- Materials
-
- 1.5-3 µg (or 150-300 fmol) high molecular weight genomic DNA for R10.3 flow cells
- Consumables
-
- Nuclease-free water (e.g. ThermoFisher, AM9937)
- NEBNext FFPE DNA Repair Mix (NEB, M6630)
- NEBNext® Ultra II End Repair / dA-tailing Module (NEB, E7546)
- Agencourt AMPure XP beads (Beckman Coulter™, A63881)
- Freshly prepared 80% ethanol in nuclease-free water
- Thermo Scientific™ Abgene™ 96 Well 0.8 ml Polypropylene Deepwell Storage Plate (Thermo Scientific™, cat # AB0859)
- Sarstedt Inc Screw Cap Micro tube 2 ml, PP 1000/case (e.g. FisherScientific, Cat# NC0418367)
- Roche Diagnotics MagNA Pure LC Medium Reagent Tubs 20 (Cat# 03004058001)
- Bio-Rad Hard-Shell® 96-Well PCR Plates (Cat# HSP9601)
- Hamilton PCR ComfortLid (Cat# 814300)
- Hamilton 60 ml Reagent Reservoir, Self-Standing with Lid (Cat# 56694-01)
- Hamilton 1000 µl CO-RE tips with filter (Cat# 235905)
- Hamilton 300 µl CO-RE tips with filter (Cat# 235903)
- Hamilton 50 µl CO-RE tips with filter (Cat# 235948)
- Equipment
-
- Ice bucket with ice
- P1000 pipette and tips
- P200 pipette and tips
- P100 pipette and tips
- P10 pipette and tips
- Microplate centrifuge, e.g. Fisherbrand™ Mini Plate Spinner Centrifuge (Fisher Scientific, 11766427)
- Vortex mixer
-
Consumables and equipment quantities:
Consumable/equipment X24 samples X48 samples X96 samples Hamilton 50 µl CO-RE tips with filter 96 192 384 Hamilton 300 µl CO-RE tips with filter 201 298 490 Hamilton 1000 µl CO-RE tips with filter 32 32 32 Hamilton 60 ml Reagent Reservoir, Self-Standing with Lid 2 (1 EtOH & H20) 2 (1 EtOH & H20) 2 (1 EtOH & H20) Hamilton PCR ComfortLid 1 1 1 Bio-Rad Hard-Shell® 96-Well PCR Plate 1 (1 input sample & 1 end prepped sample) 1 (1 input sample & 1 end prepped sample) 1 (1 input sample & 1 end prepped sample) Hamilton 20 ml Reagent Reservoirs 1 1 1 Sarstedt Inc Screw Cap Micro Tube 2 ml 1 2 2 Abgene™ 96 Well 0.8 ml Polypropylene Deepwell Storage Plate 1 1 1 -
Reagents quantities:
Reagents X24 samples X48 samples X96 samples 80% ethanol 16.5 ml 28 ml 51 ml AMPure XP Beads 3.7 ml 5.4 ml 8.9 ml NEBNext Companion Module for Oxford Nanopore Technologies Ligation Sequencing (Cat# E7180S)
regarding the reagents below2 tubes 3 tubes 5 tubes Alternatively: - - - NEBNext FFPE DNA Repair Buffer 1 tube 1 tube 1 tube NEBNext FFPE DNA Repair Mix 1 tube 1 tube 2 tubes Ultra II End-prep Reaction Buffer 1 tube 2 tubes 3 tubes Ultra II End-prep Enzyme Mix 1 tube 1 tube 2 tubes Note: Dead volumes are included.
-
Prepare the NEBNext FFPE DNA Repair Mix and NEBNext Ultra II End Repair / dA-tailing Module reagents in accordance with manufacturer’s instructions, and place on ice.
For optimal performance, NEB recommend the following:
Thaw all reagents on ice.
Flick and/or invert the reagent tubes to ensure they are well mixed.
Note: Do not vortex the FFPE DNA Repair Mix or Ultra II End Prep Enzyme Mix.Always spin down tubes before opening for the first time each day.
The Ultra II End Prep Buffer and FFPE DNA Repair Buffer may have a little precipitate. Allow the mixture to come to room temperature and pipette the buffer up and down several times to break up the precipitate, followed by vortexing the tube for 30 seconds to solubilise any precipitate.
