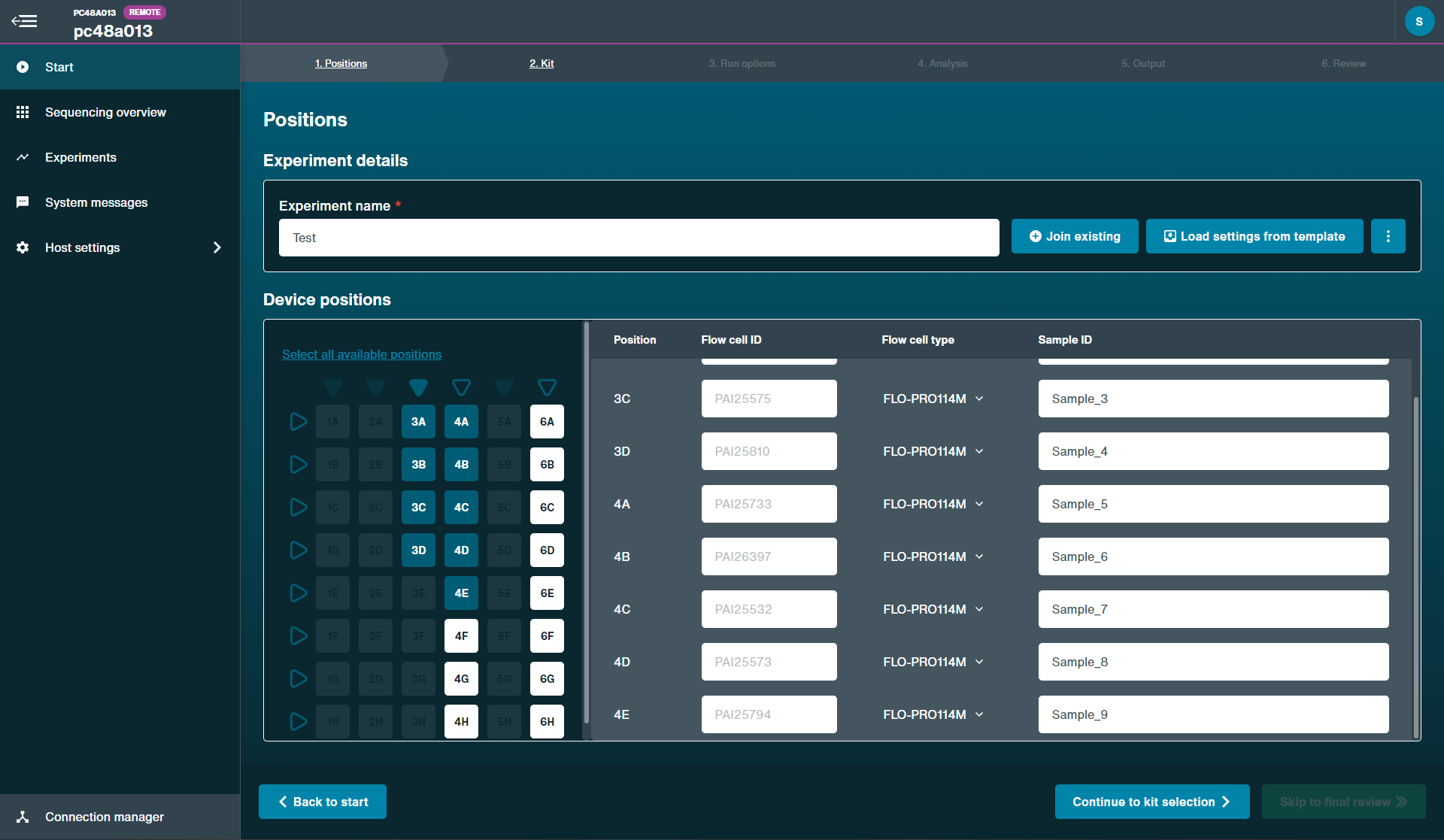
- Materials
-
- Flush Buffer (FB)
- Flush Tether (FLT)
- Loading Beads II (LBII)
- Sequencing Buffer II (SBII)
- Loading Solution (LS)
- Consumables
-
- PromethION Flow Cell
- 1.5 ml Eppendorf DNA LoBind tubes
- 2 ml Eppendorf DNA LoBind tubes
- Equipment
-
- PromethION 2 Solo device
- PromethION 24/48 device
- PromethION Flow Cell Light Shield
- P1000 pipette and tips
- P200 pipette and tips
- P20 pipette and tips
-
Using the Loading Solution
We recommend using the Loading Beads II (LBII) for loading your library onto the flow cell for most sequencing experiments. However, if you have previously used water to load your library, you must use Loading Solution (LS) instead of water.
