-
Overview
Oxford Nanopore Technologies' range of sequencing kits is designed to prepare DNA and RNA libraries using 10 ng input material or more. However, users with less than 1 ng can use whole genome amplification to generate more starting material in order to use our sequencing kits. We have previously released a Premium whole genome amplification protocol. This protocol recommends a lengthy amplification step using the REPLI-g Midi Kit (Qiagen, cat # 150043), followed by a T7 endonuclease I digestion and library preparation using the Ligation Sequencing Kit; typically the whole process takes more than a day. We have now developed a much shorter protocol allowing users to go from “cells to sequence” in under two hours. Using the QIAGEN REPLI-g UltraFast Mini Kit, whole genome amplification is performed in ~100 minutes using the REPLI-g UltraFast Mini Kit (Qiagen, cat # 150033) followed by library preparation using the Rapid Sequencing Kit.
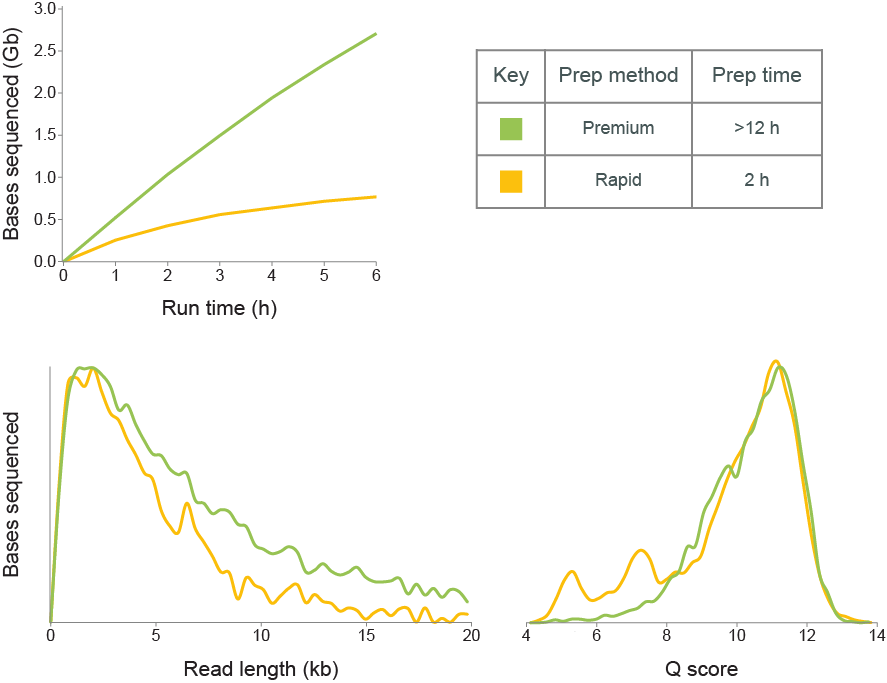
The Rapid method has been designed to generate sequencing libraries from a low number of cells, with minimal sample manipulation and minimal lab equipment, in under 2 hours. However, there is a significant compromise to throughput using this method over the ligation-based library prep, and is generally recommended only for applications where throughput is of low importance.
Although cells can be used in the REPLI-g kit, the chemical lysis step may not be sufficient to release DNA from all cell types. Additional steps to ensure all cells are lysed may be advised, depending on sample type.
-
Input recommendation
The REPLI-g UltraFast Mini Kit suggests an input of either 300 cells‡, or 1-10 ng‡‡ of gDNA. The method describing how to perform whole genome amplification differs, depending on sample type (cells or DNA), and details for each are provided. Please also refer to manufacturer's guides for further details.
Notes:
‡ Although cells can be used in the REPLI-g kit, the chemical lysis step may not be sufficient to release the DNA from all cell types. Additional steps to ensure all cells are lysed may be required, depending on sample type.
‡‡ We have found that it is possible to reduce the input amount to, for example, 10 pg of gDNA. However, lowering the input may require the time of the amplification step to be increased. We recommend amplifying until the DNA concentration is >80 ng/μl (quantification performed using the Qubit dsDNA BR Assay Kit). -
Introduction to the Rapid whole genome amplification protocol
This protocol describes how to carry out whole genome amplification and sequencing of genomic DNA using the Rapid Sequencing Kit (SQK-RAD004). It is highly recommended that a Lambda control experiment is completed first to become familiar with the technology.
Steps in the sequencing workflow:
Prepare for your experiment
You will need to:
- Extract your DNA. Alternatively, start with whole cells
- Ensure you have your sequencing kit, the correct equipment and third-party reagents
- Download the software for acquiring and analysing your data
- Check your flow cell to ensure it has enough pores for a good sequencing runLibrary preparation
You will need to:
- Amplify the genomic DNA from cells using the Qiagen REPLI-g UltraFast Kit
- Tagment your DNA using the Fragmentation Mix in the kit
- Attach sequencing adapters supplied in the kit to the DNA ends
- Prime the flow cell, and load your DNA library into the flow cellSequencing and analysis
You will need to:
- Start a sequencing run using the MinKNOW software, which will collect raw data from the device and convert it into basecalled reads
- Start the EPI2ME software and select a workflow for further analysis (this step is optional)