Note: It is important the buffers are mixed well by vortexing.The FFPE DNA Repair Buffer may have a yellow tinge and is fine to use if yellow.
-
Prepare each DNA sample per well with nuclease-free water in the input plate.
- Per sample, transfer 1 μg (or 100-200 fmol) of genomic DNA into a well of the input plate
- Adjust the volume to 48 μl with nuclease-free water
- Mix thoroughly by pipetting
- Spin down briefly in a microfuge
-
Quantify 1 µl of each eluted sample using a Qubit fluorometer plate reader off deck.
-
Switch on the Hamilton NGS STAR 96 robot and open 'Hamilton Run Control' on the computer by clicking the icon:
-
Click 'File' and 'Open' to choose the method to run on the liquid handling robot.
-
Click 'Process01: DNA repair and end-prep' to start.
-
Click 'Process02: DNA repair and end-prep clean-up' to stop the automated library preparation and quantify the samples before the adapter ligation step.
-
Click 'Browse' to choose the Input File Worklist for the specific number of samples in the run and click 'OK'.
An input file worklist for the number of samples in the run must be generated before the run.
Example of an input file worklist:Source_SampleIDSource_WellTarget_WellSample_01 A1 A1 Sample_02 B1 B1 Sample_03 C1 C1 -
Prepare the End Prep Mastermix with the following reagents according to the Hamilton user interface. Click either 'Yes' or 'No' to continue.
Note: It is user preference whether to print and save the instructions.
Reagent volumes for all sample numbers:
Reagent Volume X24 samples Volume X48 samples Volume X96 samples NEBNext FFPE DNA Repair Buffer 106.6 µl 213.3 µl 414.8 µl NEBNext FFPE DNA Repair Mix 60.9 µl 121.8 µl 237 µl Ultra II End-prep Reaction Buffer 106.6 µl 213.3 µl 414.8 µl Ultra II End-prep Enzyme Mix 91.4 µl 182.8 µl 355.6 µl Total 365.6 µl 731.2 µl 1422.2 µl 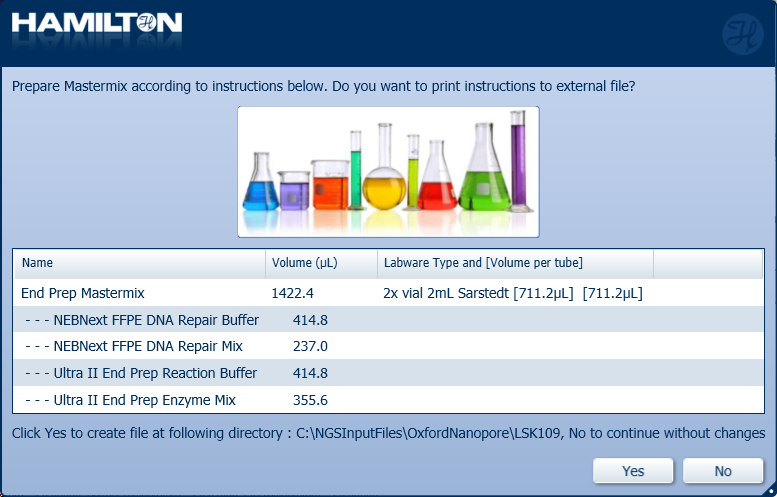
-
Insert the ComfortLid position as displayed on screen. Click 'Ok' to continue.