Note: some customers have noticed that viscous libraries can be loaded more easily when not using Loading Beads II. -
Thaw the Sequencing Buffer II (SBII), Loading Beads II (LBII), Flush Tether (FLT) and on tube of Flush Buffer (FB) at room temperature before mixing the reagents by vortexing and spin down at room temperature.
-
Prepare the flow cell priming mix in a suitable vial for the number of flow cells to flush. Once combined, mix well by briefly vortexing.
Reagent Volume per flow cell Flush Tether (FLT) 30 µl Flush Buffer (FB) 1,170 µl -
For PromethION 2 Solo, load the flow cell(s) as follows:
Place the flow cell flat on the metal plate.
Slide the flow cell into the docking port until the gold pins or green board cannot be seen.
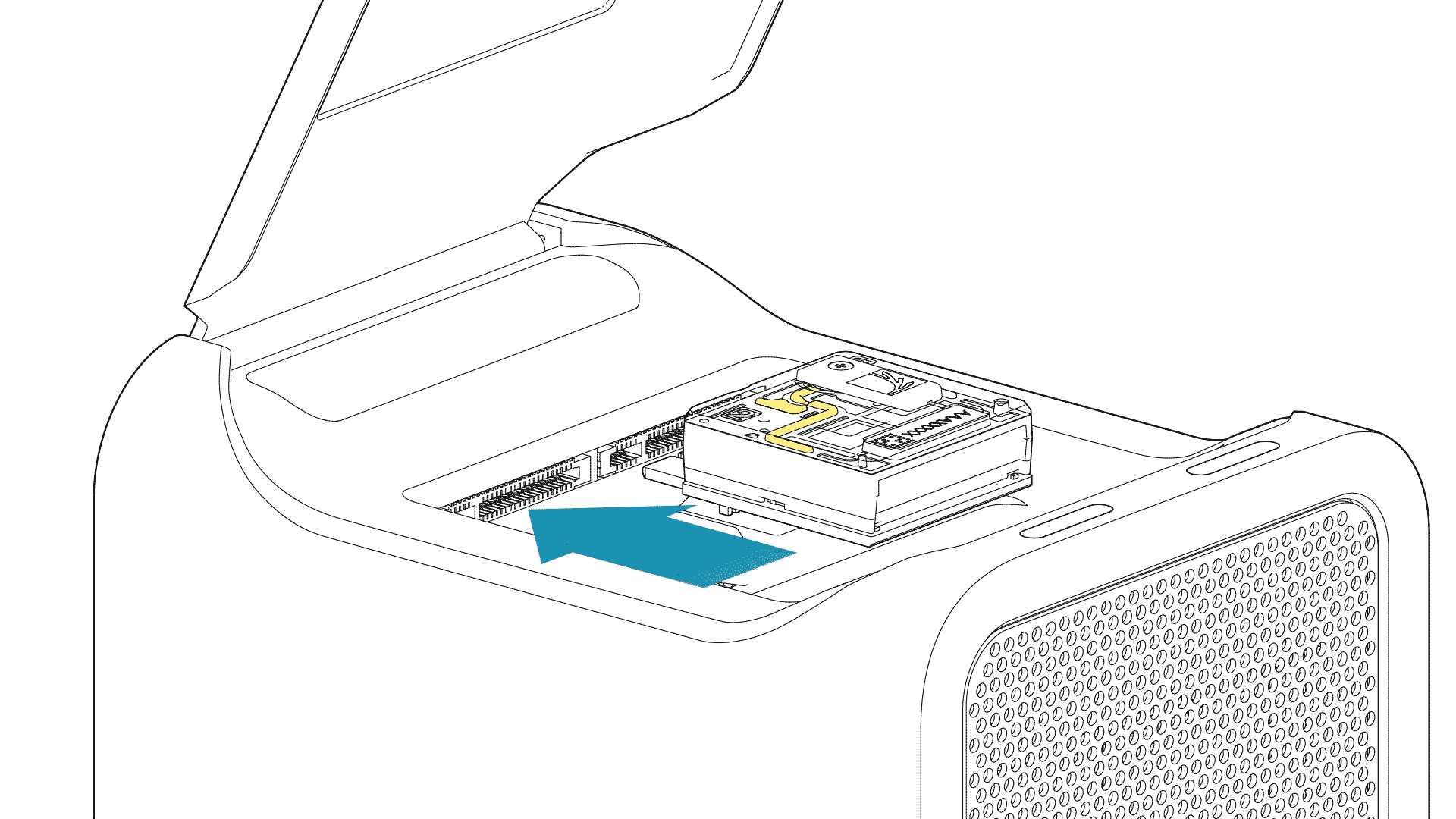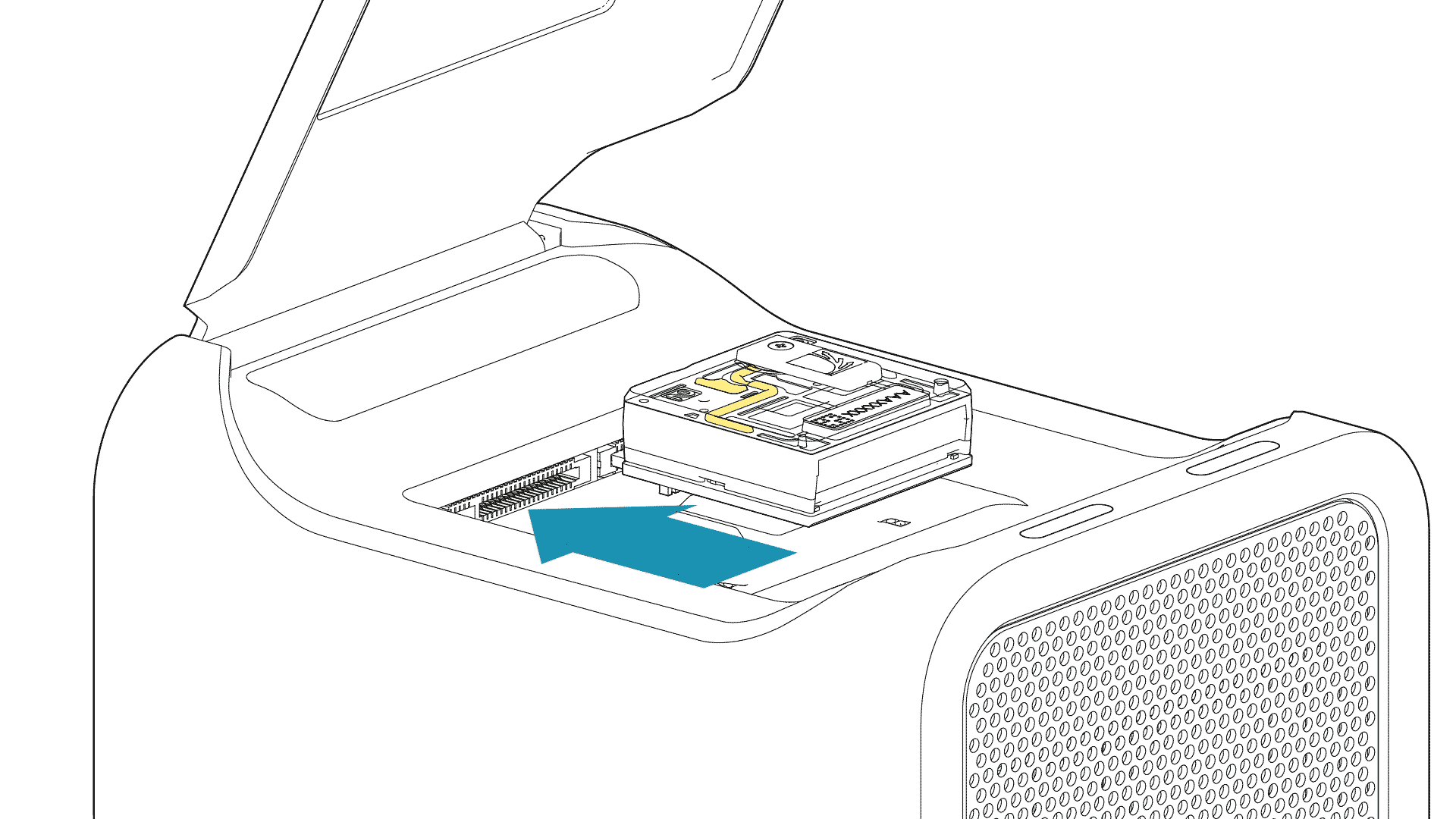