Welcome to the Nanopore Community
Order MinION devices and consumables
Visit vwr.comLigation sequencing influenza whole genome (SQK-LSK109 with EXP-NBD196)
Version for device: MinION
Introduction to the protocol
Overview of the protocol
Overview of the protocol
-
Before starting
This protocol outlines how to carry out PCR amplification and native barcoding of influenza amplicons on a 96-well plate using the Native Barcoding Expansion 96 (EXP-NBD196) in conjunction with Ligation Sequencing Kit (SQK-LSK109).
When processing multiple samples at once, we recommend making master mixes with an additional 10% of the volume. We also recommend using a template-free pre-PCR hood for making up the master mixes, and a separate template pre-PCR hood for handling the samples. It is important to clean and/or UV irradiate these hoods between sample batches. Furthermore, to track and monitor cross-contamination events, it is important to run a negative control reaction at the reverse transcription stage using nuclease-free water instead of sample, and carrying this control through the rest of the prep.
All post-PCR procedures must be carried out in a separate area to the pre-PCR preparation, with dedicated equipment for liquid handling in each area.
-
Introduction to the influenza whole genome sequencing protocol
To enable support for the rapidly expanding user requests, the team at Oxford Nanopore Technologies have put together a workflow describing how to carry out PCR amplification and native barcoding of influenza amplicons using the Native Barcoding Expansion 96 (EXP-NBD196) in conjunction with Ligation Sequencing Kit (SQK-LSK109). There are 96 unique barcodes available, allowing the user to pool up to 96 different samples in one sequencing experiment.
While this protocol is available in the Nanopore Community, we kindly ask users to ensure they are citing the following references, that this protocol is based on.
Single-reaction genomic amplification accelerates sequencing and vaccine production for classical and Swine origin human influenza A viruses by Bin Zhou et al., 2009. and Universal influenza B virus genomic amplification facilitates sequencing, diagnostics, and reverse genetics by Bin Zhou et al., 2014.
Steps in the sequencing workflow:
Prepare for your experiment
You will need to:
- Ensure you have the sequencing kit, the correct equipment and third-party reagents
- Download the software for acquiring and analysing your data
- Check your flow cell to ensure it has enough pores for a good sequencing run
Prepare your library
You will need to:
- Perform RT-PCR amplification on influenza samples
- Prepare the DNA ends for adapter attachment
- Ligate native barcodes supplied in the kit to the DNA ends
- Ligate sequencing adapters supplied in the kit to the DNA ends
- Prime the flow cell, and load your DNA library into the flow cell
Sequencing
You will need to:
- Start a sequencing run using the MinKNOW software, which will collect raw data from the device and convert it into basecalled reads
- Demultiplex barcoded reads in MinKNOW or Guppy basecalling, choosing the EXP-NBD196 kit option
- Use the wf-flu workflow in EPI2ME Labs to analyse your data
Equipment and consumables
- Materials
-
- Input influenza RNA
- Influenza A primers
- Influenza B primers
- Ligation Sequencing Kit (SQK-LSK109)
- Native Barcoding Expansion 96 (EXP-NBD196)
- Or Native Barcoding Expansion 1-12 (EXP-NBD104) and 13-24 (EXP-NBD114) if multiplexing 1-24 samples.
- Flow Cell Priming Kit (EXP-FLP002)
- Adapter Mix II Expansion (EXP-AMII001)
- SFB Expansion (EXP-SFB001)
- Sequencing Auxiliary Vials (EXP-AUX001)
- Consumables
-
- SuperScript III One-Step RT-PCR System with Platinum Taq DNA Polymerase (ThermoFisher, cat # 12574018 or 12574026)
- Nuclease-free water (e.g. ThermoFisher, AM9937)
- Agencourt AMPure XP beads (Beckman Coulter, A63881)
- Freshly prepared 80% ethanol in nuclease-free water
- NEB Blunt/TA Ligase Master Mix (NEB, M0367)
- NEBNext Ultra II End repair/dA-tailing Module (NEB, E7546)
- NEBNext Quick Ligation Module (NEB, E6056)
- 1.5 ml Eppendorf DNA LoBind tubes
- 5 ml Eppendorf DNA LoBind tubes
- 15 ml Eppendorf DNA LoBind tubes
- Reagent reservoirs for multichannel pipetting
- Qubit dsDNA HS Assay Kit (Invitrogen, Q32851)
- Qubit™ Assay Tubes (Invitrogen, Q32856)
- Eppendorf twin.tec® PCR plate 96 LoBind, semi-skirted (Cat # 0030129504) with heat seals
- Equipment
-
- Magnetic rack, suitable for 1.5 ml Eppendorf tubes
- Magnetic rack suitable for 96 well plates, e.g. DynaMag™-96 Side Skirted Magnet (Thermo Fisher CAT#12027)
- Microfuge
- Vortex mixer
- Thermal cycler
- Hula mixer (gentle rotator mixer)
- Microplate centrifuge, e.g. Fisherbrand™ Mini Plate Spinner Centrifuge (Fisher Scientific, 11766427)
- Qubit fluorometer (or equivalent for QC check)
- Multichannel pipettes suitable for dispensing 0.5–10 μl, 2–20 μl and 20–200 μl, and tips
- P1000 pipette and tips
- P200 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- P10 pipette and tips
- P2 pipette and tips
- Ice bucket with ice
- Timer
- Optional Equipment
-
- PCR hoods with UV steriliser
- PCR-Cooler (Eppendorf)
- Eppendorf 5424 centrifuge (or equivalent)
-
Ligation Sequencing Kit (SQK-LSK109) contents
Name Acronym Cap colour No. of vials Fill volume per vial (µl) DNA CS DCS Yellow 1 50 Adapter Mix AMX Green 1 40 Ligation Buffer LNB Clear 1 200 L Fragment Buffer LFB White cap, orange stripe on label 2 1,800 S Fragment Buffer SFB Grey 2 1,800 Sequencing Buffer SQB Red 2 300 Elution Buffer EB Black 1 200 Loading Beads LB Pink 1 360 -
Flow Cell Priming Kit (EXP-FLP002) contents
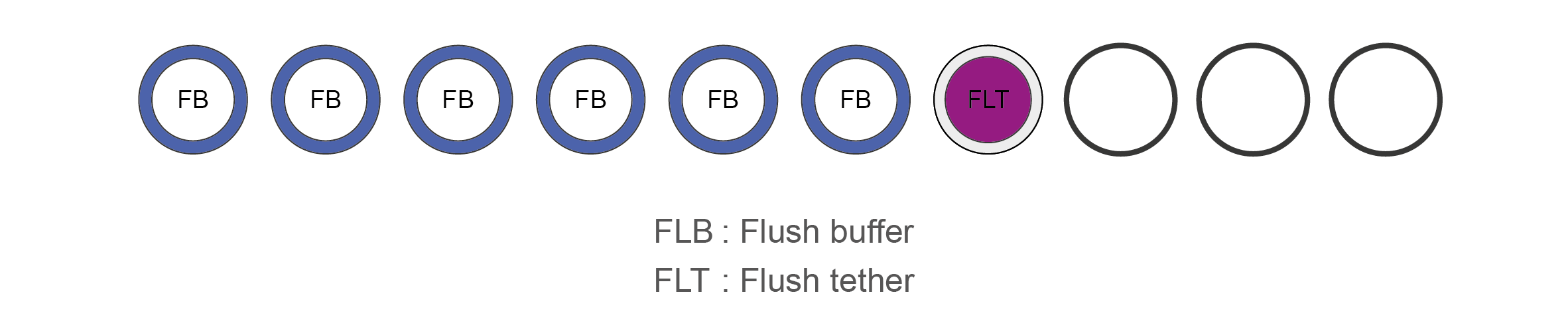
Name Acronym Cap colour No. of vial Fill volume per vial (μl) Flush Buffer FB Blue 6 1,170 Flush Tether FLT Purple 1 200 -
Native Barcoding Expansion 96 (EXP-NBD196) contents
Kits in batches NBD196.10.0007 onwards have barcodes ordered in columns on the plate:
Kits in batches prior to NBD196.10.0007 have barcodes ordered in rows:
Name Acronym Cap colour No. of vials Fill volume per vial (μl) Native Barcode 01-96 NB01-96 - 1 plate 40 μl per well Adapter Mix II AMII Green 1 70 -
SFB Expansion contents (EXP-SFB001)
Name Acronym Cap colour No. of vials Fill volume per vial (μl) Short Fragment Buffer SFB Grey 4 1,800 -
Adapter Mix II Expansion contents (EXP-AMII001)
Name Acronym Cap colour No. of tubes Fill volume per vial (μl) Adapter Mix II AMII Green 2 40 -
Adapter Mix II Expansion use
Protocols that use the Native Barcoding Expansions require 5 μl of AMII per reaction. Native Barcoding Expansions EXP-NBD104/NBD114 contain sufficient AMII for 6 reactions (or 12 reactions when sequencing on Flongle). This assumes that all barcodes are used in one sequencing run.
The Adapter Mix II expansion provides additional AMII for customers who are running subsets of barcodes, and allows a further 12 reactions (24 on Flongle).
-
Sequencing Auxiliary Vials contents (EXP-AUX001)
Name Acronym Cap colour No. of vials Fill volume per vial (μl) Sequencing Buffer SQB Red 6 300 Elution Buffer EB Black 2 200 Loading Beads LB Pink 2 360 -
Native barcode sequences
Component Forward sequence Reverse sequence NB01 CACAAAGACACCGACAACTTTCTT AAGAAAGTTGTCGGTGTCTTTGTG NB02 ACAGACGACTACAAACGGAATCGA TCGATTCCGTTTGTAGTCGTCTGT NB03 CCTGGTAACTGGGACACAAGACTC GAGTCTTGTGTCCCAGTTACCAGG NB04 TAGGGAAACACGATAGAATCCGAA TTCGGATTCTATCGTGTTTCCCTA NB05 AAGGTTACACAAACCCTGGACAAG CTTGTCCAGGGTTTGTGTAACCTT NB06 GACTACTTTCTGCCTTTGCGAGAA TTCTCGCAAAGGCAGAAAGTAGTC NB07 AAGGATTCATTCCCACGGTAACAC GTGTTACCGTGGGAATGAATCCTT NB08 ACGTAACTTGGTTTGTTCCCTGAA TTCAGGGAACAAACCAAGTTACGT NB09 AACCAAGACTCGCTGTGCCTAGTT AACTAGGCACAGCGAGTCTTGGTT NB10 GAGAGGACAAAGGTTTCAACGCTT AAGCGTTGAAACCTTTGTCCTCTC NB11 TCCATTCCCTCCGATAGATGAAAC GTTTCATCTATCGGAGGGAATGGA NB12 TCCGATTCTGCTTCTTTCTACCTG CAGGTAGAAAGAAGCAGAATCGGA NB13 AGAACGACTTCCATACTCGTGTGA TCACACGAGTATGGAAGTCGTTCT NB14 AACGAGTCTCTTGGGACCCATAGA TCTATGGGTCCCAAGAGACTCGTT NB15 AGGTCTACCTCGCTAACACCACTG CAGTGGTGTTAGCGAGGTAGACCT NB16 CGTCAACTGACAGTGGTTCGTACT AGTACGAACCACTGTCAGTTGACG NB17 ACCCTCCAGGAAAGTACCTCTGAT ATCAGAGGTACTTTCCTGGAGGGT NB18 CCAAACCCAACAACCTAGATAGGC GCCTATCTAGGTTGTTGGGTTTGG NB19 GTTCCTCGTGCAGTGTCAAGAGAT ATCTCTTGACACTGCACGAGGAAC NB20 TTGCGTCCTGTTACGAGAACTCAT ATGAGTTCTCGTAACAGGACGCAA NB21 GAGCCTCTCATTGTCCGTTCTCTA TAGAGAACGGACAATGAGAGGCTC NB22 ACCACTGCCATGTATCAAAGTACG CGTACTTTGATACATGGCAGTGGT NB23 CTTACTACCCAGTGAACCTCCTCG CGAGGAGGTTCACTGGGTAGTAAG NB24 GCATAGTTCTGCATGATGGGTTAG CTAACCCATCATGCAGAACTATGC NB25 GTAAGTTGGGTATGCAACGCAATG CATTGCGTTGCATACCCAACTTAC NB26 CATACAGCGACTACGCATTCTCAT ATGAGAATGCGTAGTCGCTGTATG NB27 CGACGGTTAGATTCACCTCTTACA TGTAAGAGGTGAATCTAACCGTCG NB28 TGAAACCTAAGAAGGCACCGTATC GATACGGTGCCTTCTTAGGTTTCA NB29 CTAGACACCTTGGGTTGACAGACC GGTCTGTCAACCCAAGGTGTCTAG NB30 TCAGTGAGGATCTACTTCGACCCA TGGGTCGAAGTAGATCCTCACTGA NB31 TGCGTACAGCAATCAGTTACATTG CAATGTAACTGATTGCTGTACGCA NB32 CCAGTAGAAGTCCGACAACGTCAT ATGACGTTGTCGGACTTCTACTGG NB33 CAGACTTGGTACGGTTGGGTAACT AGTTACCCAACCGTACCAAGTCTG NB34 GGACGAAGAACTCAAGTCAAAGGC GCCTTTGACTTGAGTTCTTCGTCC NB35 CTACTTACGAAGCTGAGGGACTGC GCAGTCCCTCAGCTTCGTAAGTAG NB36 ATGTCCCAGTTAGAGGAGGAAACA TGTTTCCTCCTCTAACTGGGACAT NB37 GCTTGCGATTGATGCTTAGTATCA TGATACTAAGCATCAATCGCAAGC NB38 ACCACAGGAGGACGATACAGAGAA TTCTCTGTATCGTCCTCCTGTGGT NB39 CCACAGTGTCAACTAGAGCCTCTC GAGAGGCTCTAGTTGACACTGTGG NB40 TAGTTTGGATGACCAAGGATAGCC GGCTATCCTTGGTCATCCAAACTA NB41 GGAGTTCGTCCAGAGAAGTACACG CGTGTACTTCTCTGGACGAACTCC NB42 CTACGTGTAAGGCATACCTGCCAG CTGGCAGGTATGCCTTACACGTAG NB43 CTTTCGTTGTTGACTCGACGGTAG CTACCGTCGAGTCAACAACGAAAG NB44 AGTAGAAAGGGTTCCTTCCCACTC GAGTGGGAAGGAACCCTTTCTACT NB45 GATCCAACAGAGATGCCTTCAGTG CACTGAAGGCATCTCTGTTGGATC NB46 GCTGTGTTCCACTTCATTCTCCTG CAGGAGAATGAAGTGGAACACAGC NB47 GTGCAACTTTCCCACAGGTAGTTC GAACTACCTGTGGGAAAGTTGCAC NB48 CATCTGGAACGTGGTACACCTGTA TACAGGTGTACCACGTTCCAGATG NB49 ACTGGTGCAGCTTTGAACATCTAG CTAGATGTTCAAAGCTGCACCAGT NB50 ATGGACTTTGGTAACTTCCTGCGT ACGCAGGAAGTTACCAAAGTCCAT NB51 GTTGAATGAGCCTACTGGGTCCTC GAGGACCCAGTAGGCTCATTCAAC NB52 TGAGAGACAAGATTGTTCGTGGAC GTCCACGAACAATCTTGTCTCTCA NB53 AGATTCAGACCGTCTCATGCAAAG CTTTGCATGAGACGGTCTGAATCT NB54 CAAGAGCTTTGACTAAGGAGCATG CATGCTCCTTAGTCAAAGCTCTTG NB55 TGGAAGATGAGACCCTGATCTACG CGTAGATCAGGGTCTCATCTTCCA NB56 TCACTACTCAACAGGTGGCATGAA TTCATGCCACCTGTTGAGTAGTGA NB57 GCTAGGTCAATCTCCTTCGGAAGT ACTTCCGAAGGAGATTGACCTAGC NB58 CAGGTTACTCCTCCGTGAGTCTGA TCAGACTCACGGAGGAGTAACCTG NB59 TCAATCAAGAAGGGAAAGCAAGGT ACCTTGCTTTCCCTTCTTGATTGA NB60 CATGTTCAACCAAGGCTTCTATGG CCATAGAAGCCTTGGTTGAACATG NB61 AGAGGGTACTATGTGCCTCAGCAC GTGCTGAGGCACATAGTACCCTCT NB62 CACCCACACTTACTTCAGGACGTA TACGTCCTGAAGTAAGTGTGGGTG NB63 TTCTGAAGTTCCTGGGTCTTGAAC GTTCAAGACCCAGGAACTTCAGAA NB64 GACAGACACCGTTCATCGACTTTC GAAAGTCGATGAACGGTGTCTGTC NB65 TTCTCAGTCTTCCTCCAGACAAGG CCTTGTCTGGAGGAAGACTGAGAA NB66 CCGATCCTTGTGGCTTCTAACTTC GAAGTTAGAAGCCACAAGGATCGG NB67 GTTTGTCATACTCGTGTGCTCACC GGTGAGCACACGAGTATGACAAAC NB68 GAATCTAAGCAAACACGAAGGTGG CCACCTTCGTGTTTGCTTAGATTC NB69 TACAGTCCGAGCCTCATGTGATCT AGATCACATGAGGCTCGGACTGTA NB70 ACCGAGATCCTACGAATGGAGTGT ACACTCCATTCGTAGGATCTCGGT NB71 CCTGGGAGCATCAGGTAGTAACAG CTGTTACTACCTGATGCTCCCAGG NB72 TAGCTGACTGTCTTCCATACCGAC GTCGGTATGGAAGACAGTCAGCTA NB73 AAGAAACAGGATGACAGAACCCTC GAGGGTTCTGTCATCCTGTTTCTT NB74 TACAAGCATCCCAACACTTCCACT AGTGGAAGTGTTGGGATGCTTGTA NB75 GACCATTGTGATGAACCCTGTTGT ACAACAGGGTTCATCACAATGGTC NB76 ATGCTTGTTACATCAACCCTGGAC GTCCAGGGTTGATGTAACAAGCAT NB77 CGACCTGTTTCTCAGGGATACAAC GTTGTATCCCTGAGAAACAGGTCG NB78 AACAACCGAACCTTTGAATCAGAA TTCTGATTCAAAGGTTCGGTTGTT NB79 TCTCGGAGATAGTTCTCACTGCTG CAGCAGTGAGAACTATCTCCGAGA NB80 CGGATGAACATAGGATAGCGATTC GAATCGCTATCCTATGTTCATCCG NB81 CCTCATCTTGTGAAGTTGTTTCGG CCGAAACAACTTCACAAGATGAGG NB82 ACGGTATGTCGAGTTCCAGGACTA TAGTCCTGGAACTCGACATACCGT NB83 TGGCTTGATCTAGGTAAGGTCGAA TTCGACCTTACCTAGATCAAGCCA NB84 GTAGTGGACCTAGAACCTGTGCCA TGGCACAGGTTCTAGGTCCACTAC NB85 AACGGAGGAGTTAGTTGGATGATC GATCATCCAACTAACTCCTCCGTT NB86 AGGTGATCCCAACAAGCGTAAGTA TACTTACGCTTGTTGGGATCACCT NB87 TACATGCTCCTGTTGTTAGGGAGG CCTCCCTAACAACAGGAGCATGTA NB88 TCTTCTACTACCGATCCGAAGCAG CTGCTTCGGATCGGTAGTAGAAGA NB89 ACAGCATCAATGTTTGGCTAGTTG CAACTAGCCAAACATTGATGCTGT NB90 GATGTAGAGGGTACGGTTTGAGGC GCCTCAAACCGTACCCTCTACATC NB91 GGCTCCATAGGAACTCACGCTACT AGTAGCGTGAGTTCCTATGGAGCC NB92 TTGTGAGTGGAAAGATACAGGACC GGTCCTGTATCTTTCCACTCACAA NB93 AGTTTCCATCACTTCAGACTTGGG CCCAAGTCTGAAGTGATGGAAACT NB94 GATTGTCCTCAAACTGCCACCTAC GTAGGTGGCAGTTTGAGGACAATC NB95 CCTGTCTGGAAGAAGAATGGACTT AAGTCCATTCTTCTTCCAGACAGG NB96 CTGAACGGTCATAGAGTCCACCAT ATGGTGGACTCTATGACCGTTCAG -