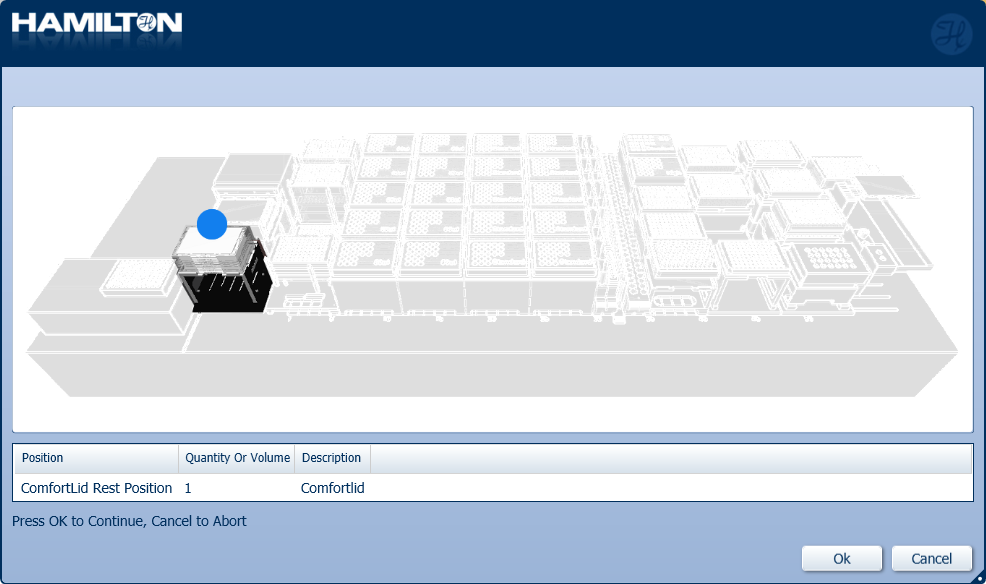
-
Insert plates to their corresponding positions. Click 'Ok' to continue.
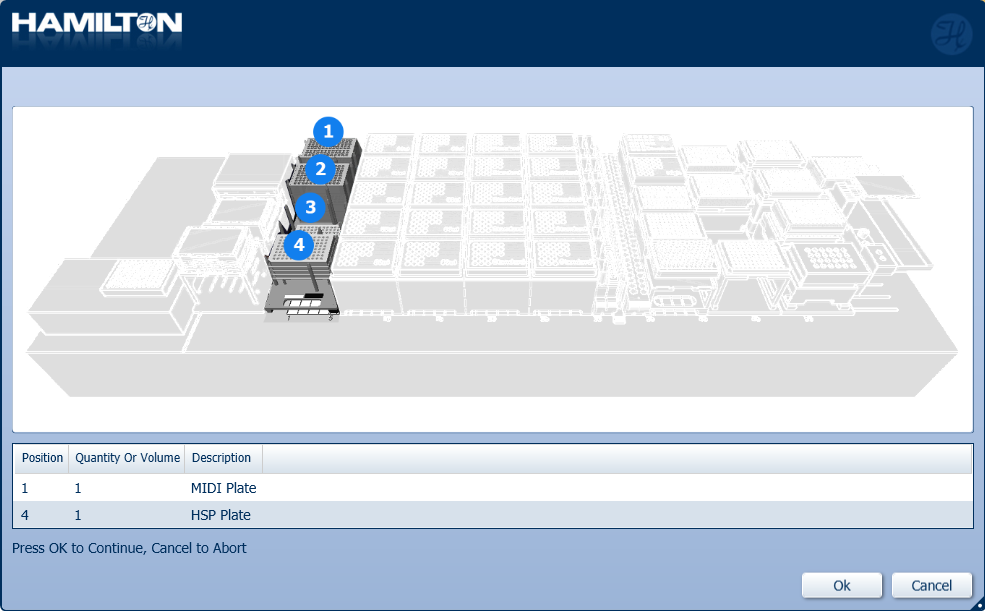
-
Load a full deck of 50 µl tips into the positions on screen. Click 'Ok' to continue.
