-
Introduction to DNA quality control
It is important that you check your input DNA for quality before beginning library preparation. Low molecular weight, incorrectly quantified and/or contaminated DNA (e.g. salt, EDTA, protein, organic solvents) can have a significant impact on downstream processes and ultimately, your sequencing runs. Below are some guidelines for how to check the DNA quality to ensure the highest possible throughput.
Access to laboratory equipment is not always possible in field conditions, so we recommend that you optimise extraction and purification in the laboratory before doing fieldwork.
To obtain similar results to those generated in the Lambda Control protocol, use high quality, pure genomic DNA (gDNA). We have had good results with DNA which meets the following criteria:
- Purity as measured using Nanodrop – OD 260/280 of 1.8 and OD 260/230 of 2.0–2.2
- Average fragment size, as measured by pulsed-field gel analysis ( >30 kb)
- Input mass, as measured by Qubit – 1 µg
For long-term storage of high molecular weight (HMW) gDNA we recommend the use of TE buffer.
There are established methods for generating high quality HMW gDNA which should be used when preparing gDNA for sequencing on the MinION Mk 1B. Several of these methods are described in detail in Sambrook J, and Russell DW, (2001) Molecular Cloning: a Laboratory Manual. 3 rd Edition, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York.
-
Correct quantification
The majority of RNA should be removed by RNase digestion. We have found RiboShredder (Epicentre RS 12500) to be particularly effective. However, since Riboshredder is being discontinued, you can use the RNase Cocktail Enzyme Mix (ThermoFisher, AM2286) instead.
Please be aware that certain RNase treatments can lead to digestion of DNA, as well as RNA.We recommend that the DNA stock is quantified using Qubit analysis. A Qubit measures DNA specifically. Even after RNase digestion, residual RNA is a common contaminant in gDNA preparations and is not well identified by Nanodrop measurements. Incorrect quantification could mean that you will proceed with less DNA than intended, resulting in poor performance. Also, contamination from bases (dNTPs and NTPs) will interfere with Nanodrop measurements. Therefore, we recommend that Qubit is used for all quantification measurements (i.e. after all clean-up steps).
Additionally, high concentration, high molecular weight DNA preparations (and those with heavy RNA contamination) can lack homogeneity, which will give rise to inaccurate quantification. If you encounter this with your RNase-treated DNA sample, we recommend that you dilute the DNA further with TE, and that you rotate the tube gently until the suspension is homogeneous. Vortexing the DNA or pipetting up and down will cause shearing, which will limit the fragment sizes available to the nanopore.
-
Assessing DNA quality
- Chemical impurities such as detergents, denaturants, chelating agents and high concentrations of salts should be avoided as these may affect the efficiency of enzymatic steps. Please refer to the Contaminants document for more details.
- Other contaminants such as single stranded DNA, RNA, proteins and dyes may also reduce the efficiency of steps in the library preparation.
- The quality of DNA may be assessed by Nanodrop (for samples with concentration >20 ng/µl).
- We recommend that sample DNA has a 260/280 ~ 1.80 and a 260/230 ~2.0-2.2.
- A 260/280 which is higher than ~1.8 indicates the presence of RNA.
- A 260/280 which is lower than ~1.8 can indicate the presence of protein or phenol.
- A 260/230 significantly lower than 2.0-2.2 indicates the presence of contaminants, and the DNA may need additional purification.
In the Nanodrop trace shown below, Sample 1 had a 260/230 of ~1.0 and the resulting library performed badly in a sequencing run. If additional purification is not possible, amplification of the library by PCR can be performed to improve library cleanliness.

-
Assessing molecular weight
Nanopore sequencing devices generate reads that reflect the lengths of the fragments loaded into the flow cell. To have control over the size of the fragments generated in the library prep, it is important to begin with high molecular weight (HMW) DNA.