Influenza primer sequences
Influenza A primer sequences described in the protocol originated from: Single-reaction genomic amplification accelerates sequencing and vaccine production for classical and Swine origin human influenza A viruses by Bin Zhou et al., 2009.
Component Sequence Tuni 12 ACGCGTGATCAGCAAAAGCAGG Tuni 12.4 ACGCGTGATCAGCGAAAGCAGG Tuni 13 ACGCGTGATCAGTAGAAACAAGG Influenza B primer sequences described in the protocol originated from: Universal influenza B virus genomic amplification facilitates sequencing, diagnostics, and reverse genetics by Bin Zhou et al., 2014.
Component Sequence B-PBs-UniF GGGGGGAGCAGAAGCGGAGC B-PBs-UniR CCGGGTTATTAGTAGAAACACGAGC B-PA-UniF GGGGGGAGCAGAAGCGGTGC B-PA-UniR CCGGGTTATTAGTAGAAACACGTGC B-HANA-UniF GGGGGGAGCAGAAGCAGAGC B-HANA-UniR CCGGGTTATTAGTAGTAACAAGAGC B-NP-UniF GGGGGGAGCAGAAGCACAGC B-NP-UniR CCGGGTTATTAGTAGAAACAACAGC B-M-Uni3F GGGGGGAGCAGAAGCACGCACTT B-Mg-Uni3F GGGGGGAGCAGAAGCAGGCACTT B-M-Uni3R CCGGGTTATTAGTAGAAACAACGCACTT B-NS-Uni3F GGGGGGAGCAGAAGCAGAGGATT B-NS-Uni3R CCGGGTTATTAGTAGTAACAAGAGGATT
Computer requirements and software
Computer requirements and software
-
MinION Mk1B IT requirements
Sequencing on a MinION Mk1B requires a high-spec computer or laptop to keep up with the rate of data acquisition. For more information, refer to the MinION Mk1B IT requirements document.
-
MinION Mk1C IT requirements
The MinION Mk1C contains fully-integrated compute and screen, removing the need for any accessories to generate and analyse nanopore data. For more information refer to the MinION Mk1C IT requirements document.
-
MinION Mk1D IT requirements
Sequencing on a MinION Mk1D requires a high-spec computer or laptop to keep up with the rate of data acquisition. For more information, refer to the MinION Mk1D IT requirements document.
-
Software for nanopore sequencing
MinKNOW
The MinKNOW software controls the nanopore sequencing device, collects sequencing data and basecalls in real time. You will be using MinKNOW for every sequencing experiment to sequence, basecall and demultiplex if your samples were barcoded.
For instructions on how to run the MinKNOW software, please refer to the MinKNOW protocol.
EPI2ME (optional)
The EPI2ME cloud-based platform performs further analysis of basecalled data, for example alignment to the Lambda genome, barcoding, or taxonomic classification. You will use the EPI2ME platform only if you would like further analysis of your data post-basecalling.
For instructions on how to create an EPI2ME account and install the EPI2ME Desktop Agent, please refer to this link.
-
Check your flow cell
We highly recommend that you check the number of pores in your flow cell prior to starting a sequencing experiment. This should be done within 12 weeks of purchasing for MinION/GridION/PromethION or within four weeks of purchasing Flongle Flow Cells. Oxford Nanopore Technologies will replace any flow cell with fewer than the number of pores in the table below, when the result is reported within two days of performing the flow cell check, and when the storage recommendations have been followed. To do the flow cell check, please follow the instructions in the Flow Cell Check document.
Flow cell Minimum number of active pores covered by warranty Flongle Flow Cell 50 MinION/GridION Flow Cell 800 PromethION Flow Cell 5000
Library preparation
Reverse transcription, PCR and clean-up
Reverse transcription, PCR and clean-up
- Materials
-
- Input influenza RNA
- Influenza A primers
- Influenza B primers
- Consumables
-
- SuperScript III One-Step RT-PCR System with Platinum Taq DNA Polymerase (ThermoFisher, cat # 12574018 or 12574026)
- Nuclease-free water (e.g. ThermoFisher, cat # AM9937)
- Eppendorf twin.tec® PCR plate 96 LoBind, semi-skirted (Cat # 0030129504) with heat seals
- 1.5 ml Eppendorf DNA LoBind tubes
- 5 ml Eppendorf DNA LoBind tubes
- 15 ml Eppendorf DNA LoBind tubes
- Agencourt AMPure XP beads (Beckman Coulter, A63881)
- Freshly prepared 80% ethanol in nuclease-free water
- Qubit dsDNA HS Assay Kit (Invitrogen, Q32851)
- Qubit™ Assay Tubes (Invitrogen, Q32856)
- Equipment
-
- Thermal cycler
- Qubit fluorometer (or equivalent for QC check)
- Multichannel pipettes suitable for dispensing 20–200 μl, and tips
- P1000 pipette and tips
- P200 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- P10 pipette and tips
- P2 pipette and tips
- Magnetic rack suitable for 96 well plates, e.g. DynaMag™-96 Side Skirted Magnet (Thermo Fisher CAT#12027)
- Microplate centrifuge, e.g. Fisherbrand™ Mini Plate Spinner Centrifuge (Fisher Scientific, 11766427)
- Microfuge
- Optional Equipment
-
- PCR hoods with UV steriliser
- PCR-Cooler (Eppendorf)
-
To reduce risk of contamination, we recommend the use of PCR hoods with a UV steriliser when setting up the PCR plates.
- When handling the primer stocks and derivatives, use a clean template-free PCR hood.
- When handling the samples and/or a positive control, use a clean template-addition PCR hood.
-
In a clean template-free pre-PCR hood, prepare the primer mixes for influenza A and influenza B as follows in 1.5 ml Eppendorf DNA LoBind tubes:
Note: The volume requirements can be adjusted according to stock concentrations and experiment needs.
Influenza A primer mix
Primer Concentration Volume Nuclease-free water - 378 µl Tuni 12 100 µM 16.8 µl Tuni 12.4 100 µM 4.2 µl Tuni 13 100 µM 21 µl Total 420 µl Influenza B primer mix
Primer Concentration Volume Nuclease-free water - 378 µl B-PBs-UniF 100 µM 5 µl B-PBs-UniR 100 µM 5 µl B-PA-UniF 100 µM 2.5 µl B-PA-UniR 100 µM 2.5 µl B-HANA-UniF 100 µM 5 µl B-HANA-UniR 100 µM 5 µl B-NP-UniF 100 µM 3 µl B-NP-UniR 100 µM 3 µl B-M-Uni3F 100 µM 1.5 µl B-Mg-Uni3F 100 µM 1.5 µl B-M-Uni3R 100 µM 3 µl B-NS-Uni3F 100 µM 2.5 µl B-NS-Uni3R 100 µM 2.5 µl Total 420 µl -
In the template-free pre-PCR hood, prepare the following master mixes in Eppendorf DNA LoBind tubes and mix thoroughly as follows:
For X12 samples, use 1.5 ml Eppendorf DNA LoBind tubes:
Component Influenza A RT-PCR Master Mix Influenza B RT-PCR Master Mix Nuclease free water 280 µl 280 µl Influenza A primer mix 28 µl - Influenza B primer mix - 28 µl 2X Reaction Mix 350 µl 350 µl SuperScript™ III RT/Platinum™ Taq Mix 28 µl 28 µl Total volume 686 µl 686 µl For X24 samples, use 1.5 ml Eppendorf DNA LoBind tubes:
Component Influenza A RT-PCR Master Mix Influenza B RT-PCR Master Mix Nuclease free water 560 µl 560 µl Influenza A primer mix 56 µl - Influenza B primer mix - 56 µl 2X Reaction Mix 700 µl 700 µl SuperScript™ III RT/Platinum™ Taq Mix 56 µl 56 µl Total volume 1372 µl 1372 µl For X48 samples, use 5 ml Eppendorf DNA LoBind tubes:
Component Influenza A RT-PCR Master Mix Influenza B RT-PCR Master Mix Nuclease free water 1120 µl 1120 µl Influenza A primer mix 112 µl - Influenza B primer mix - 112 µl 2X Reaction Mix 1400 µl 1400 µl SuperScript™ III RT/Platinum™ Taq Mix 112 µl 112 µl Total volume 2744 µl 2744 µl For X96 samples, use 15 ml Eppendorf DNA LoBind tubes:
Component Influenza A RT-PCR Master Mix Influenza B RT-PCR Master Mix Nuclease free water 2240 µl 2240 µl Influenza A primer mix 224 µl - Influenza B primer mix - 224 µl 2X Reaction Mix 2800 µl 2800 µl SuperScript™ III RT/Platinum™ Taq Mix 224 µl 224 µl Total volume 5488 µl 5488 µl -
For each influenza type, place one clean 96-well RT-PCR plate into a PCR-cooler (if using).
-
Using a stepper pipette or a multichannel pipette, aliquot 49 µl of influenza A RT-PCR Master Mix into the influenza A RT-PCR plate.
-
Using a stepper pipette or a multichannel pipette, aliquot 49 µl of influenza B RT-PCR Master Mix into the influenza B RT-PCR plate.
-
Seal the RT-PCR plate(s) and transfer to a template-addition pre-PCR hood.
-
Transfer 1 µl of influenza A samples to the wells containing influenza A RT-PCR Master Mix in the influenza A RT-PCR plate and mix thoroughly by pipetting the contents of each well up and down.
-
Transfer 1 µl of influenza B samples to the wells containing influenza B RT-PCR Master Mix in the influenza B RT-PCR plate and mix thoroughly by pipetting the contents of each well up and down.
-
Seal the RT-PCR plate(s) and spin down in a centrifuge.
-
Incubate the influenza A RT-PCR plate using the following program, with the heated lid set to 105°C:
Step Temperature Time Cycles cDNA synthesis 42°C 60 min 1 Initial denaturation 94°C 2 min 1 Denaturation
Annealing and extension94°C
45°C
68°C30 sec
30 sec
3 min
5Denaturation
Annealing and extension94°C
57°C
68°C30 sec
30 sec
3 min
31Hold 4°C ∞ -
Incubate the influenza B RT-PCR plate using the following program, with the heated lid set to 105°C:
Step Temperature Time Cycles cDNA synthesis 45°C 60 min 1 cDNA synthesis 55°C 30 min 1 Initial denaturation 94°C 2 min 1 Denaturation
Annealing and extension94°C
40°C
68°C20 sec
30 sec
3 min 30 sec
5Denaturation
Annealing and extension94°C
58°C
68°C20 sec
30 sec
3 min 30 sec
30Final extension 68°C 10 min 1 Hold 4°C ∞ -
Optional ActionIf necessary, the protocol can be paused at this point. The samples should be kept at 4°C and can be stored overnight.
-
Resuspend the AMPure XP beads by vortexing.
-
Add 50 µl of resuspended AMPure XP beads to each well of the RT-PCR plate(s) and mix by gently pipetting.
-
Incubate the RT-PCR plate(s) at room temperature for 10 minutes.
-
Prepare at least 500 µl 80% ethanol in nuclease-free water per sample.
-
Spin down the RT-PCR plate(s) and pellet the beads on a magnet for 5 minutes. Keep the plate on the magnet until the eluate is clear and colourless, and pipette off the supernatant.
-
Keep the plate on the magnet and wash the beads in each well with 200 µl of freshly prepared 80% ethanol without disturbing the pellet. Remove the ethanol using a pipette and discard.
