- Materials
-
- 10 ng of cDNA amplicons prepared using 10X Genomics Next GEM Single Cell 3' Kits (V3.1)
- Custom ordered-oligo at 10 μM: [Btn]Fwd_3580_partial_read1_defined_for_3'_cDNA (sequence provided in Equipment and Consumables)
- Custom-ordered oligo at 10 μM: Rev_PR2_partial_TSO_defined_for_3'_cDNA (sequence provided in Equipment and Consumables)
- Consumables
-
- LongAmp Hot Start Taq 2X Master Mix (NEB, M0533S)
- Agencourt AMPure XP beads (Beckman Coulter™ cat # A63881)
- Freshly prepared 80% ethanol in nuclease-free water
- Nuclease-free water (e.g. ThermoFisher, cat #AM9937)
- 1.5 ml Eppendorf DNA LoBind tubes
- 0.2 ml thin-walled PCR tubes
- Equipment
-
- Thermal cycler
- Microfuge
- Hula mixer (gentle rotator mixer)
- Magnetic rack (e.g. Invitrogen DynaMag-2 Magnet, Cat # 12321D)
- Ice bucket with ice
- P1000 pipette and tips
- P200 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- P2 pipette and tips
-
Check your flow cell.
We recommend performing a flow cell check before starting your library prep to ensure you have a flow cell with enough pores for a good sequencing run.
See the flow cell check instructions in the MinKNOW protocol for more information.
-
Prepare the cDNA amplicons in nuclease-free water:
- Transfer 10 ng of cDNA amplicons into a 0.2 ml thin-walled PCR tube.
- Adjust the volume to 21 µl with nuclease-free water.
- Mix thoroughly by flicking the tube to avoid unwanted shearing.
- Spin down briefly in a microfuge.
-
In the same 0.2 ml thin-walled PCR tube, set up the following biotin tagging reaction:
Reagent Stock Final Volume cDNA template 0.48 ng/μl 0.2 ng/μl 21 μl [Btn]Fwd_3580_partial_read1_defined_for_3'_cDNA 10 μM 0.4 μM 2 μl Rev_PR2_partial_TSO_defined_for_3'_cDNA 10 μM 0.4 μM 2 μl LongAmp Hot Start Taq 2X Master Mix 2X 1X 25 μl Total - - 50 μl -
Mix by pipetting and spin down.
-
Amplify using the following cycling conditions:
Cycle step Temperature Ramp rate Time No. of cycles Initial denaturation 94°C max 3 min 1 Denaturation
Annealing ramp-down
Annealing
Extension94°C
66°C down to 58°C
58°C
65°Cmax
0.2°C/s
max
max30 sec
40 sec
50 sec
6 mins
4Final extension 65°C max 10 min 1 Hold 4°C - ∞ - Below is a schematic of the cycling conditions.
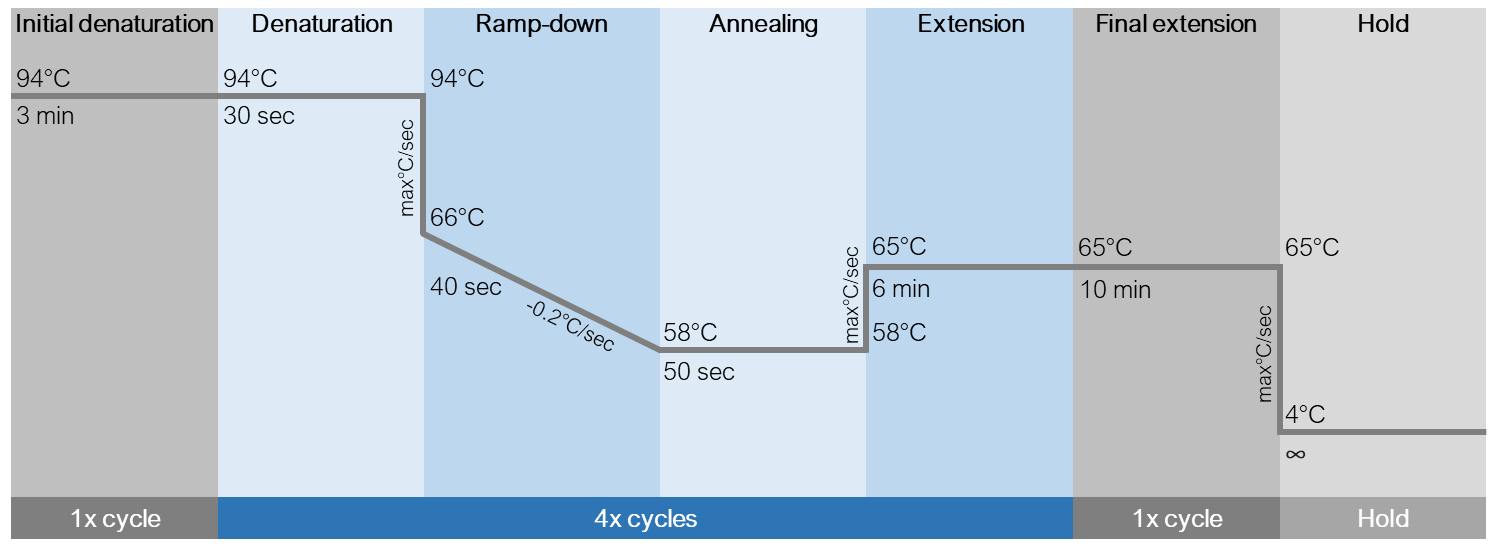
-
Transfer the sample to a clean 1.5 ml Eppendorf DNA LoBind tube.
-
Resuspend the AMPure XP beads by vortexing.
-
Add 40 µl of resuspended AMPure XP beads to the reaction and mix by flicking the tube.
-
Incubate on a Hula mixer (rotator mixer) for 5 minutes at room temperature.
-
Prepare 500 μl of fresh 80% ethanol in nuclease-free water.
-
Spin down the samples and pellet the beads on a magnet until the eluate is clear and colourless. Keep the tubes on the magnet and pipette off the supernatant.
-
Keep the tube on the magnet and wash the beads with 200 µl of freshly prepared 80% ethanol in nuclease-free water without disturbing the pellet. Remove the ethanol using a pipette and discard.
-
Repeat the previous step.
-
Briefly spin down and place the tubes back on the magnet. Pipette off any residual ethanol. Allow to dry for 30 seconds, but do not dry the pellet to the point of cracking.
-
Remove the tube from the magnetic rack and resuspend the pellet in 10 µl nuclease-free water. Spin down and incubate for 2 minutes at room temperature.
-
Pellet the beads on a magnet until the eluate is clear and colourless.
-
Remove and retain 10 µl of eluate into a clean 1.5 ml Eppendorf DNA LoBind tube.
