-
Basecalling Kit 14 simplex data
Simplex basecalling is where the single template DNA strand passes through the nanopore and is basecalled with sequencing accuracies of up to 99% when using our super-accurate basecaller (SUP). We recommend basecalling Kit 14 simplex data on MinKNOW.
For optimal performance on our devices, please see the specifications below:
- 64 bit Linux or Windows 10
- Intel i7, i9, Xeon, or better processor
- At least 16 GB of RAM
- An NVIDIA GPU, at least RTX 2070 or better, with at least 16 GB of GPU memory
- At least 1 TB SSD
Make sure you are using the most recent version of MinKNOW.
Below are our basecaller and output recommendations for setting up a Kit 14 sequencing run on MinKNOW to generate POD5 files to use as input for duplex basecalling on Dorado.
-
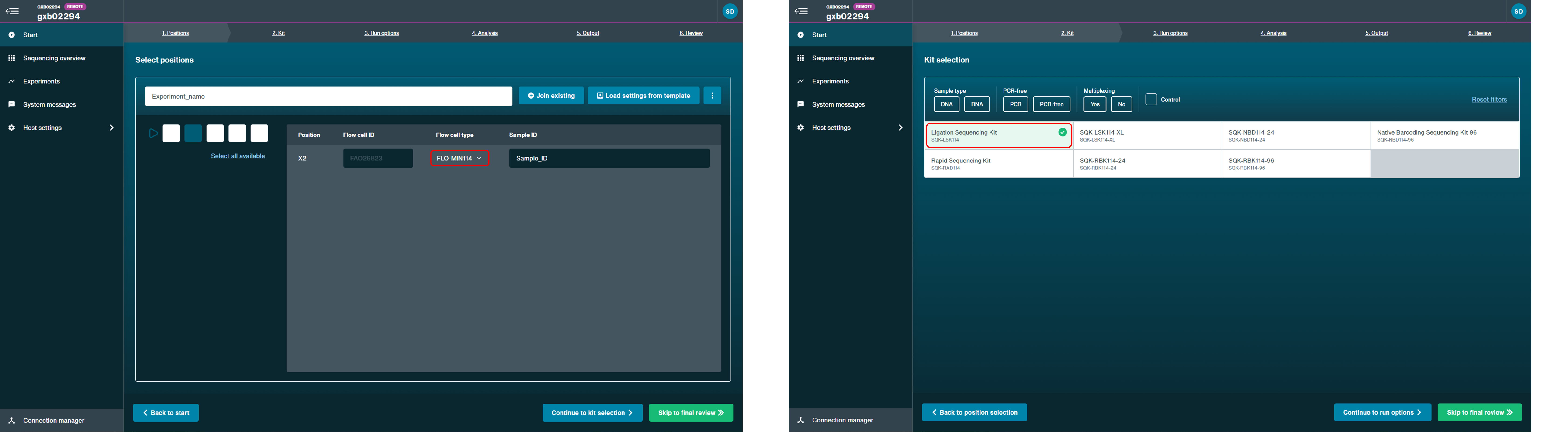
When setting up a sequencing run, select the R10.4.1 flow cell (FLO-MIN114 or FLO-PRO114M) and choose the Kit 14 used (e.g. SQK-LSK114) for your library prep.
For detailed instructions on setting up a run in MinKNOW, please see the "Starting a sequencing run" in the MinKNOW protocol.