-
Repeat the previous step.
-
Spin down and place the plate back on the magnet. Pipette off any residual ethanol. Allow to dry for ~30 seconds, but do not dry the pellet to the point of cracking.
-
Remove the plate from the magnetic rack and resuspend each pellet in 15 µl nuclease-free water. Incubate for 2 minutes at room temperature.
-
Pellet the beads on a magnet until the eluate is clear and colourless.
-
Remove and retain 15 µl of eluate containing the DNA per well, into a clean 96-well plate(s).
Dispose of the pelleted beads.
-
Quantify 1 µl of each eluted sample using a Qubit fluorometer.
End-prep
End-prep
- Consumables
-
- Nuclease-free water (e.g. ThermoFisher, cat # AM9937)
- NEBNext® Ultra™ II End Repair/dA-Tailing Module (E7546)
- 1.5 ml Eppendorf DNA LoBind tubes
- Eppendorf twin.tec® PCR plate 96 LoBind, semi-skirted (Cat # 0030129504) with heat seals
- Equipment
-
- Multichannel pipette capable of dispensing 0.5 – 10 µL, and tips
- P1000 pipette and tips
- P200 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- P10 pipette and tips
- Thermal cycler
- Microfuge
- Ice bucket with ice
- Microplate centrifuge, e.g. Fisherbrand™ Mini Plate Spinner Centrifuge (Fisher Scientific, 11766427)
- Vortex mixer
-
Determine the volume of the cleaned-up PCR reaction that yields 200 fmol of DNA per sample and aliquot into a clean 96-well plate (End-prep plate).
-
Prepare the NEBNext Ultra II End Repair / dA-tailing Module reagents in accordance with manufacturer's instructions, and place on ice:
For optimal performance, NEB recommend the following:
Thaw all reagents on ice.
Ensure the reagents are well mixed.
Note: Do not vortex the Ultra II End Prep Enzyme Mix.Always spin down tubes before opening for the first time each day.
The NEBNext Ultra II End Prep Reaction Buffer may contain a white precipitate. If this occurs, allow the mixture(s) to come to room temperature and pipette the buffer several times to break up the precipitate, followed by a quick vortex to mix.
-
Make up each sample per well to 12.5 µl using nuclease-free water.
-
Prepare the following end-prep master mix in 1.5 ml Eppendorf DNA LoBind tube and mix thoroughly by pipetting:
Reagent Volume per reaction For X24 samples For X48 samples For X96 samples Ultra II End-prep reaction buffer 1.75 µl 52.5 µl 105 µl 210 µl Ultra II End-prep enzyme mix 0.75 µl 22.5 µl 45 µl 90 µl Total 2.5 µl 75 µl 150 µl 300 µl -
Using a stepper pipette or multi-channel pipette, add 2.5 µl of the end-prep master mix to each well containing 12.5 µl sample.
-
Ensure the reactions are thoroughly mixed by pipetting. Seal the End-prep plate and spin down briefly.
-
Using a thermal cycler, incubate the plate at 20°C for 5 minutes and 65°C for 5 minutes.
Native barcode ligation
Native barcode ligation
- Materials
-
- Native Barcoding Expansion 96 (EXP-NBD196)
- Short Fragment Buffer (SFB)
- Consumables
-
- 1.5 ml Eppendorf DNA LoBind tubes
- Eppendorf twin.tec® PCR plate 96 LoBind, semi-skirted (Eppendorf™, cat # 0030129504) with heat seals
- Nuclease-free water (e.g. ThermoFisher, cat # AM9937)
- Freshly prepared 80% ethanol in nuclease-free water
- Agencourt AMPure XP Beads (Beckman Coulter™, A63881)
- NEB Blunt/TA Ligase Master Mix (M0367)
- Equipment
-
- Magnetic rack, suitable for 1.5 ml Eppendorf tubes
- Thermal cycler
- Hula mixer (gentle rotator mixer)
- Vortex mixer
- Ice bucket with ice
- Microplate centrifuge, e.g. Fisherbrand™ Mini Plate Spinner Centrifuge (Fisher Scientific, 11766427)
- Microfuge
- P1000 pipette and tips
- P200 pipette and tips
- P100 pipette and tips
- P10 pipette and tips
- P2 pipette and tips
- Optional Equipment
-
- Qubit fluorometer (or equivalent for QC check)
-
Thaw Native Barcodes at room temperature. Use one barcode per sample.
- Spin down barcodes before opening tubes/piercing plates.
- Pipette mix the entire content of each barcode 10 times before use.
- Once thawed, keep the barcodes on ice.
-
Prepare the NEB Blunt/TA Ligase Master Mix according to the manufacturer's instructions, and place on ice:
Thaw the reagents at room temperature.
Spin down the reagent tubes for 5 seconds.
Ensure the reagents are fully mixed by performing 10 full volume pipette mixes.
-
Thaw the Short Fragment Buffer (SFB) at room temperature and mix by vortexing. Then spin down and place on ice.
-
In a clean 96-well plate (Native barcode ligation plate), add the following reagents in the following order per well:
Between each addition, pipette mix 10-20 times.
Reagent Volume Nuclease-free water 3 µl End-prepped DNA 0.75 µl Native Barcode 1.25 µl Blunt/TA Ligase Master Mix 5 µl Total 10 µl -
Mix contents thoroughly by pipetting, seal the Native barcode ligation plate and spin down briefly.
-
Using a thermal cycler, incubate the Native barcode ligation plate at 20°C for 20 mins and at 65°C for 10 mins.
-
Pool all the barcoded samples in a clean 1.5 ml Eppendorf DNA LoBind tube, checking the base of the plate to ensure all the liquid has been pooled.
Recovery aim ~ 10 µl per sample:
24 samples 48 samples 96 samples Total volume ~ 240 µl ~ 480 µl ~ 960 µl -
Resuspend the AMPure XP beads by vortexing.
-
Add a 0.4X volume of AMPure XP beads to the pooled reaction and mix thoroughly by pipetting:
Volume per sample For X24 samples For X48 samples For X96 samples Volume of AMPure XP beads 4 µl 96 µl 192 µl 384 µl -
Incubate on a Hula mixer (rotator mixer) for 10 minutes at room temperature.
-
Prepare 500 µl of fresh 80% ethanol in nuclease-free water.
-
Spin down the sample and pellet the beads on a magnet for 5 minutes. Keep the tube on the magnet until the eluate is clear, and pipette off the supernatant.
-
Wash the beads by adding 700 μl Short Fragment Buffer (SFB). Flick the beads to resuspend, then return the tube to the magnetic rack and allow the beads to pellet. Keep the tube on the magnet until the eluate is clear and colourless. Remove the supernatant using a pipette and discard.
-
Repeat the previous step.
-
Keep the tube on the magnetic rack and wash the beads with 100 µl of freshly prepared 80% ethanol without disturbing the pellet. Remove the ethanol using a pipette and discard.
-
Spin down and place the tube back on the magnetic rack. Pipette off any residual ethanol. Allow the pellet to dry for ~30 seconds, but do not dry the pellet to the point of cracking.
-
Remove the tube from the magnetic rack and resuspend the pellet in 35 µl nuclease-free water. Incubate for 2 minutes at room temperature.
-
Pellet the beads on a magnetic rack until the eluate is clear and colourless.
-
Remove and retain 35 µl of eluate into a clean 1.5 ml Eppendorf DNA LoBind tube.
Dispose of the pelleted beads
-
Quantify 1 µl of eluted sample using a Qubit fluorometer.
Adapter ligation and clean-up
Adapter ligation and clean-up
- Materials
-
- Elution Buffer (EB)
- Short Fragment Buffer (SFB)
- Adapter Mix II (AMII)
- Consumables
-
- NEBNext Quick Ligation Module (NEB, E6056)
- Agencourt AMPure XP beads (Beckman Coulter, A63881)
- 1.5 ml Eppendorf DNA LoBind tubes
- Equipment
-
- Microfuge
- Magnetic rack, suitable for 1.5 ml Eppendorf tubes
- P200 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- P10 pipette and tips
- Vortex mixer
- Hula mixer (gentle rotator mixer)
- Ice bucket with ice
- Optional Equipment
-
- Qubit fluorometer (or equivalent for QC check)
-
Adapter Mix II Expansion use
Protocols that use the Native Barcoding Expansions require 5 μl of AMII per reaction. Native Barcoding Expansions EXP-NBD104/NBD114 and EXP-NBD196 contain sufficient AMII for 6 and 12 reactions, respectively (or 12 and 24 reactions when sequencing on Flongle). This assumes that all barcodes are used in one sequencing run.
The Adapter Mix II expansion provides additional AMII for customers who are running subsets of barcodes, and allows a further 12 reactions (24 on Flongle).
-
Prepare the NEBNext Quick Ligation Reaction Module according to the manufacturer's instructions, and place on ice:
Thaw the reagents at room temperature.
Spin down the reagent tubes for 5 seconds.
Ensure the reagents are fully mixed by performing 10 full volume pipette mixes.
Note: Do NOT vortex the Quick T4 DNA Ligase.
The NEBNext Quick Ligation Reaction Buffer (5x) may have a little precipitate. Allow the mixture to come to room temperature and pipette the buffer up and down several times to break up the precipitate, followed by vortexing the tube for several seconds to ensure the reagent is thoroughly mixed.
-
Thaw the Elution Buffer (EB) and the Short Fragment Buffer (SFB) at room temperature, before mixing by vortexing. Then spin down and place on ice.
-
Spin down the Adapter Mix II (AMII), pipette mix and place on ice.
-
Perform the adapter ligation of the pooled and barcoded DNA. In a clean 1.5 ml Eppendorf LoBind tube, add the reagents in the following order:
Between each addition, pipette mix 10-20 times.
Reagent Volume Pooled barcoded sample 30 µl Adapter Mix II (AMII) 5 µl NEBNext Quick Ligation Reaction Buffer (5X) 10 µl Quick T4 DNA Ligase 5 µl Total 50 µl -
Ensure the components are thoroughly mixed by pipetting, and spin down.
-
Incubate the reaction for 10 minutes at room temperature.
-
Resuspend the AMPure XP beads by vortexing.
-
Add 20 µl of resuspended AMPure XP beads to the reaction and mix by pipetting.
-
Incubate on a Hula mixer (rotator mixer) for 10 minutes at room temperature.
-
Spin down the sample and pellet the beads on a magnet for 5 minutes. Keep the tube on the magnet until the eluate is clear and colourless, and pipette off the supernatant.
-
Wash the beads by adding 125 μl Short Fragment Buffer (SFB). Flick the beads to resuspend, then return the tube to the magnetic rack and allow the beads to pellet. Keep the tube on the magnet until the eluate is clear and colourless. Remove the supernatant using a pipette and discard.
-
Repeat the previous step.
-
Spin down and place the tube back on the magnet. Pipette off any residual supernatant. Allow to dry for ~30 seconds, but do not dry the pellet to the point of cracking.
-
Remove the tube from the magnetic rack and resuspend the pellet by pipetting in 15 µl Elution Buffer (EB). Spin down and incubate for 5 minutes at room temperature.
-
Pellet the beads on a magnet until the eluate is clear and colourless, for at least 1 minute.
-
Remove and retain 15 µl of eluate containing the DNA library into a clean 1.5 ml Eppendorf DNA LoBind tube.
Dispose of the pelleted beads
-
Quantify 1 µl of eluted sample using a Qubit fluorometer.
-
Optional ActionIf quantities allow, the library may be diluted in Elution Buffer (EB) for splitting across multiple flow cells.
Additional buffer for doing this can be found in the Sequencing Auxiliary Vials expansion (EXP-AUX001), available to purchase separately. This expansion also contains additional vials of Sequencing Buffer (SQB) and Loading Beads (LB), required for loading the libraries onto flow cells.
Priming and loading the SpotON flow cell
Priming and loading the SpotON flow cell
- Materials
-
- Flush Tether (FLT)
- Flush Buffer (FB)
- Sequencing Buffer (SQB)
- Loading Beads (LB)
- Consumables
-
- 1.5 ml Eppendorf DNA LoBind tubes
- Equipment
-
- MinION Mk1B or Mk1C
- SpotON Flow Cell
- P1000 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- Vortex mixer
- Microfuge
-
Thaw the Sequencing Buffer (SQB), Loading Beads (LB), Flush Tether (FLT) and one tube of Flush Buffer (FB) at room temperature before mixing the reagents by vortexing, and spin down at room temperature.
-
To prepare the flow cell priming mix, add 30 µl of thawed and mixed Flush Tether (FLT) directly to the tube of thawed and mixed Flush Buffer (FB), and mix by vortexing at room temperature.
-
Open the MinION device lid and slide the flow cell under the clip.
Press down firmly on the flow cell to ensure correct thermal and electrical contact.
-
Optional ActionComplete a flow cell check to assess the number of pores available before loading the library.
This step can be omitted if the flow cell has been checked previously.
See the flow cell check instructions in the MinKNOW protocol for more information.
-
Slide the flow cell priming port cover clockwise to open the priming port.
-
After opening the priming port, check for a small air bubble under the cover. Draw back a small volume to remove any bubbles:
- Set a P1000 pipette to 200 µl
- Insert the tip into the priming port
- Turn the wheel until the dial shows 220-230 µl, to draw back 20-30 µl, or until you can see a small volume of buffer entering the pipette tip
Note: Visually check that there is continuous buffer from the priming port across the sensor array.