The shearing of HMW DNA can be minimised by:
- Using wide-bore pipette tips to handle the gDNA
- Mixing gently but thoroughly by flicking the tube, as opposed to vortexing or pipetting
- Avoiding unnecessary freeze-thaw cycles
- Avoiding pH <6 and >9
- Avoiding high temperatures, which can lead to degradation
Conventional agarose gels cannot resolve DNA fragments greater than 15–20 kb, but the molecular weight of starting material can be measured by pulsed-field gel analysis.
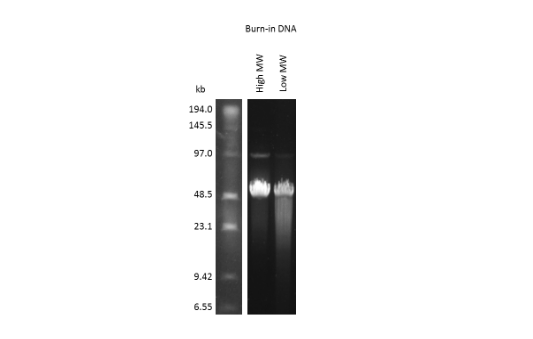
The figure shows two samples of Lambda phage DNA: one of intact high molecular weight fragments and one containing a significant proportion of low molecular weight fragments.
Low % agarose gel analysis can be used to detect substantial degradation/shearing:
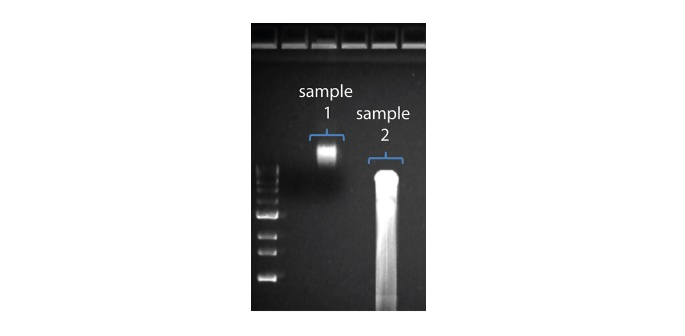
The figure shows two samples of input DNA: Sample #1 is of high molecular weight and sample #2 is of lower molecular weight and has sheared.
-
Assessing fragmentation
Post-fragmentation, the quality of the fragmented material may be assessed by different methods e.g. Agilent Bioanalzyer.

The figure above shows successful (sample 1) and unsuccessful (sample 2) fragmentation, demonstrated by an Agilent Bioanalyzer trace of two DNA samples. Sample #2 contains a substantial proportion of low molecular weight fragments. This is possibly as a result of improper fragmentation, or these low MW fragments may have been present in the input sample.
-
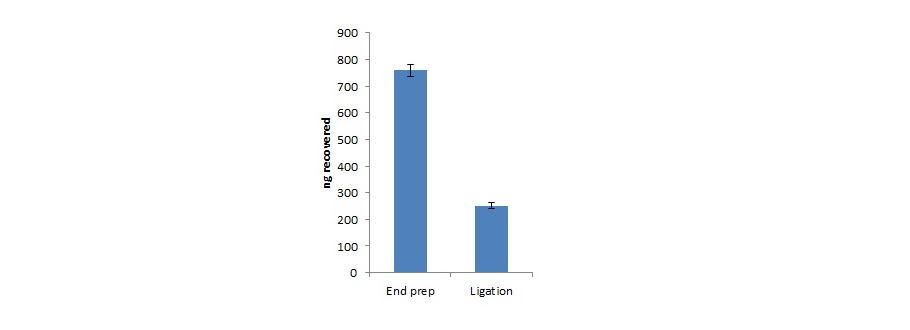
Assessing DNA recovery during library preparation
We recommend performing QC checks at certains points in the library preparation process. This allow users to know how much DNA they have at each stage. Starting with 1 µg (as measured by Qubit), the graph below outlines what should be expected after end repair, dA-tailing and adapter ligation with the Ligation Sequencing Kit. Throughput of a library that follows this quantification is expected to give good throughput.