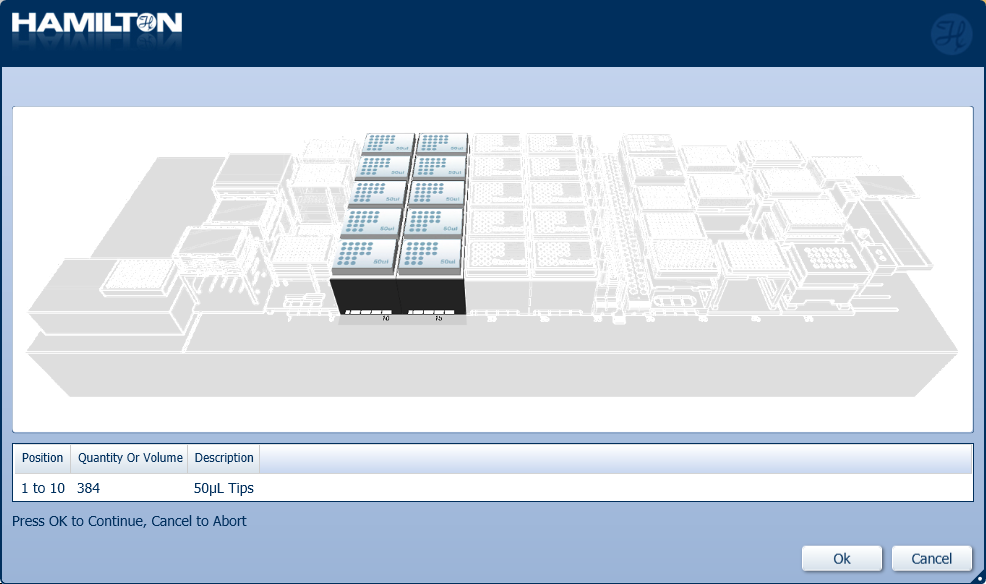
-
Highlight the 50 µl tips available to use on the 'Edit Tip Count' window and click 'Ok' to continue.
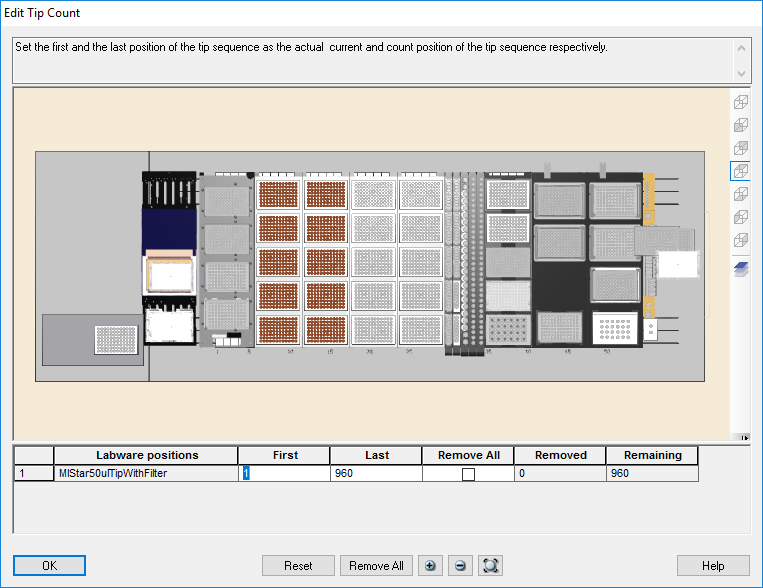
-
Load a full deck of 300 µl tips in the positions on screen. Click 'Ok' to continue.
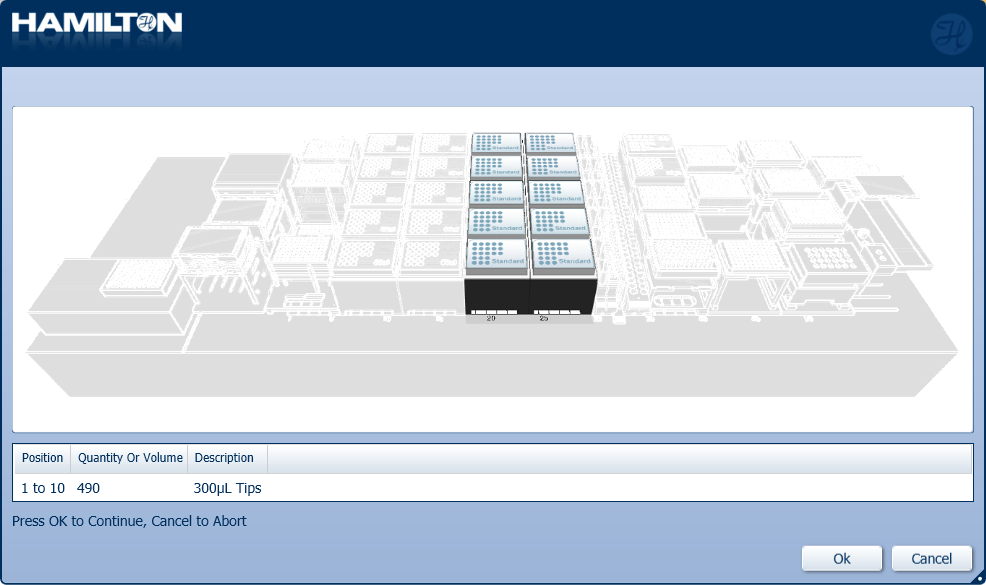
-
Highlight the 300 µl tips available to use on the 'Edit Tip Count' window and click 'Ok' to continue.