-
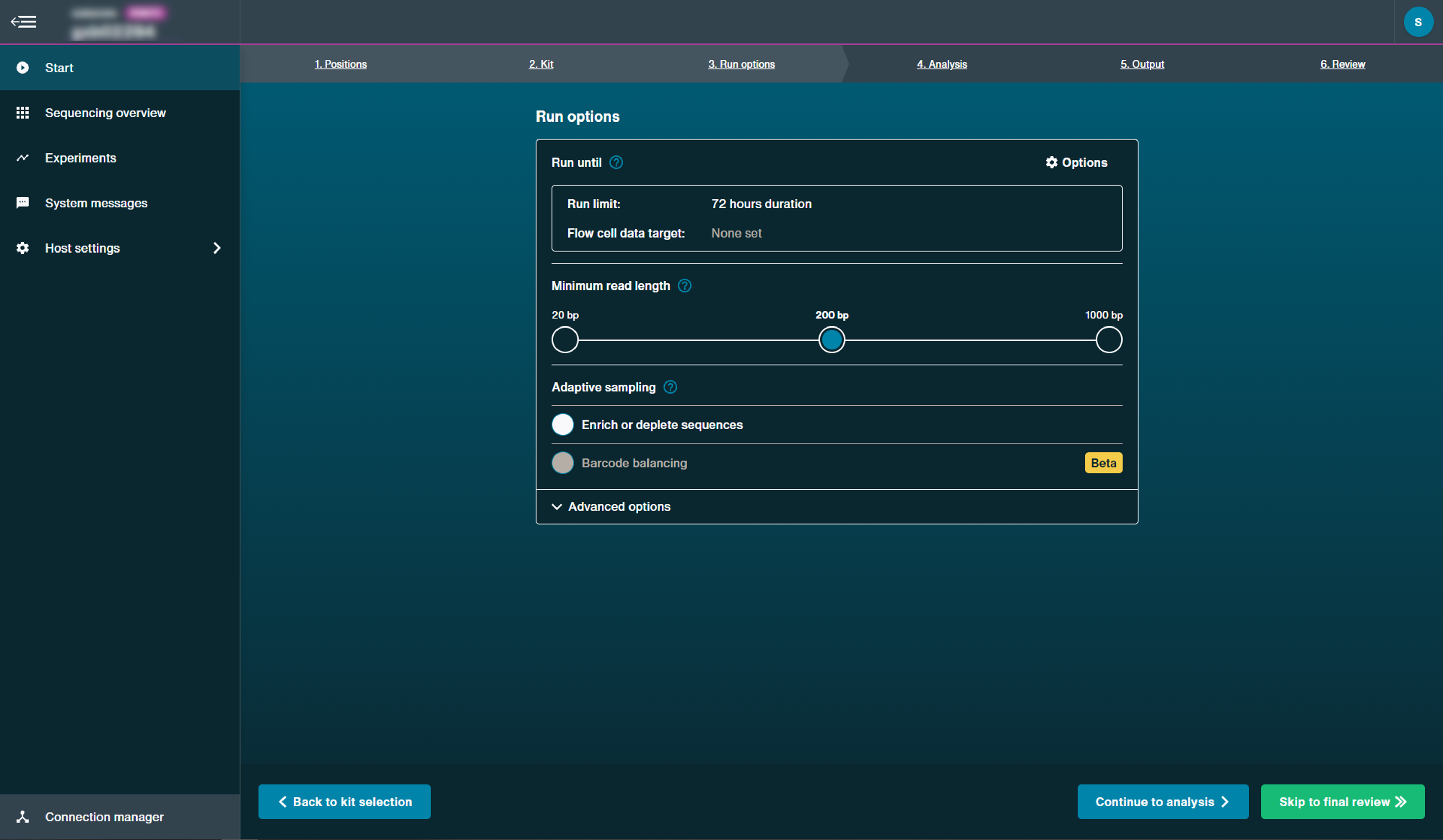
The parameters on the run options tab can be left to their default settings.
-
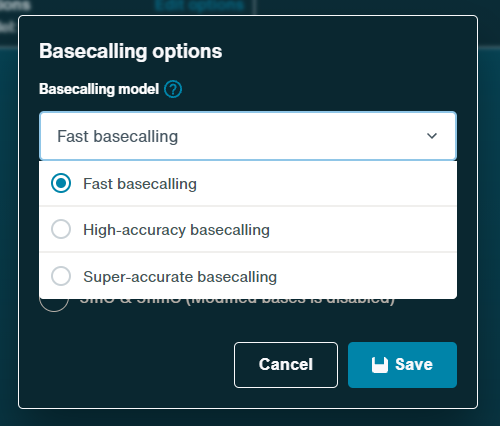
On the analysis tab, we recommend using the Fast basecaller.
Other basecallers are compatible but the Fast basecaller will be quicker during the sequencing run. The High-accuracy (HAC) and Super-accurate (SUP) basecallers can be used when rebasecalling your data on Dorado.