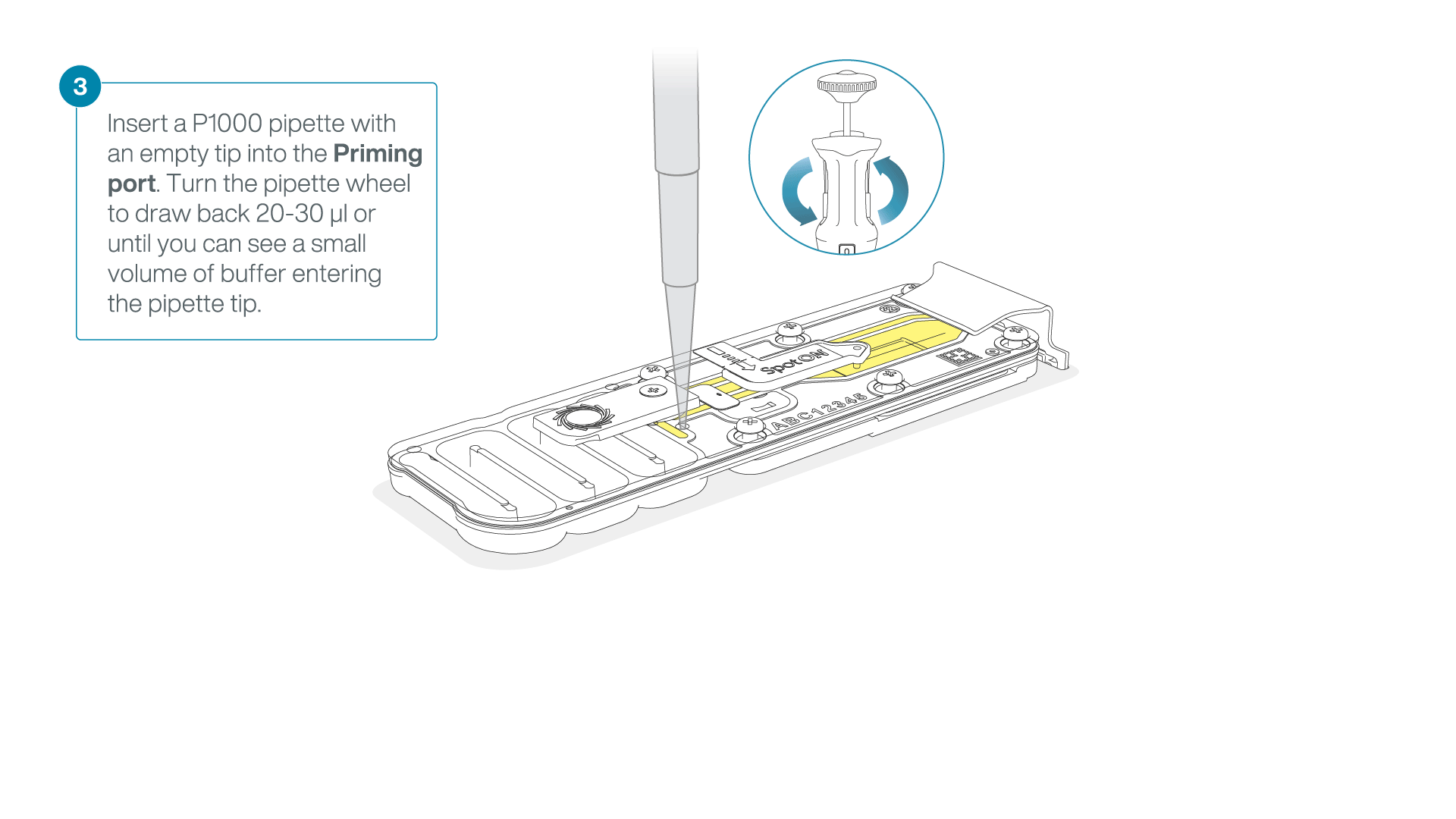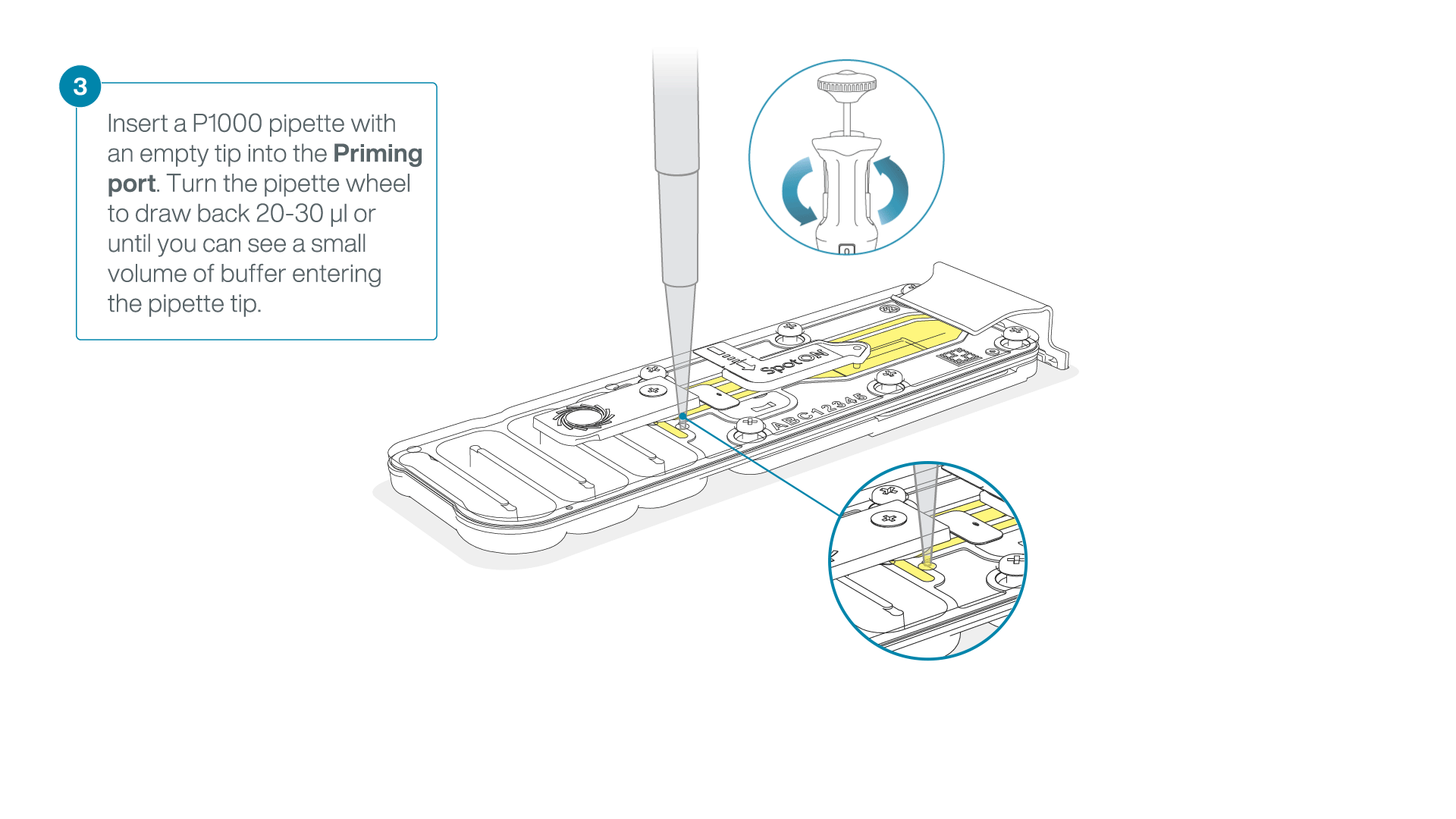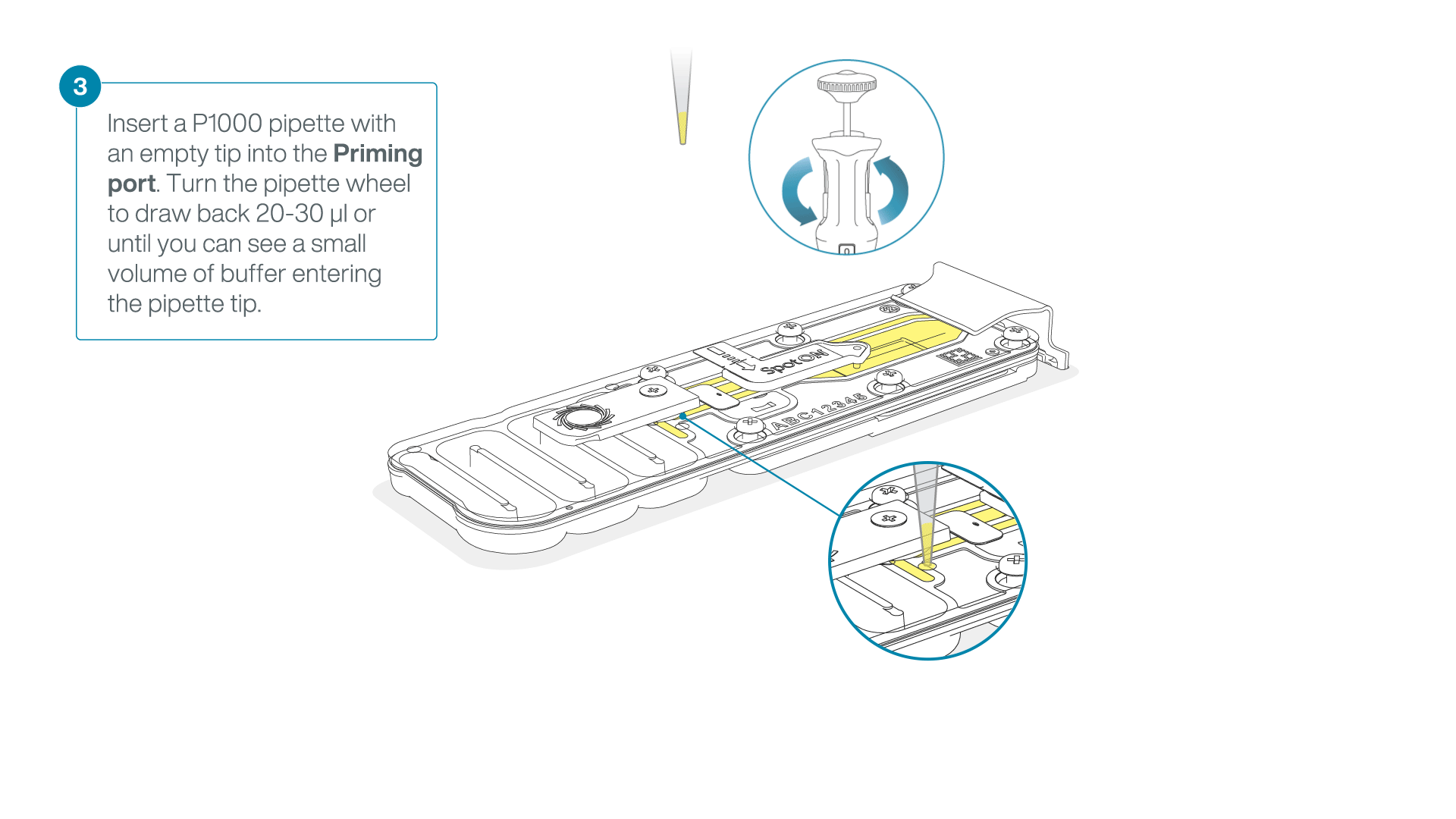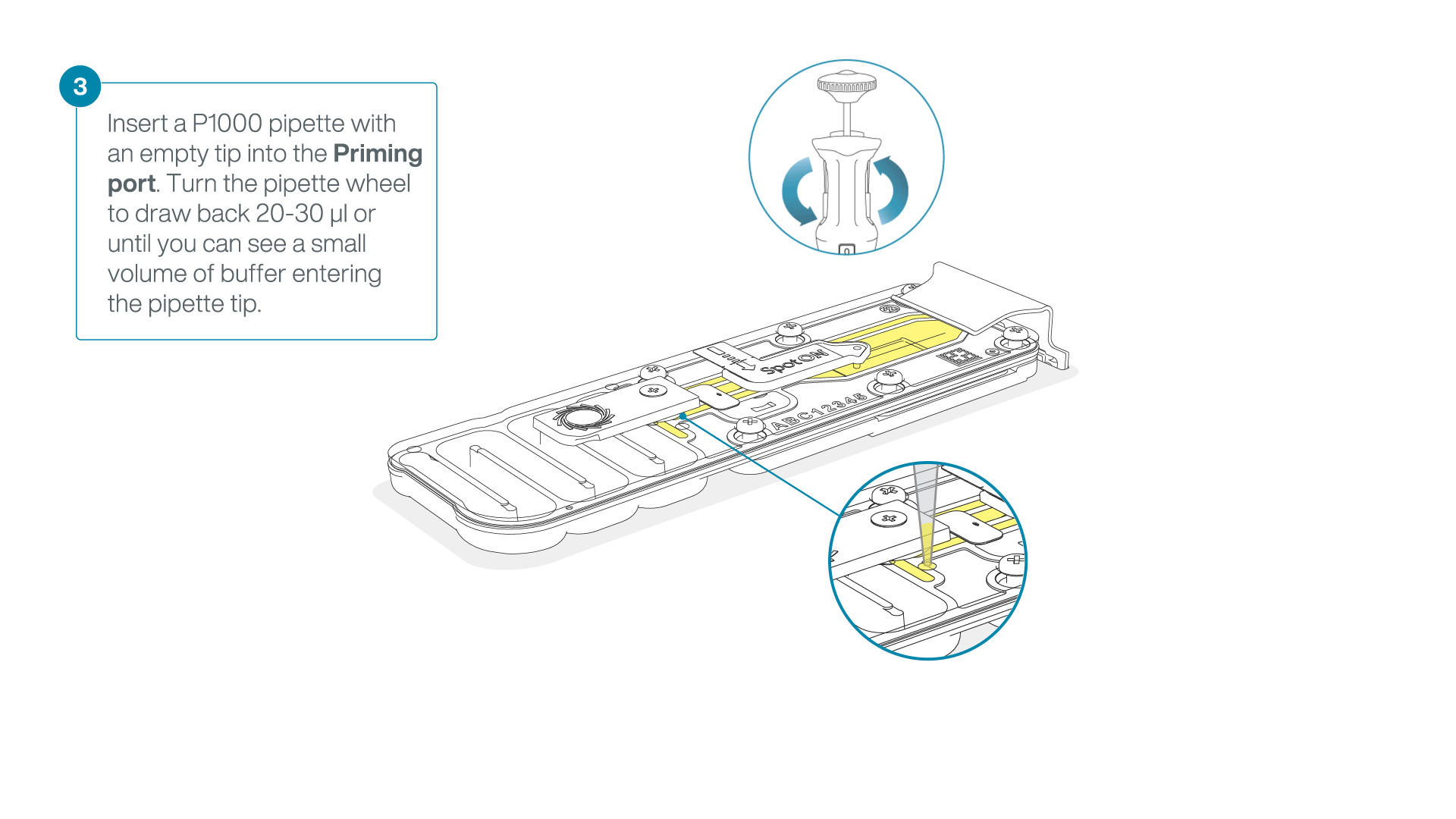
-
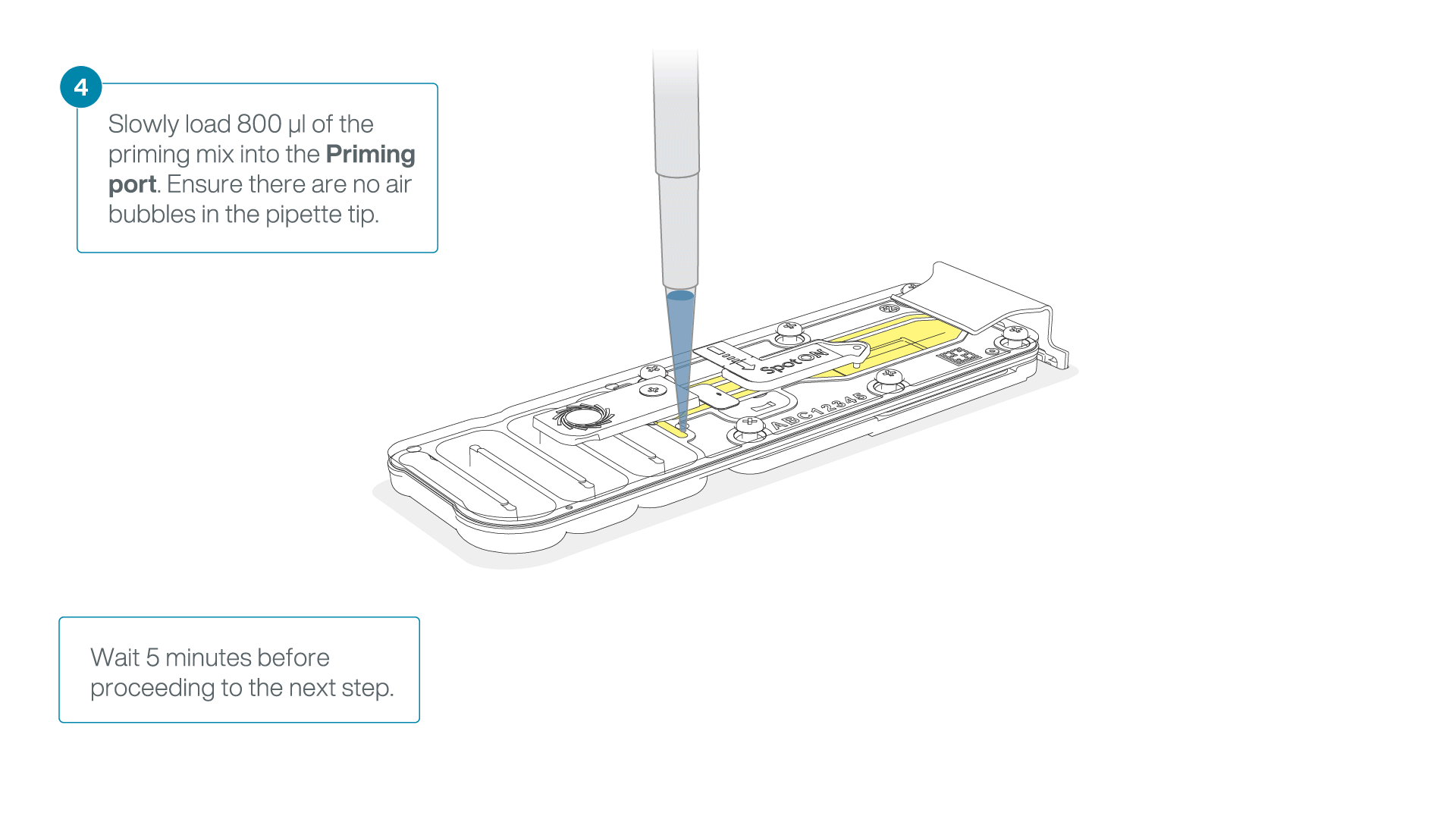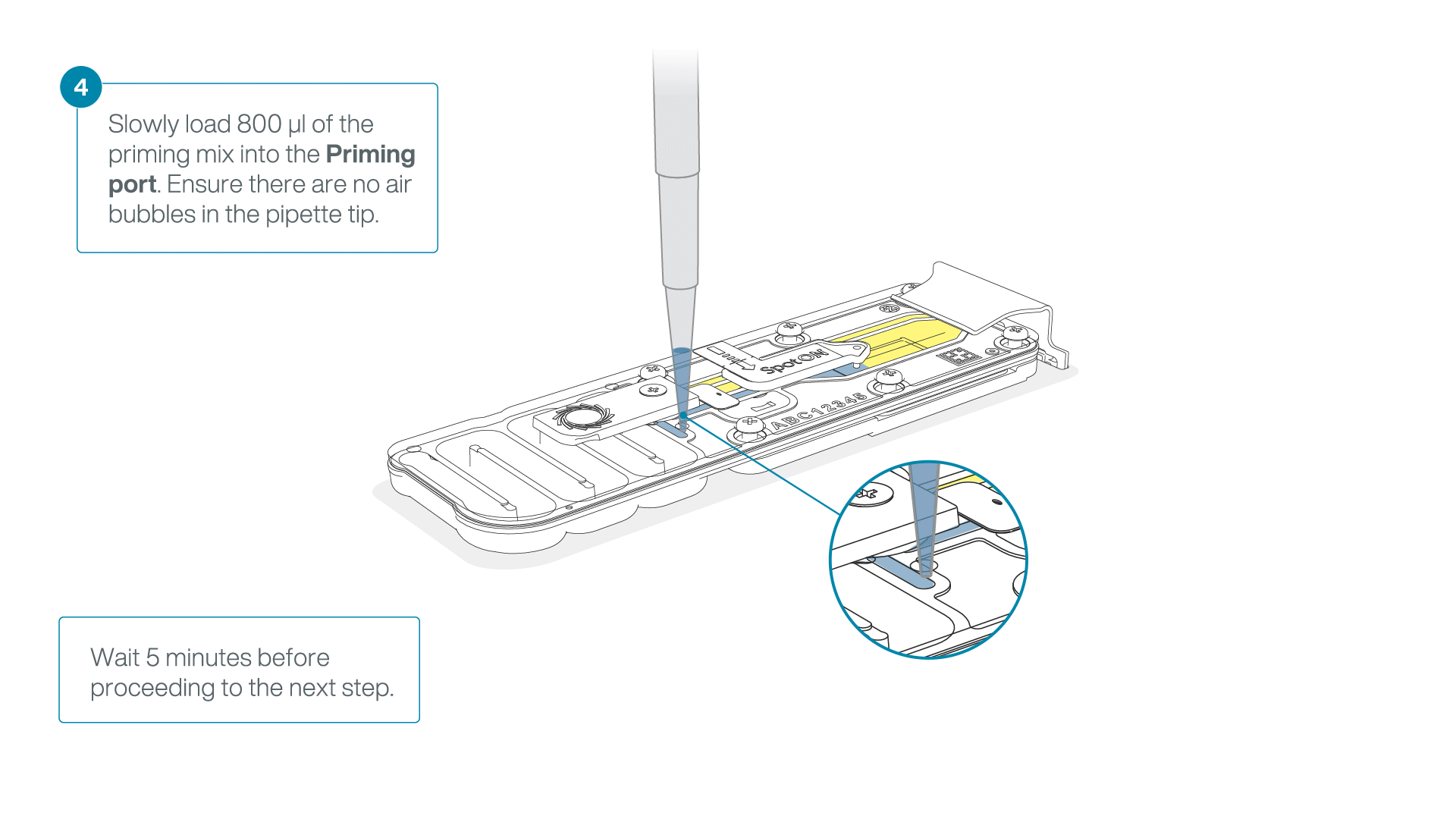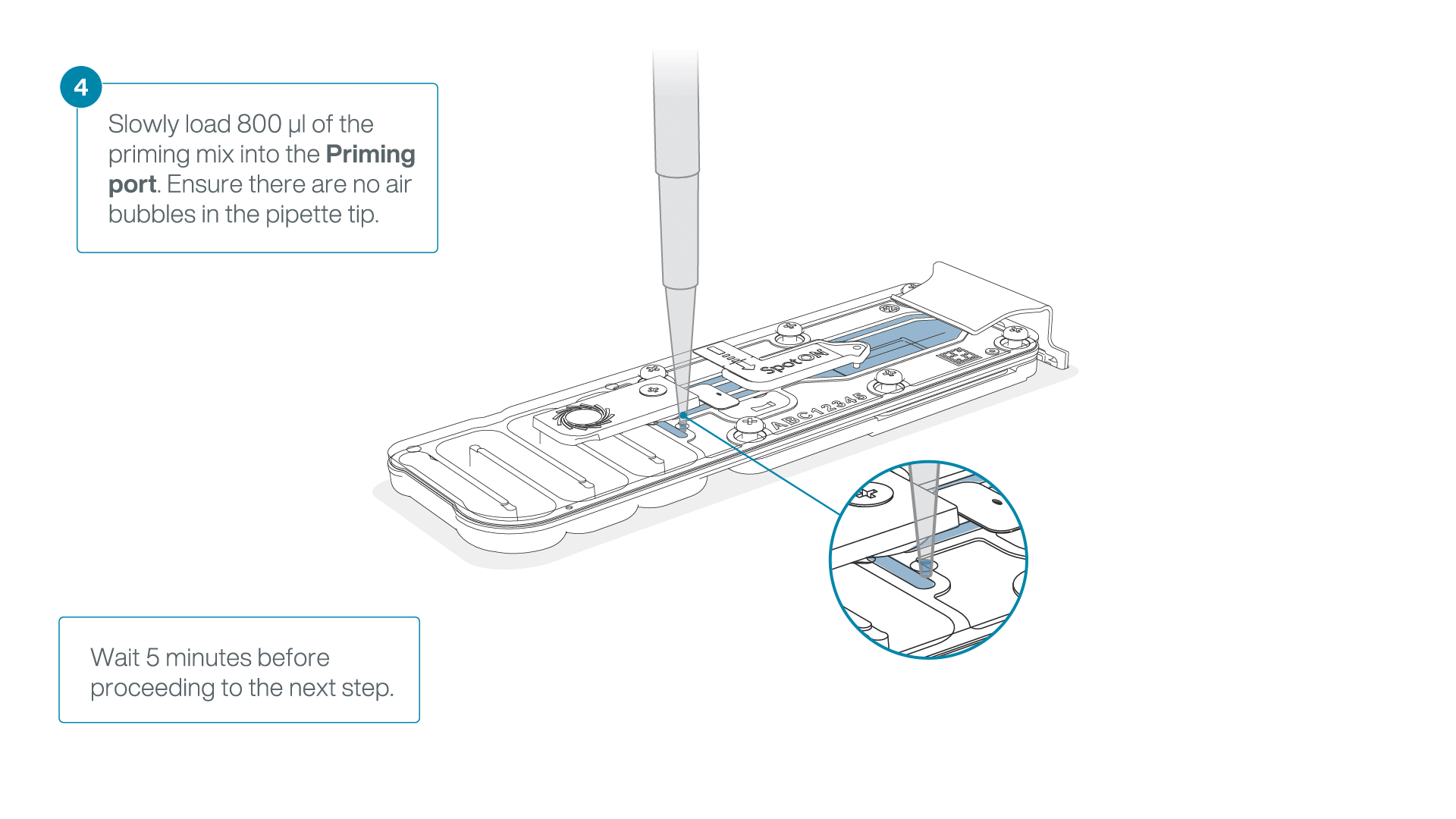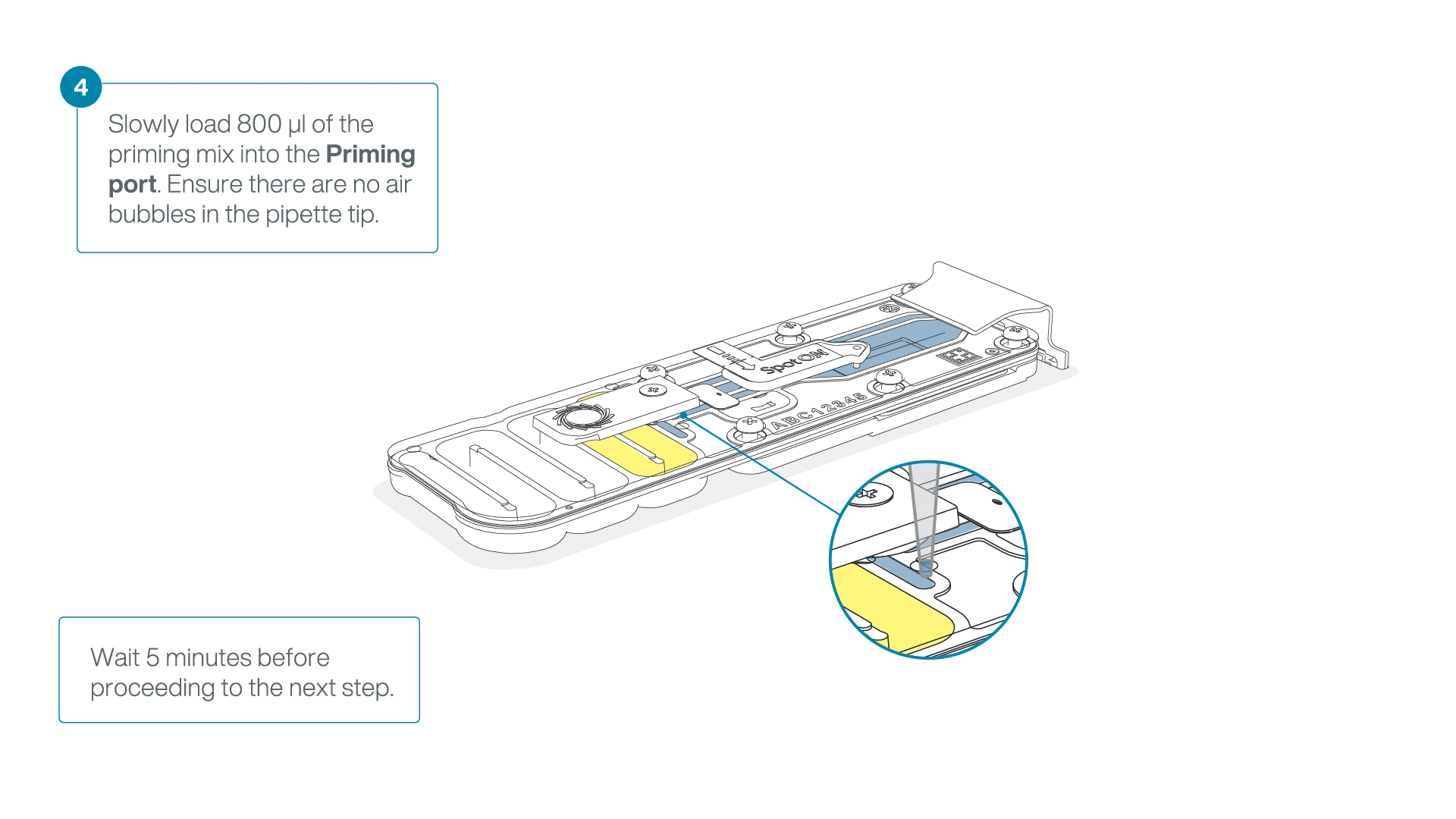
Load 800 µl of the priming mix into the flow cell via the priming port, avoiding the introduction of air bubbles. Wait for five minutes. During this time, prepare the library for loading by following the steps below.
-
Thoroughly mix the contents of the Loading Beads (LB) by pipetting.
-
In a new tube, prepare the library for loading as follows:
Reagent Volume per flow cell Sequencing Buffer (SQB) 37.5 µl Loading Beads (LB), mixed immediately before use 25.5 µl DNA library 12 µl Total 75 µl Note: Load the library onto the flow cell immediately after adding the Sequencing Buffer (SQB) and Loading Beads (LB) because the fuel in the buffer will start to be consumed by the adapter.
-
Complete the flow cell priming:
- Gently lift the SpotON sample port cover to make the SpotON sample port accessible.
- Load 200 µl of the priming mix into the flow cell priming port (not the SpotON sample port), avoiding the introduction of air bubbles.