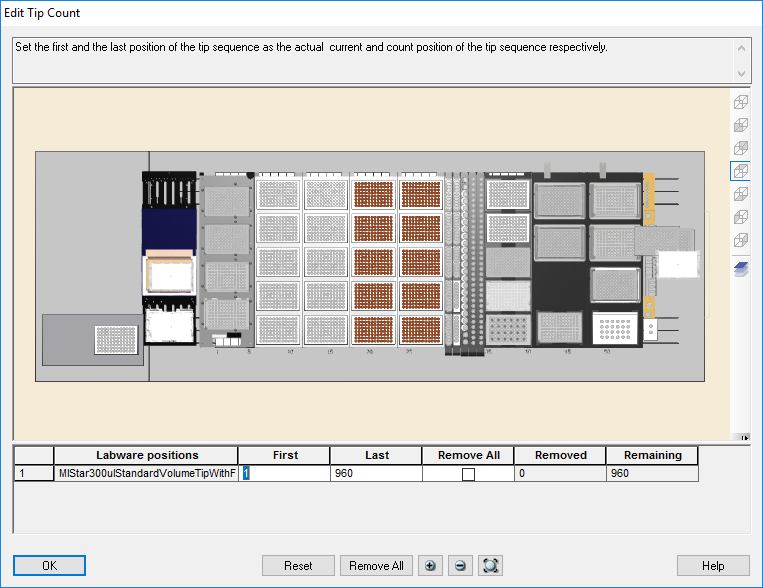
-
Freshly prepare 80% ethanol in nuclease-free water in a trough.
Reagents Volume X24 samples Volume X48 samples Volume X96 samples 80% ethanol 16.5 ml 28 ml 51 ml -
Insert the trough of 80% ethanol in the position on screen and click 'Ok' to continue.
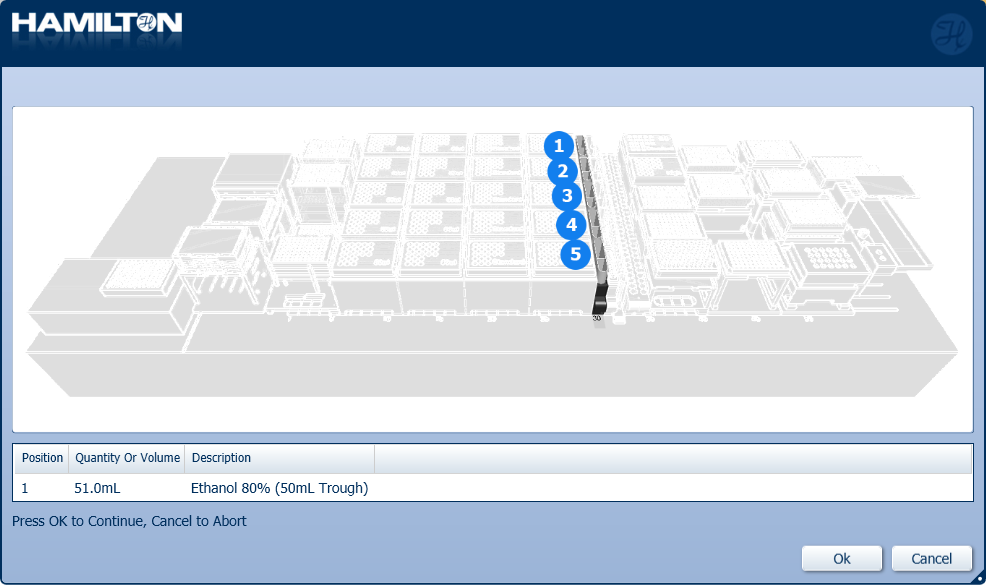
-
Prepare the AMPure XP beads by vortexing and load the 20 ml trough with the volume required:
Reagents Volume X24 samples Volume X48 samples Volume X96 samples Beads 3.7 ml 5.4 ml 8.9 ml -
Insert the trough of AMPure XP beads and nuclease-free water in their positions on screen. Click 'Ok' to continue.