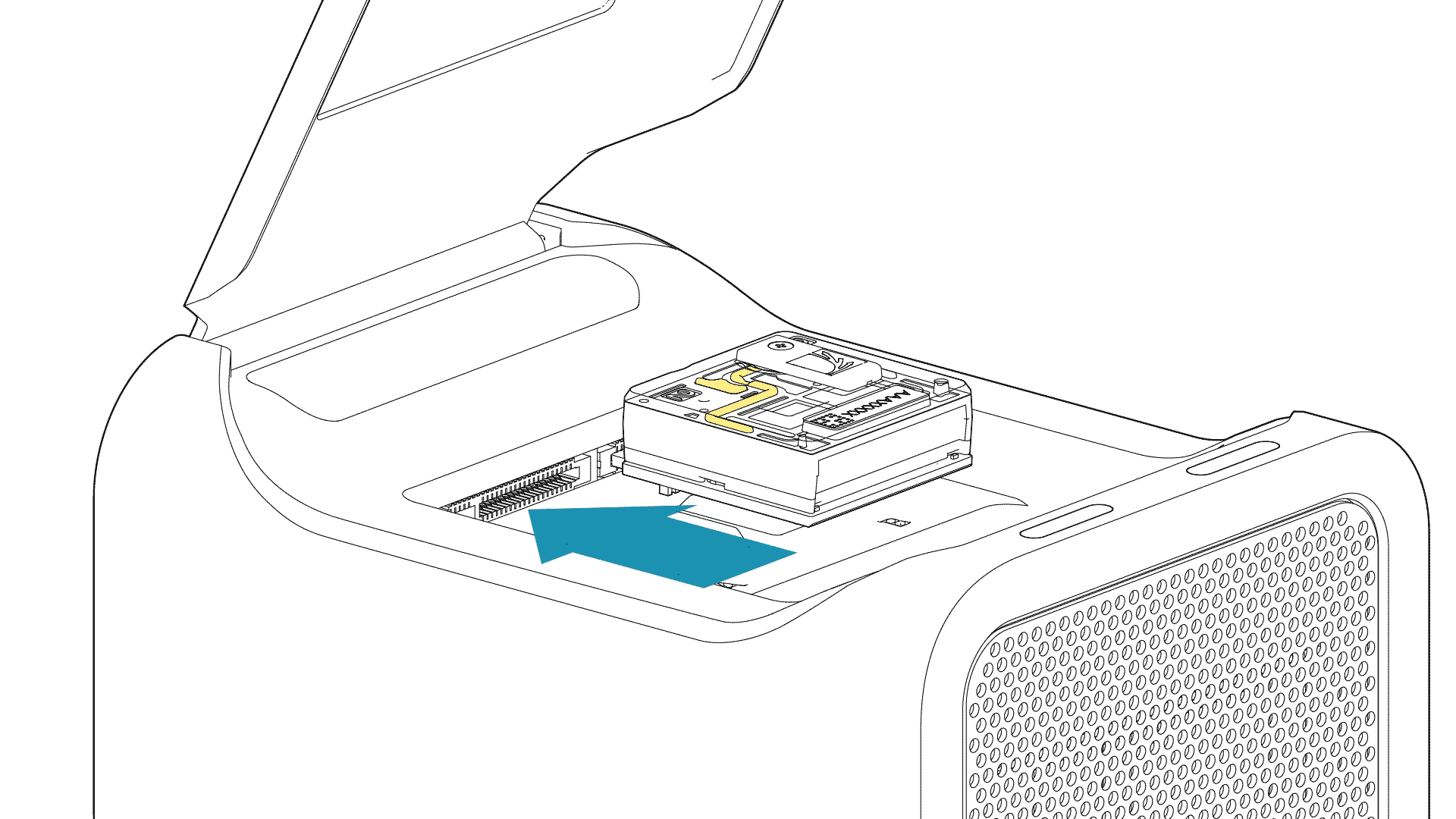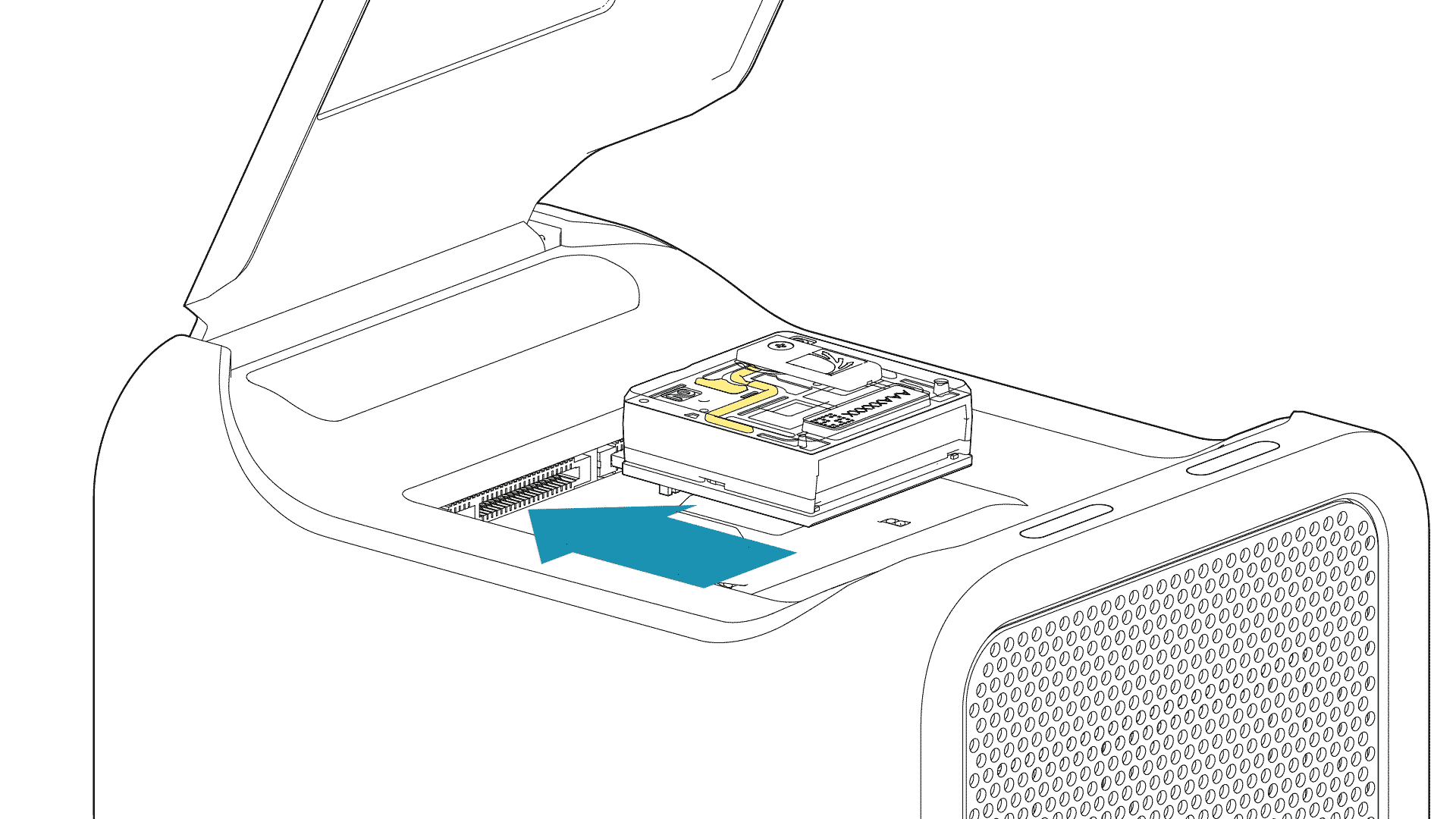
-
For the PromethION 24/48, load the flow cell(s) into the docking ports:
- Line up the flow cell with the connector horizontally and vertically before smoothly inserting into position.