-
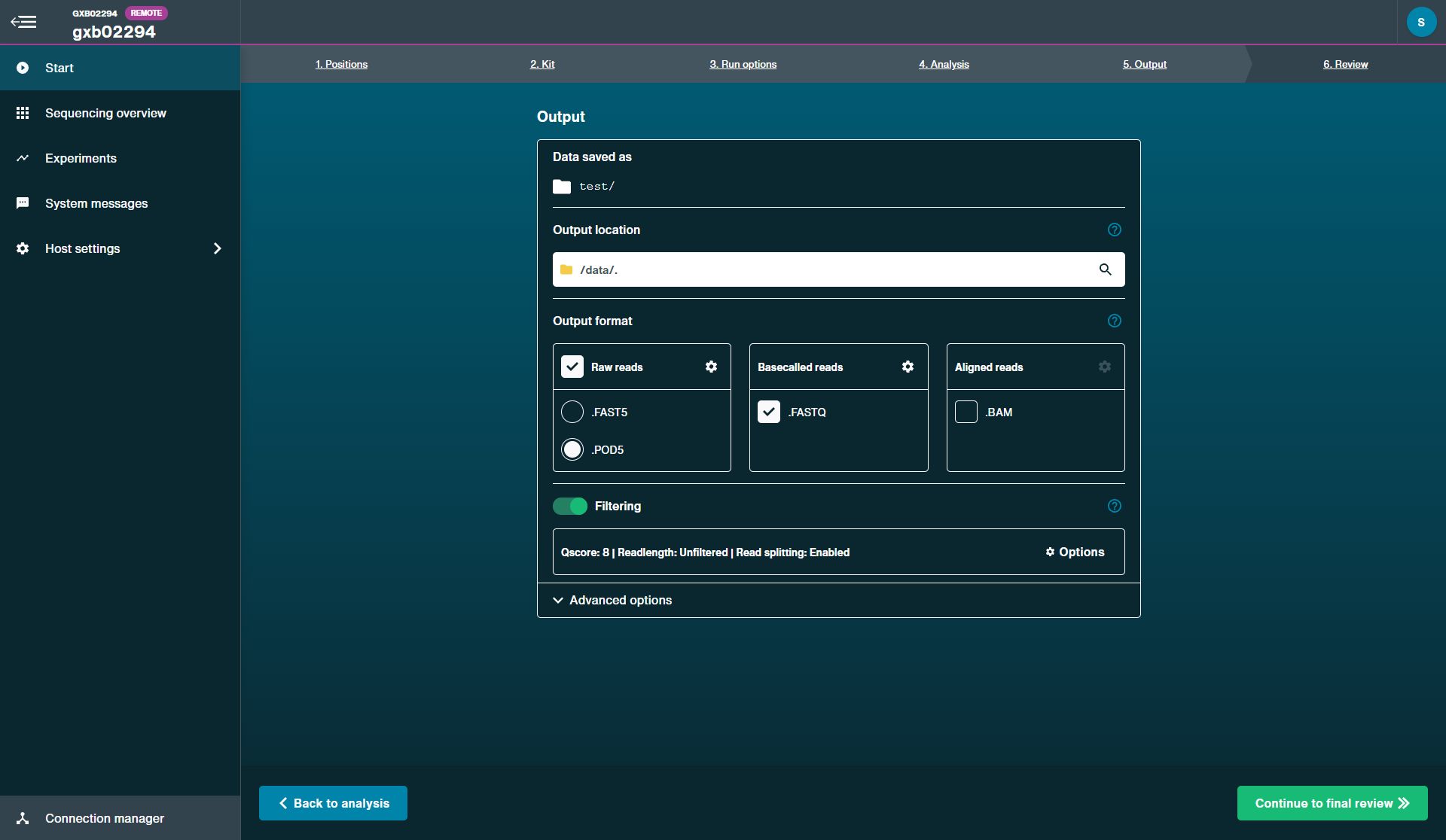
On the output tab, ensure POD5 is selected as the output format and read splitting is on by default.
For more information about read splitting, please see the section Introduction to read splitting in the appendix.
FAST5 files can be used as an output file but POD5 files are required for optimal performance on Dorado.