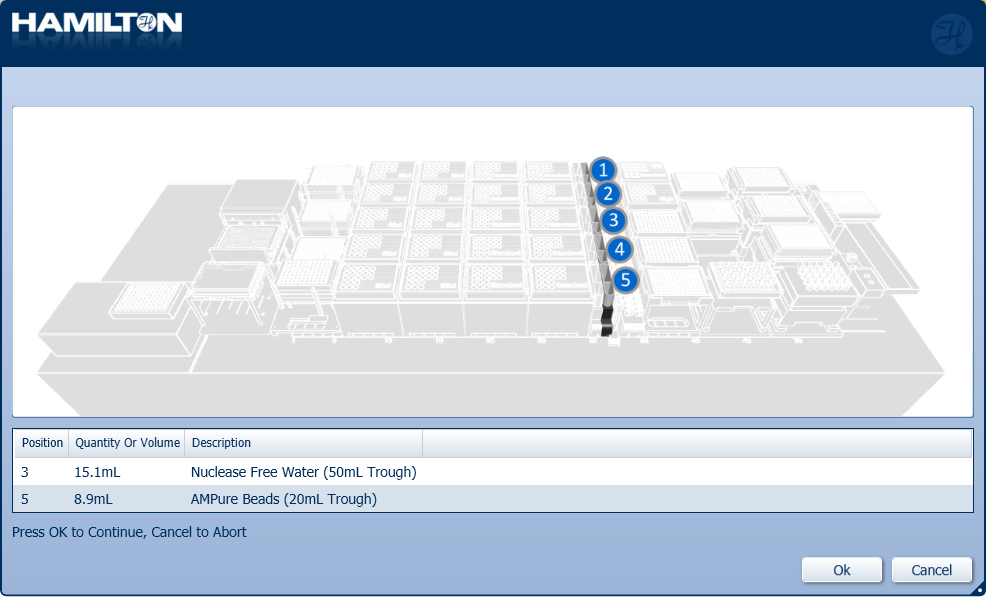
-
Load 1000 µl tips and insert the input plate of DNA samples into the position on screen. Click 'Ok' to continue.
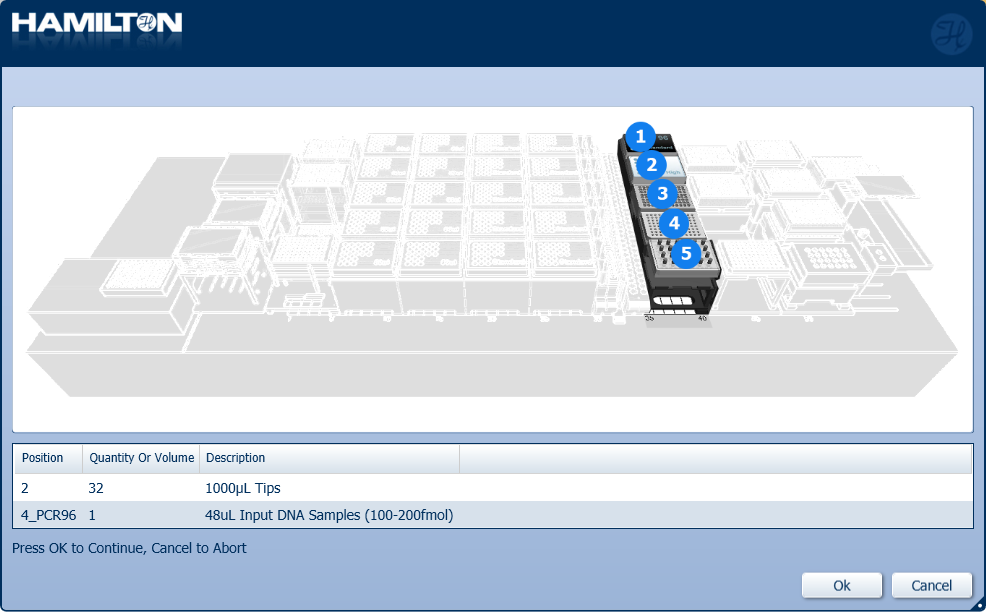
-
Highlight the 1000 µl tips available to use on the 'Edit Tip Count' window and click 'Ok' to continue.