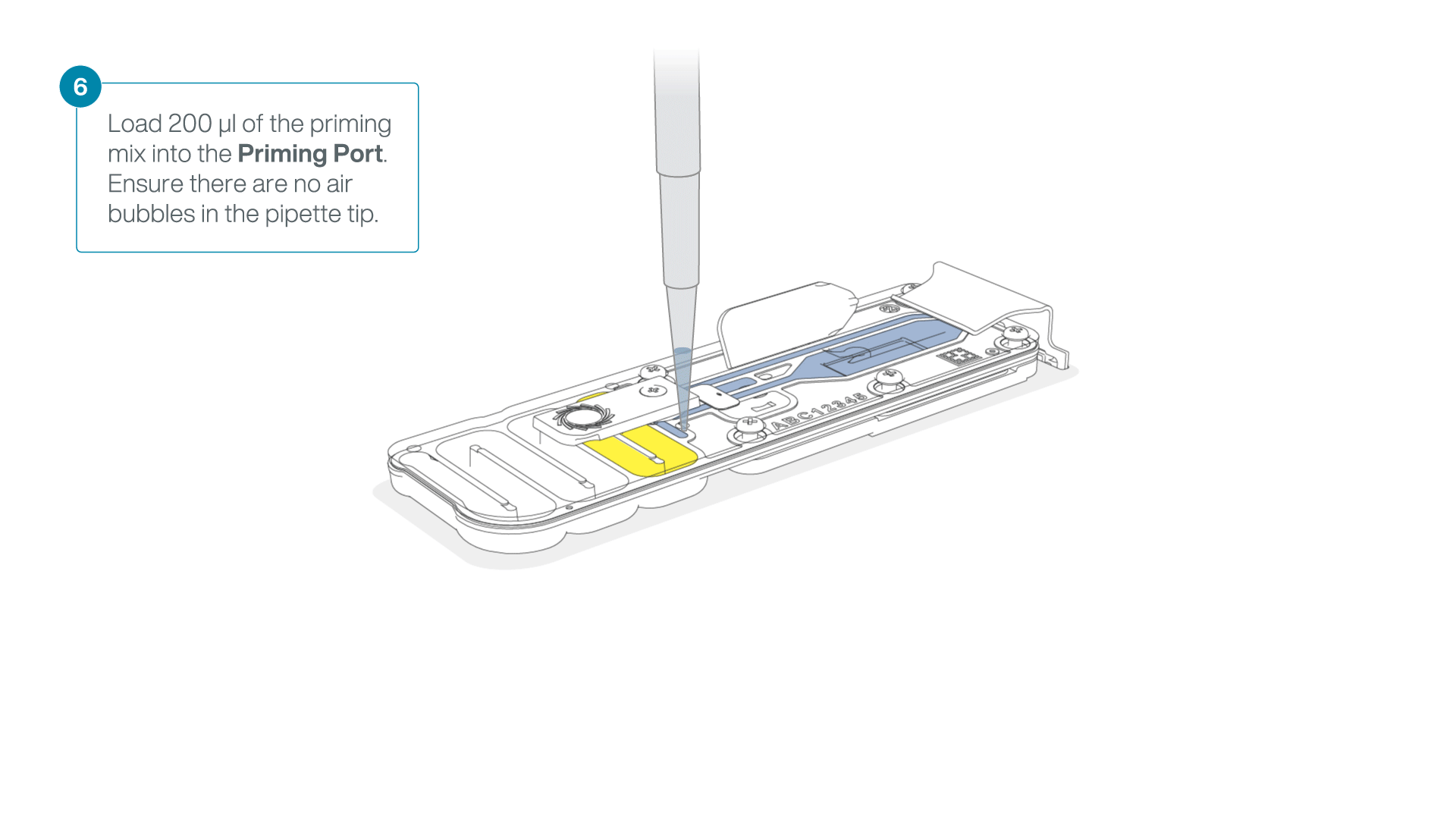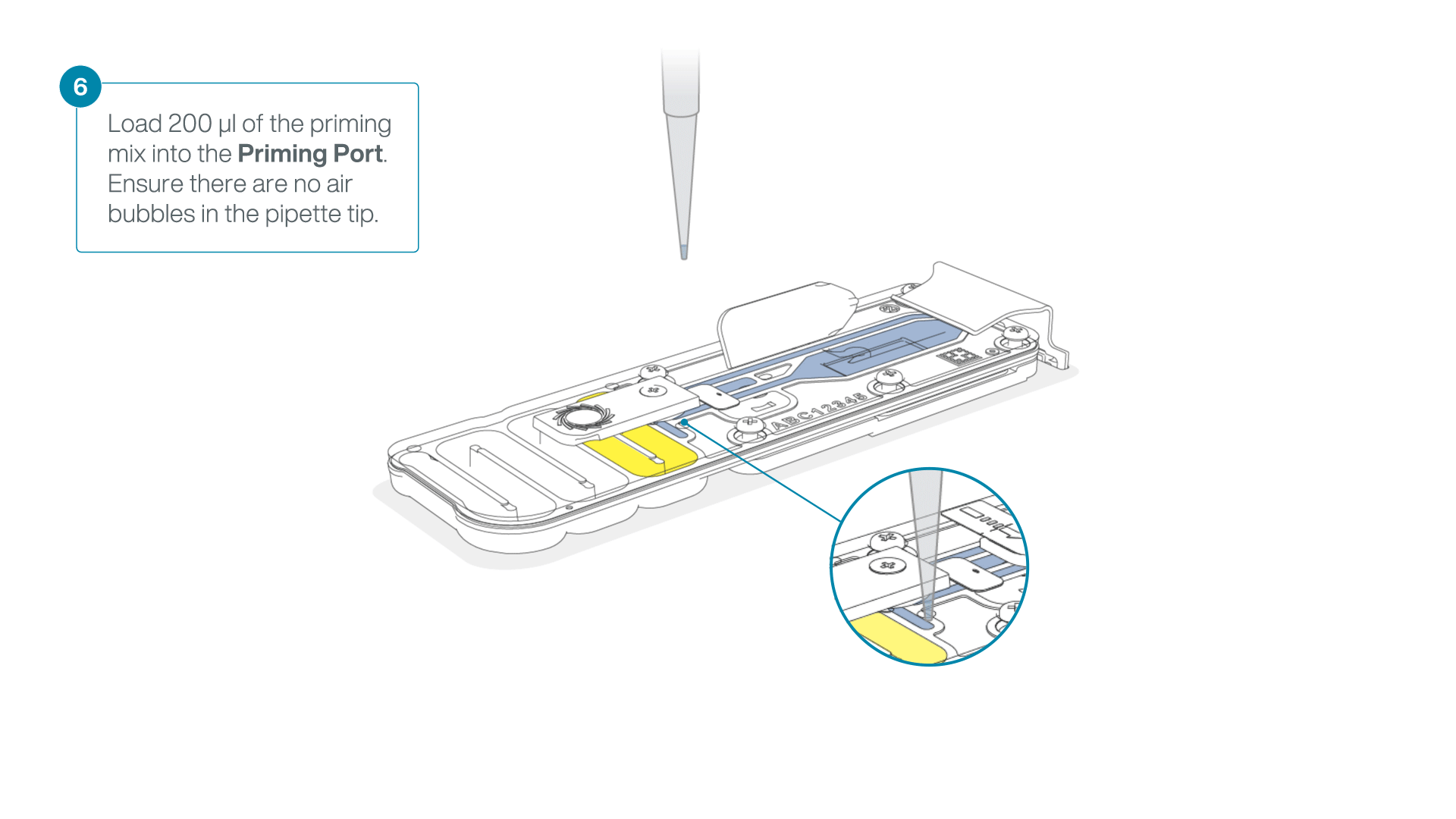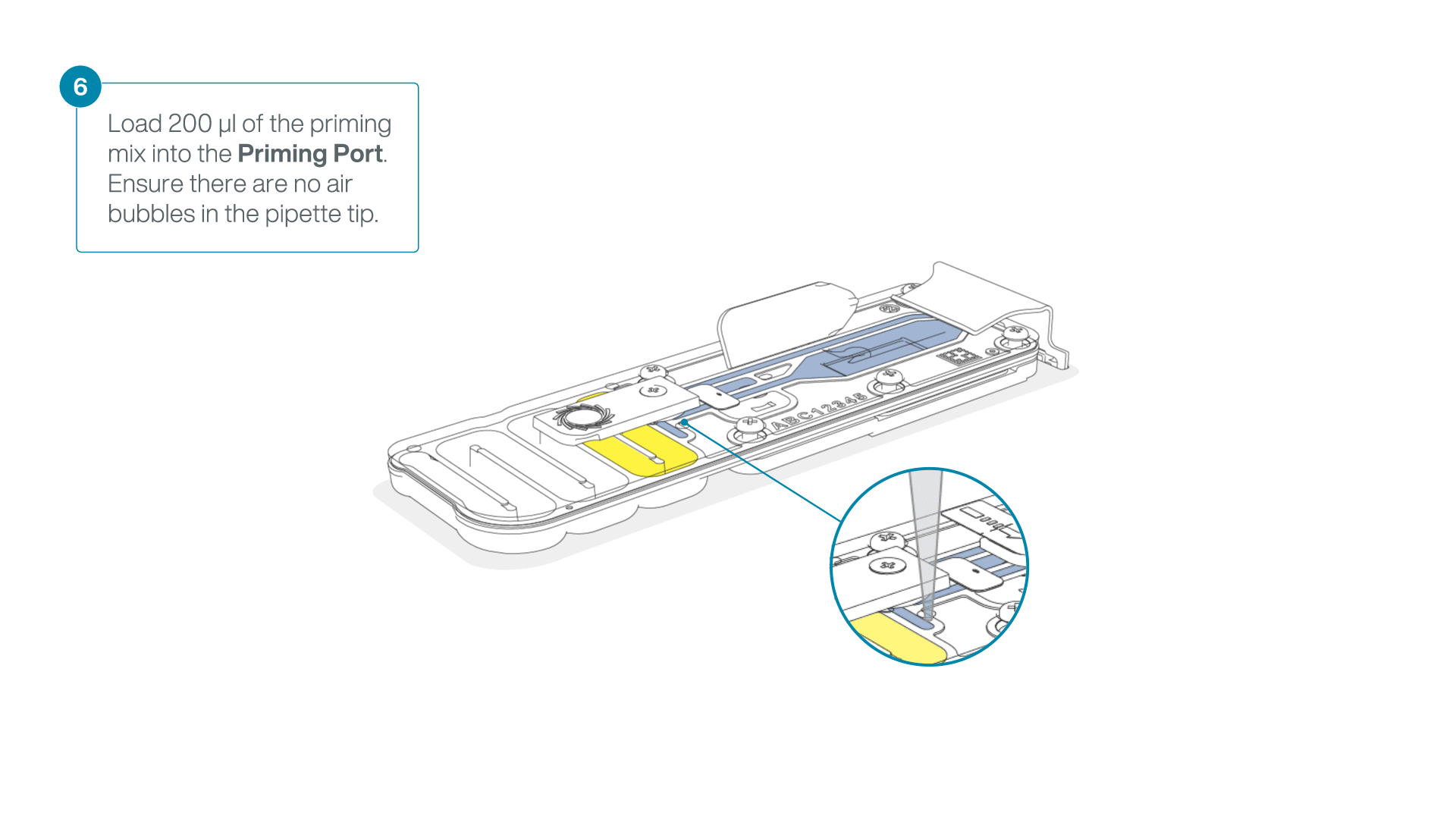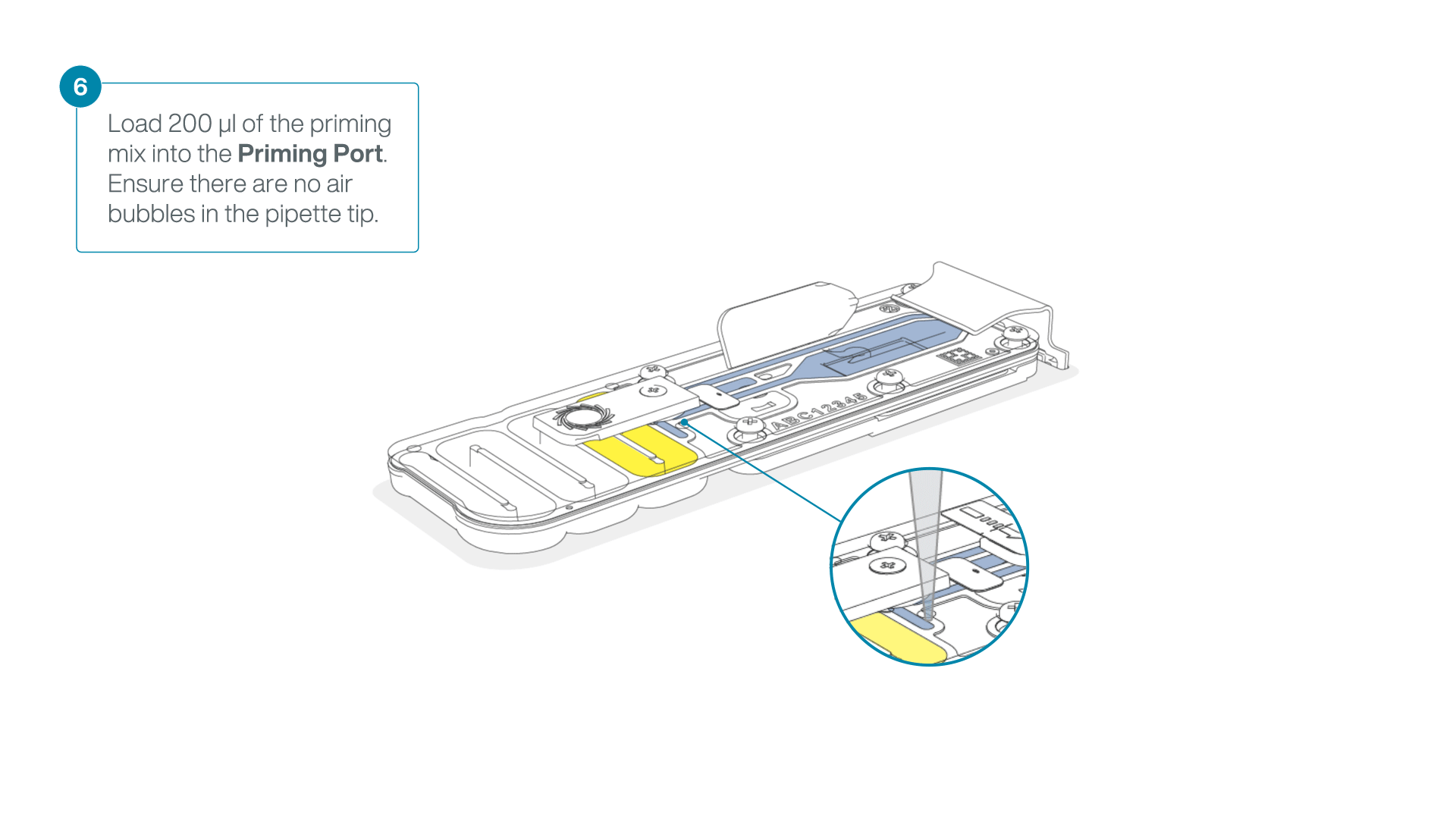
-
Mix the prepared library gently by pipetting up and down just prior to loading.
-
Add 75 μl of the prepared library to the flow cell via the SpotON sample port in a dropwise fashion. Ensure each drop flows into the port before adding the next.
