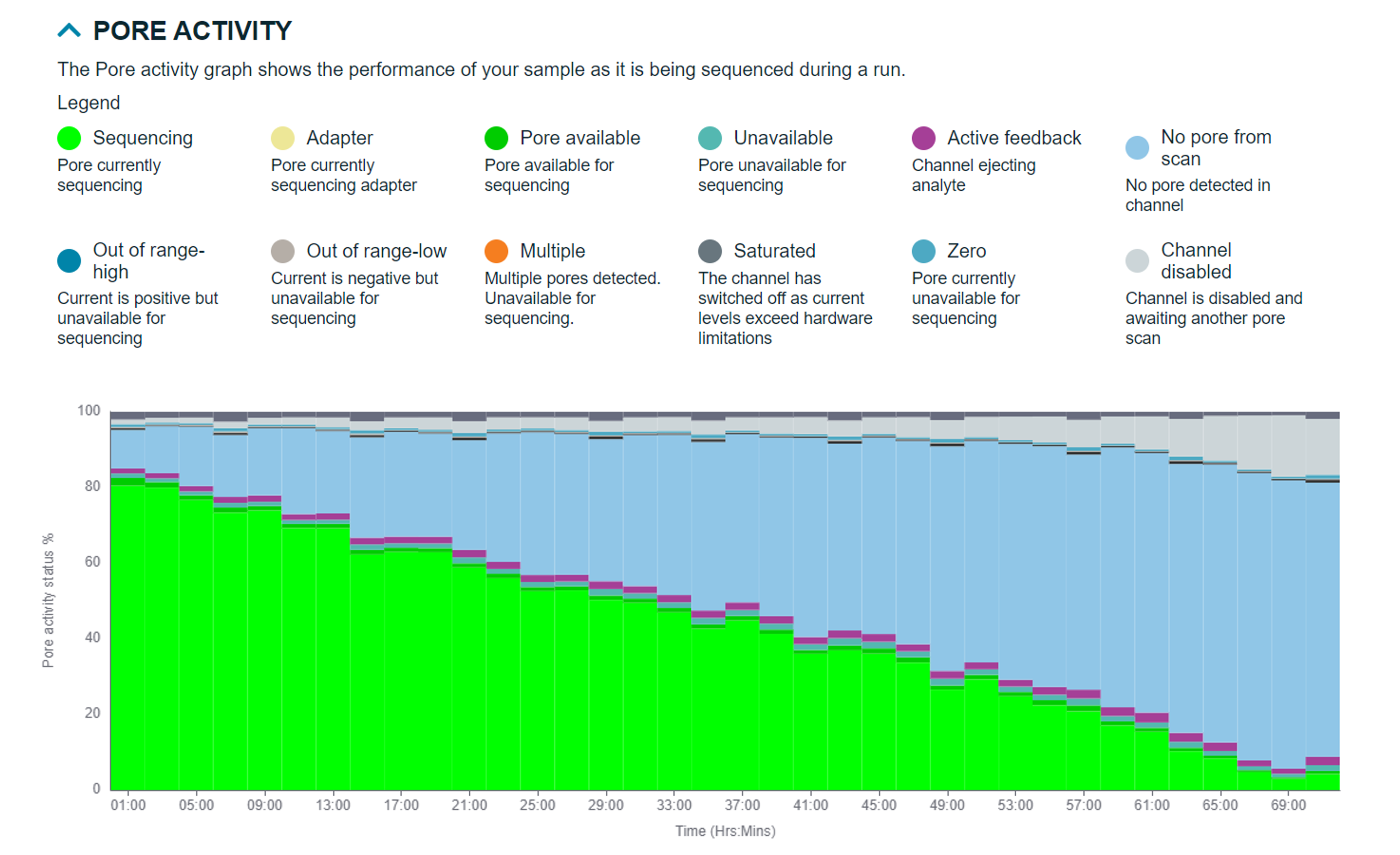
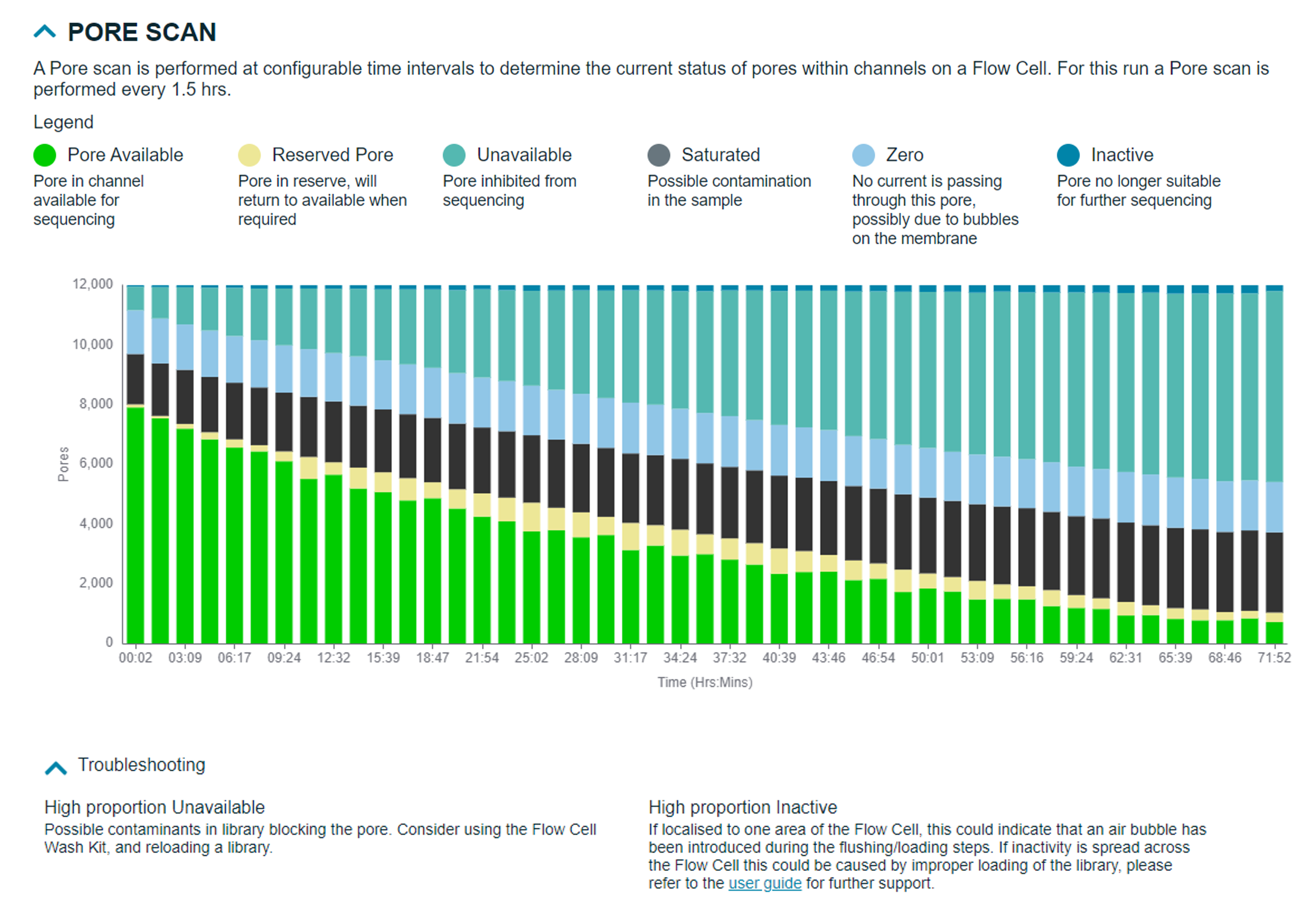
Pore scan states:
- Pore available: Pore in channel available for sequencing
- Reserved pore: Pore in reserve, will return to available when required
- Unavailable: Pore inhibited from sequencing
- Saturated: Possible contamination in the sample
- Zero: No current is passing through this pore, possibly due to bubbles on the membrane
- Inactive: Pore no longer suitable for sequencing
-
Run log
The run log includes system messages sent during the sequencing run regarding errors, warning, and any events.