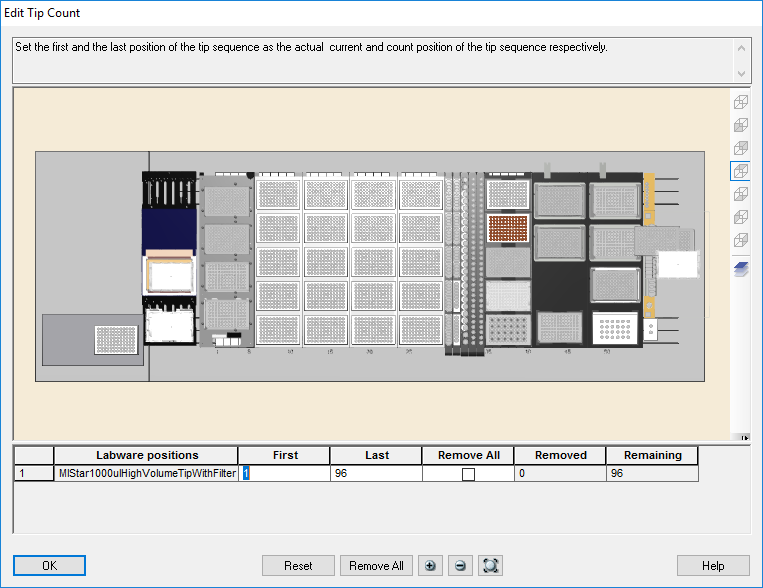
-
Mix and insert the prepared End Prep Mastermix into the positions on screen.
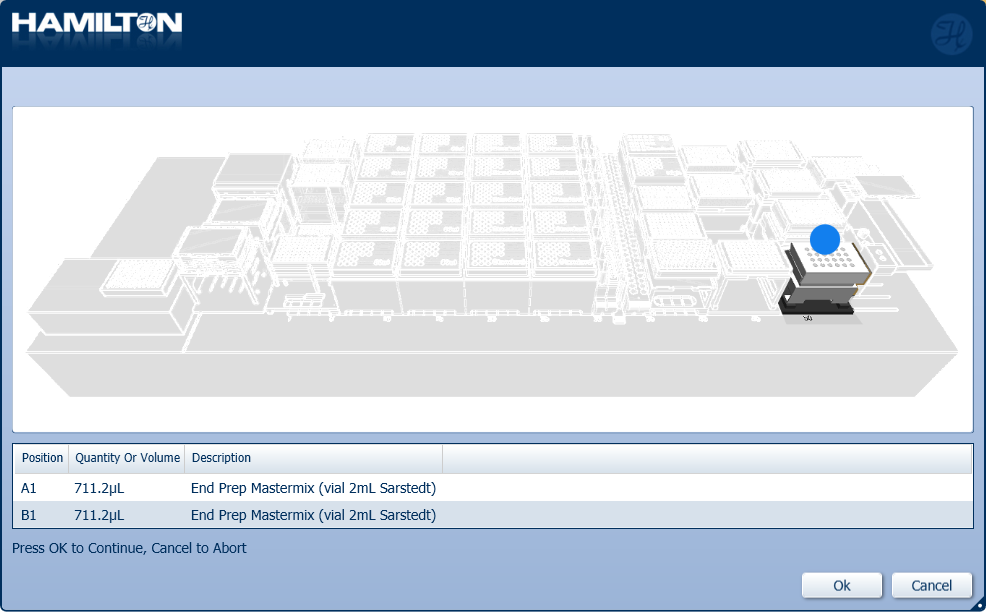
-
Click 'Ok' to start the DNA repair and end-prep automation process.
-
Once the automation process has finished, there will be an on screen prompt to unload the plate. Click 'Ok' to continue.

-
Quantify 1 µl of each eluted sample using a Qubit fluorometer plate reader off deck.

