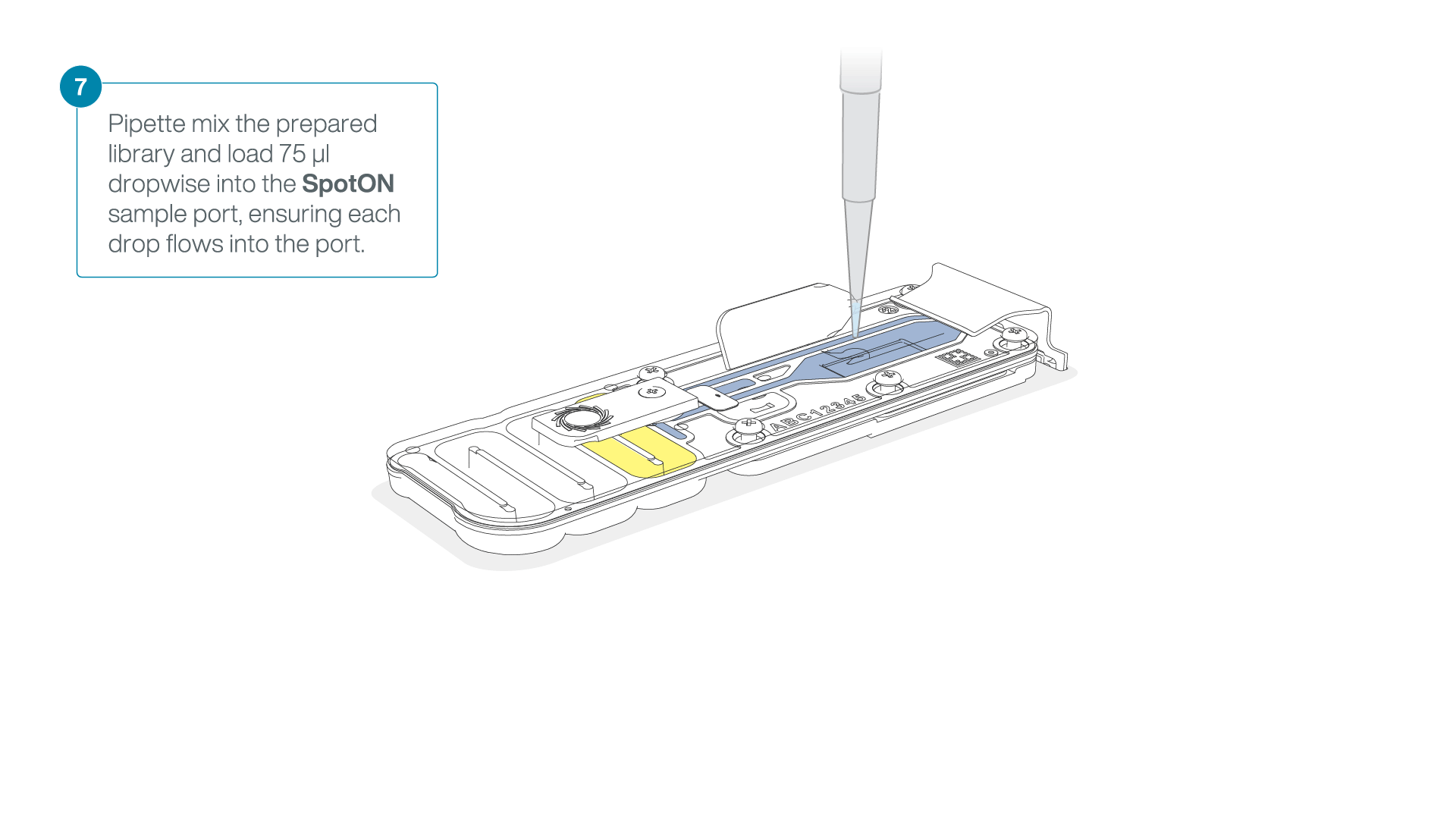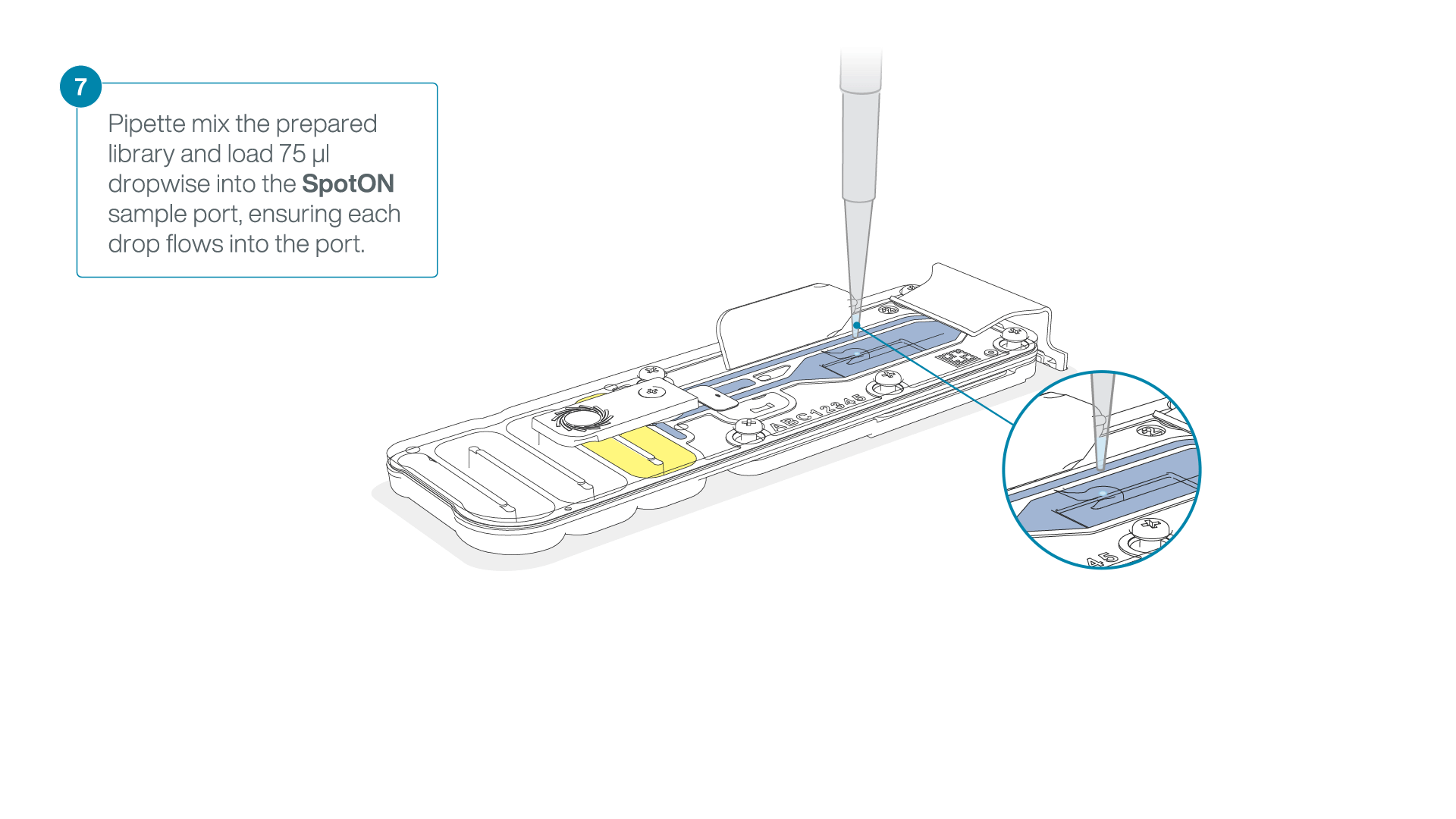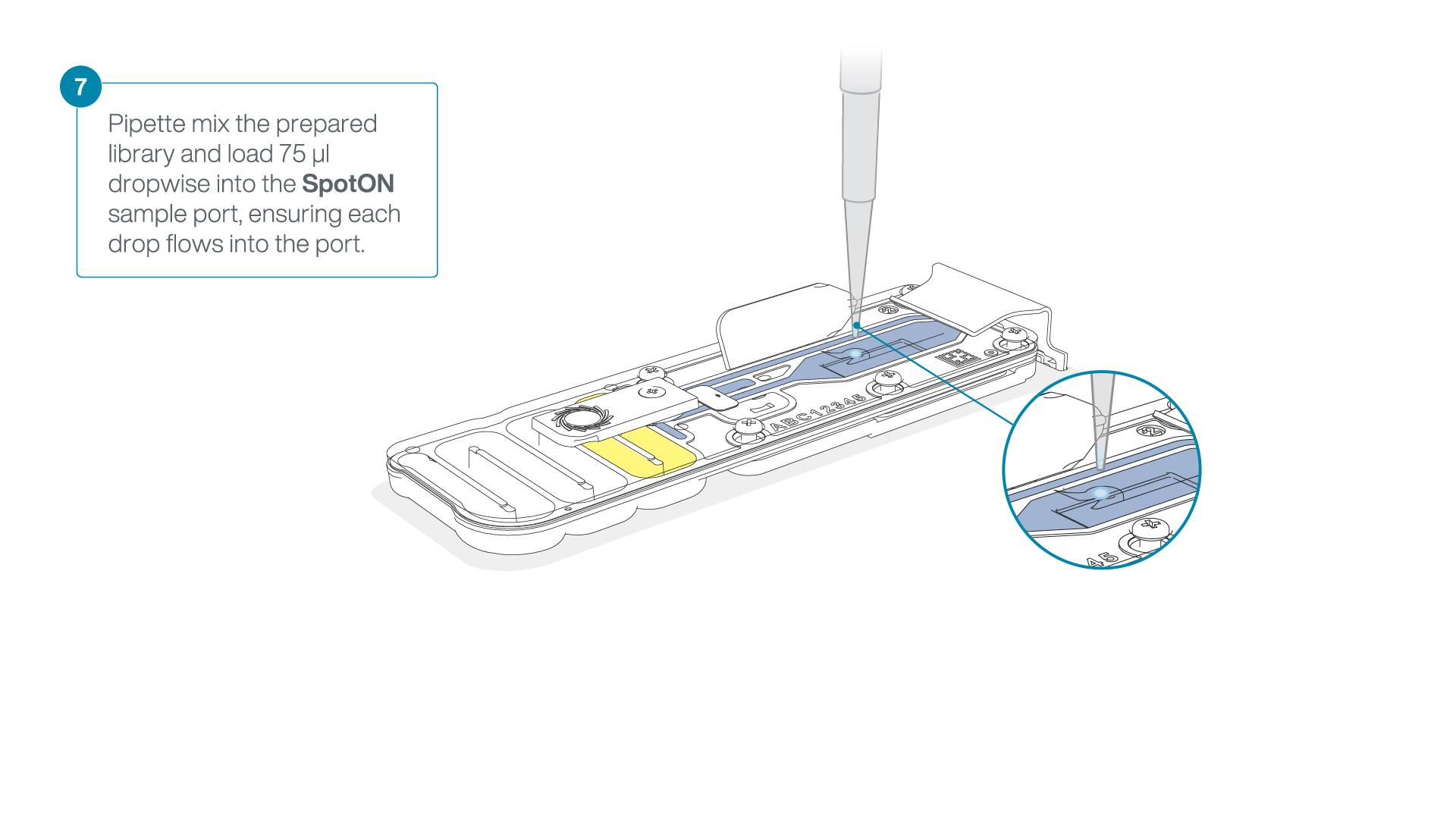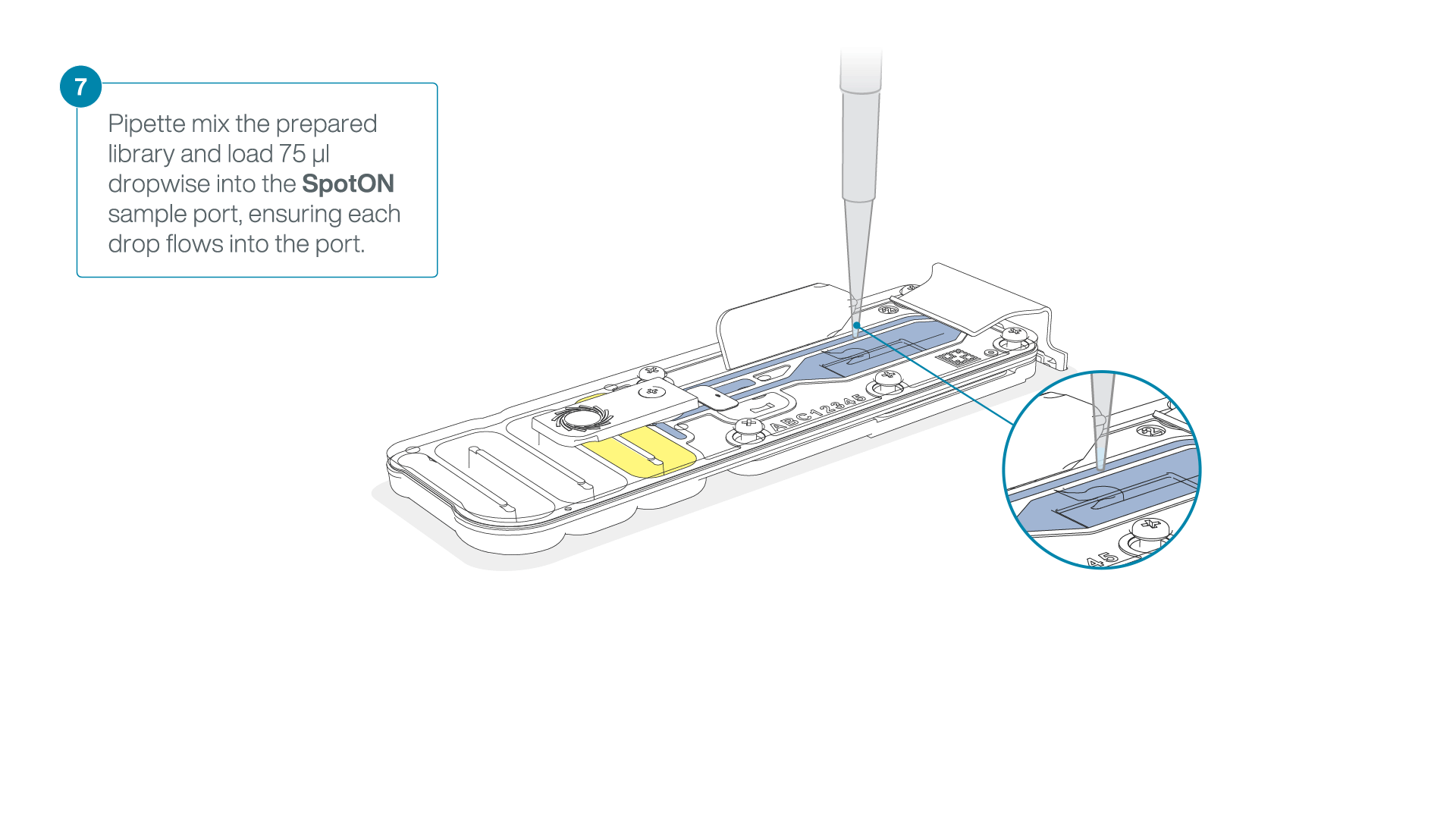
-
Gently replace the SpotON sample port cover, making sure the bung enters the SpotON port, close the priming port and replace the MinION device lid.
Sequencing and data analysis
Data acquisition and basecalling
Data acquisition and basecalling
-
Overview of nanopore data analysis
For a full overview of nanopore data analysis, which includes options for basecalling and post-basecalling analysis, please refer to the Data Analysis document.
-
How to start sequencing
The sequencing device control, data acquisition and real-time basecalling are carried out by the MinKNOW software. Please ensure MinKNOW is installed on your computer or device. There are multiple options for how to carry out sequencing:
1. Data acquisition and basecalling in real-time using MinKNOW on a computer
Follow the instructions in the MinKNOW protocol beginning from the "Starting a sequencing run" section until the end of the "Completing a MinKNOW run" section.
2. Data acquisition and basecalling in real-time using the MinION Mk1B/Mk1D device
Follow the instructions in the MinION Mk1B user manual or the MinION Mk1D user manual.
3. Data acquisition and basecalling in real-time using the MinION Mk1C device
Follow the instructions in the MinION Mk1C user manual.
4. Data acquisition and basecalling in real-time using the GridION device
Follow the instructions in the GridION user manual.
5. Data acquisition and basecalling in real-time using the PromethION device
Follow the instructions in the PromethION user manual or the PromethION 2 Solo user manual.
6. Data acquisition using MinKNOW on a computer and basecalling at a later time using MinKNOW
Follow the instructions in the MinKNOW protocol beginning from the "Starting a sequencing run" section until the end of the "Completing a MinKNOW run" section. When setting your experiment parameters, set the Basecalling tab to OFF. After the sequencing experiment has completed, follow the instructions in the Post-run analysis section of the MinKNOW protocol.
Downstream analysis
Downstream analysis
-
Recommended analysis pipeline
The analysis of the FASTQ format sequence data is performed using a Nextflow workflow called wf-flu. This workflow is accessible through the Nextflow command-line software and may also be run using the graphical interface provided by EPI2ME Labs.
The workflow processes the basecalled and demultiplexed DNA sequence data output by MinKNOW. The sequences are filtered for a minimum length and quality thresholds (200 nucleotides and Q9 respectively) prior to sequence alignment to the CDC multi-fasta Influenza reference. The alignment is performed using the Minimap2 software. Depth of coverage across the mapped sequences is measured using Samtools before genetic variants are called using Medaka. A coverage-masked consensus sequence is prepared for each sample using bcftools. The influenza strain typing is then performed using the abricate software with an insaflu database. The influenza strains included in the database are listed in the project documentation pages at https://github.com/epi2me-labs/wf-flu.
The workflow returns a per-run HTML-format summary report along with a CSV file of typing results. Additional files that include mapping BAM files and VCF files of Medaka variants are also included in the workflow output.
For more information, please refer to the Influenza workflow blog.
-
Software set-up and installation
The wf-flu workflow requires the Nextflow and either the Docker or Conda software to have been installed. The EPI2ME Labs Workflow quick start guide provides instructions for the installation of these requirements for GridION, PromethION and general Ubuntu Linux users and provides a little more introduction to the Nextflow software.
To run the EPI2ME Labs via the GUI instead of the command-line, download the executable for your operating system here: https://labs.epi2me.io/downloads and consult the Quick Start Guide to set up and run the software.
Flow cell reuse and returns
Flow cell reuse and returns
- Materials
-
- Flow Cell Wash Kit (EXP-WSH004)
-
After your sequencing experiment is complete, if you would like to reuse the flow cell, please follow the Flow Cell Wash Kit protocol and store the washed flow cell at +2°C to +8°C.
The Flow Cell Wash Kit protocol is available on the Nanopore Community.
-
Alternatively, follow the returns procedure to send the flow cell back to Oxford Nanopore.
Instructions for returning flow cells can be found here.
Troubleshooting
Issues during DNA extraction and library preparation
Issues during DNA extraction and library preparation
-
Below is a list of the most commonly encountered issues, with some suggested causes and solutions.
We also have an FAQ section available on the Nanopore Community Support section.
If you have tried our suggested solutions and the issue still persists, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community.
-
Low sample quality
Observation Possible cause Comments and actions Low DNA purity (Nanodrop reading for DNA OD 260/280 is <1.8 and OD 260/230 is <2.0–2.2) The DNA extraction method does not provide the required purity The effects of contaminants are shown in the Contaminants document. Please try an alternative extraction method that does not result in contaminant carryover.
Consider performing an additional SPRI clean-up step.Low RNA integrity (RNA integrity number <9.5 RIN, or the rRNA band is shown as a smear on the gel) The RNA degraded during extraction Try a different RNA extraction method. For more info on RIN, please see the RNA Integrity Number document. Further information can be found in the DNA/RNA Handling page. RNA has a shorter than expected fragment length The RNA degraded during extraction Try a different RNA extraction method. For more info on RIN, please see the RNA Integrity Number document. Further information can be found in the DNA/RNA Handling page.
We recommend working in an RNase-free environment, and to keep your lab equipment RNase-free when working with RNA. -
Low output from PCR
Observation Possible cause Comments and actions Low output from PCR Insufficient primer Increase the amount of primer pool in the PCR reaction by 0.25-1 µl, decrease water proportionally. - Insufficient DNA template Use undiluted DNA. - Thermocycler fault Ensure that you use a thermocycler that has been recently calibrated. - Insufficient viral DNA and an abundance of host DNA Use a host depletion method such as NEBNext® Microbiome DNA Enrichment Kit before PCR*. - Low representation of a particular primer in the sequencing data Spike in a small quantity of the primer in the 100 µM pool before future runs * This may be prohibitively expensive at scale, but for precious samples this will have a noticeable impact on amplicon yield.
-
Low DNA recovery after AMPure bead clean-up
Observation Possible cause Comments and actions Low recovery DNA loss due to a lower than intended AMPure beads-to-sample ratio 1. AMPure beads settle quickly, so ensure they are well resuspended before adding them to the sample.
2. When the AMPure beads-to-sample ratio is lower than 0.4:1, DNA fragments of any size will be lost during the clean-up.Low recovery DNA fragments are shorter than expected The lower the AMPure beads-to-sample ratio, the more stringent the selection against short fragments. Please always determine the input DNA length on an agarose gel (or other gel electrophoresis methods) and then calculate the appropriate amount of AMPure beads to use. 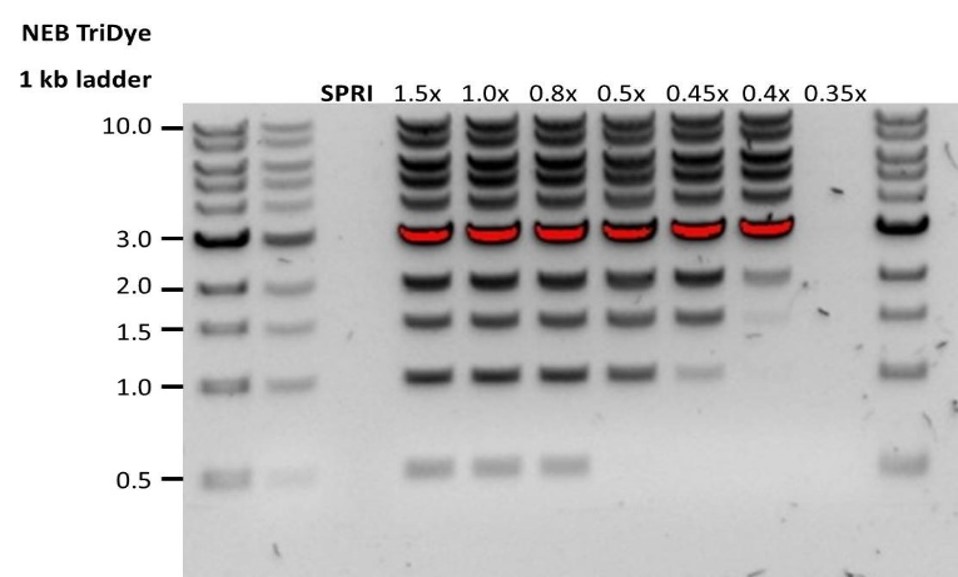
Low recovery after end-prep The wash step used ethanol <70% DNA will be eluted from the beads when using ethanol <70%. Make sure to use the correct percentage.
Issues during the sequencing run
Issues during the sequencing run
-
Below is a list of the most commonly encountered issues, with some suggested causes and solutions.
We also have an FAQ section available on the Nanopore Community Support section.
If you have tried our suggested solutions and the issue still persists, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community.
-
Fewer pores at the start of sequencing than after Flow Cell Check
Observation Possible cause Comments and actions MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check An air bubble was introduced into the nanopore array After the Flow Cell Check it is essential to remove any air bubbles near the priming port before priming the flow cell. If not removed, the air bubble can travel to the nanopore array and irreversibly damage the nanopores that have been exposed to air. The best practice to prevent this from happening is demonstrated in this video. MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check The flow cell is not correctly inserted into the device Stop the sequencing run, remove the flow cell from the sequencing device and insert it again, checking that the flow cell is firmly seated in the device and that it has reached the target temperature. If applicable, try a different position on the device (GridION/PromethION). MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check Contaminations in the library damaged or blocked the pores The pore count during the Flow Cell Check is performed using the QC DNA molecules present in the flow cell storage buffer. At the start of sequencing, the library itself is used to estimate the number of active pores. Because of this, variability of about 10% in the number of pores is expected. A significantly lower pore count reported at the start of sequencing can be due to contaminants in the library that have damaged the membranes or blocked the pores. Alternative DNA/RNA extraction or purification methods may be needed to improve the purity of the input material. The effects of contaminants are shown in the Contaminants Know-how piece. Please try an alternative extraction method that does not result in contaminant carryover. -
MinKNOW script failed
Observation Possible cause Comments and actions MinKNOW shows "Script failed" Restart the computer and then restart MinKNOW. If the issue persists, please collect the MinKNOW log files and contact Technical Support. If you do not have another sequencing device available, we recommend storing the flow cell and the loaded library at 4°C and contact Technical Support for further storage guidance. -
Pore occupancy below 40%
Observation Possible cause Comments and actions Pore occupancy <40% Not enough library was loaded on the flow cell Ensure you load the recommended amount of good quality library in the relevant library prep protocol onto your flow cell. Please quantify the library before loading and calculate mols using tools like the Promega Biomath Calculator, choosing "dsDNA: µg to pmol" Pore occupancy close to 0 The Ligation Sequencing Kit was used, and sequencing adapters did not ligate to the DNA Make sure to use the NEBNext Quick Ligation Module (E6056) and Oxford Nanopore Technologies Ligation Buffer (LNB, provided in the sequencing kit) at the sequencing adapter ligation step, and use the correct amount of each reagent. A Lambda control library can be prepared to test the integrity of the third-party reagents. Pore occupancy close to 0 The Ligation Sequencing Kit was used, and ethanol was used instead of LFB or SFB at the wash step after sequencing adapter ligation Ethanol can denature the motor protein on the sequencing adapters. Make sure the LFB or SFB buffer was used after ligation of sequencing adapters. Pore occupancy close to 0 No tether on the flow cell Tethers are adding during flow cell priming (FLT/FCT tube). Make sure FLT/FCT was added to FB/FCF before priming. -
Shorter than expected read length
Observation Possible cause Comments and actions Shorter than expected read length Unwanted fragmentation of DNA sample Read length reflects input DNA fragment length. Input DNA can be fragmented during extraction and library prep.
1. Please review the Extraction Methods in the Nanopore Community for best practice for extraction.
2. Visualise the input DNA fragment length distribution on an agarose gel before proceeding to the library prep.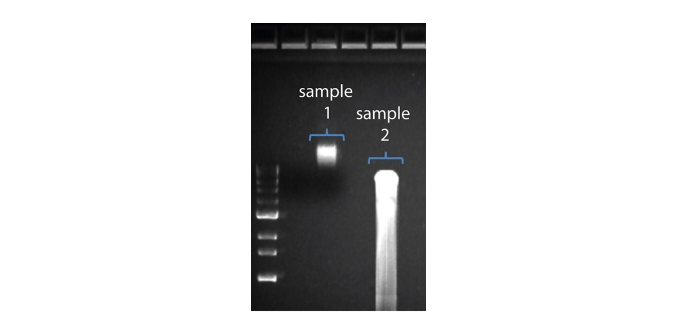 In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented.
In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented.
3. During library prep, avoid pipetting and vortexing when mixing reagents. Flicking or inverting the tube is sufficient. -
Large proportion of inactive pores
Observation Possible cause Comments and actions Large proportion of inactive/unavailable pores (shown as light blue in the channels panel and pore activity plot. Pores or membranes are irreversibly damaged) Air bubbles have been introduced into the flow cell Air bubbles introduced through flow cell priming and library loading can irreversibly damage the pores. Watch the Priming and loading your flow cell video for best practice Large proportion of inactive/unavailable pores Certain compounds co-purified with DNA Known compounds, include polysaccharides, typically associate with plant genomic DNA.
1. Please refer to the Plant leaf DNA extraction method.
2. Clean-up using the QIAGEN PowerClean Pro kit.
3. Perform a whole genome amplification with the original gDNA sample using the QIAGEN REPLI-g kit.Large proportion of inactive/unavailable pores Contaminants are present in the sample The effects of contaminants are shown in the Contaminants Know-how piece. Please try an alternative extraction method that does not result in contaminant carryover. -
Reduction in sequencing speed and q-score later into the run
Observation Possible cause Comments and actions Reduction in sequencing speed and q-score later into the run For Kit 9 chemistry (e.g. SQK-LSK109), fast fuel consumption is typically seen when the flow cell is overloaded with library (please see the appropriate protocol for your DNA library to see the recommendation). Add more fuel to the flow cell by following the instructions in the MinKNOW protocol. In future experiments, load lower amounts of library to the flow cell. -
Temperature fluctuation
Observation Possible cause Comments and actions Temperature fluctuation The flow cell has lost contact with the device Check that there is a heat pad covering the metal plate on the back of the flow cell. Re-insert the flow cell and press it down to make sure the connector pins are firmly in contact with the device. If the problem persists, please contact Technical Services. -
Failed to reach target temperature
Observation Possible cause Comments and actions MinKNOW shows "Failed to reach target temperature" The instrument was placed in a location that is colder than normal room temperature, or a location with poor ventilation (which leads to the flow cells overheating) MinKNOW has a default timeframe for the flow cell to reach the target temperature. Once the timeframe is exceeded, an error message will appear and the sequencing experiment will continue. However, sequencing at an incorrect temperature may lead to a decrease in throughput and lower q-scores. Please adjust the location of the sequencing device to ensure that it is placed at room temperature with good ventilation, then re-start the process in MinKNOW. Please refer to this link for more information on MinION temperature control. -
Guppy – no input .fast5 was found or basecalled
Observation Possible cause Comments and actions No input .fast5 was found or basecalled input_path did not point to the .fast5 file location The --input_path has to be followed by the full file path to the .fast5 files to be basecalled, and the location has to be accessible either locally or remotely through SSH. No input .fast5 was found or basecalled The .fast5 files were in a subfolder at the input_path location To allow Guppy to look into subfolders, add the --recursive flag to the command -
Guppy – no Pass or Fail folders were generated after basecalling
Observation Possible cause Comments and actions No Pass or Fail folders were generated after basecalling The --qscore_filtering flag was not included in the command The --qscore_filtering flag enables filtering of reads into Pass and Fail folders inside the output folder, based on their strand q-score. When performing live basecalling in MinKNOW, a q-score of 7 (corresponding to a basecall accuracy of ~80%) is used to separate reads into Pass and Fail folders. -
Guppy – unusually slow processing on a GPU computer
Observation Possible cause Comments and actions Unusually slow processing on a GPU computer The --device flag wasn't included in the command The --device flag specifies a GPU device to use for accelerate basecalling. If not included in the command, GPU will not be used. GPUs are counted from zero. An example is --device cuda:0 cuda:1, when 2 GPUs are specified to use by the Guppy command.
Become a full member
Purchase a MinION Starter Pack from Avantor to get full community access and benefit from:
- News - hear about the latest product updates
- Posts - interact with thousands of nanopore users from around the globe
- Software - download the latest sequencing and analysis software
Already have a Nanopore Community account?
Log in hereNeed more help?
Request a call with our experts for detailed advice on implementing nanopore sequencing.
Request a callInterested in microbiology?
Visit our microbial sequencing spotlight page on vwr.com.
Microbial sequencing






 In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented.
In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented.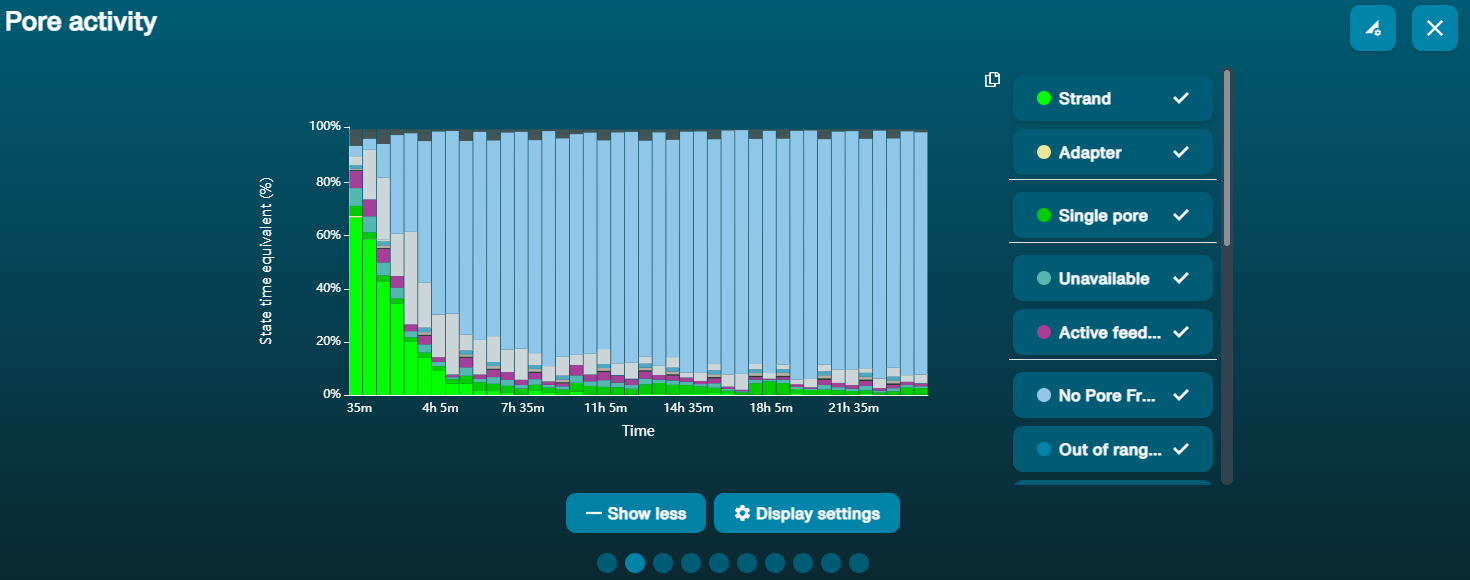 The pore activity plot above shows an increasing proportion of "unavailable" pores over time.
The pore activity plot above shows an increasing proportion of "unavailable" pores over time.