- Press down firmly onto the flow cell and ensure the latch engages and clicks into place.
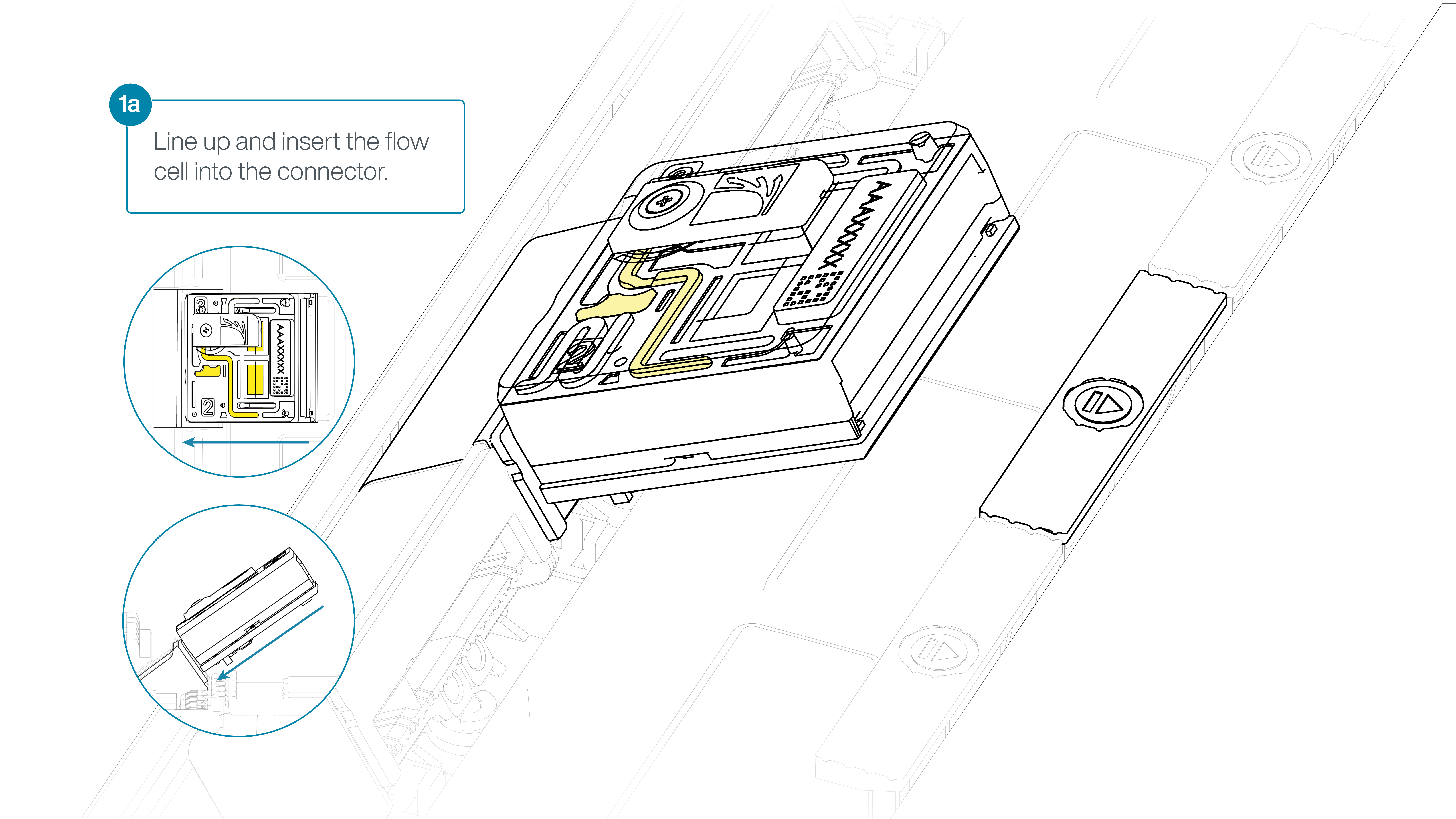
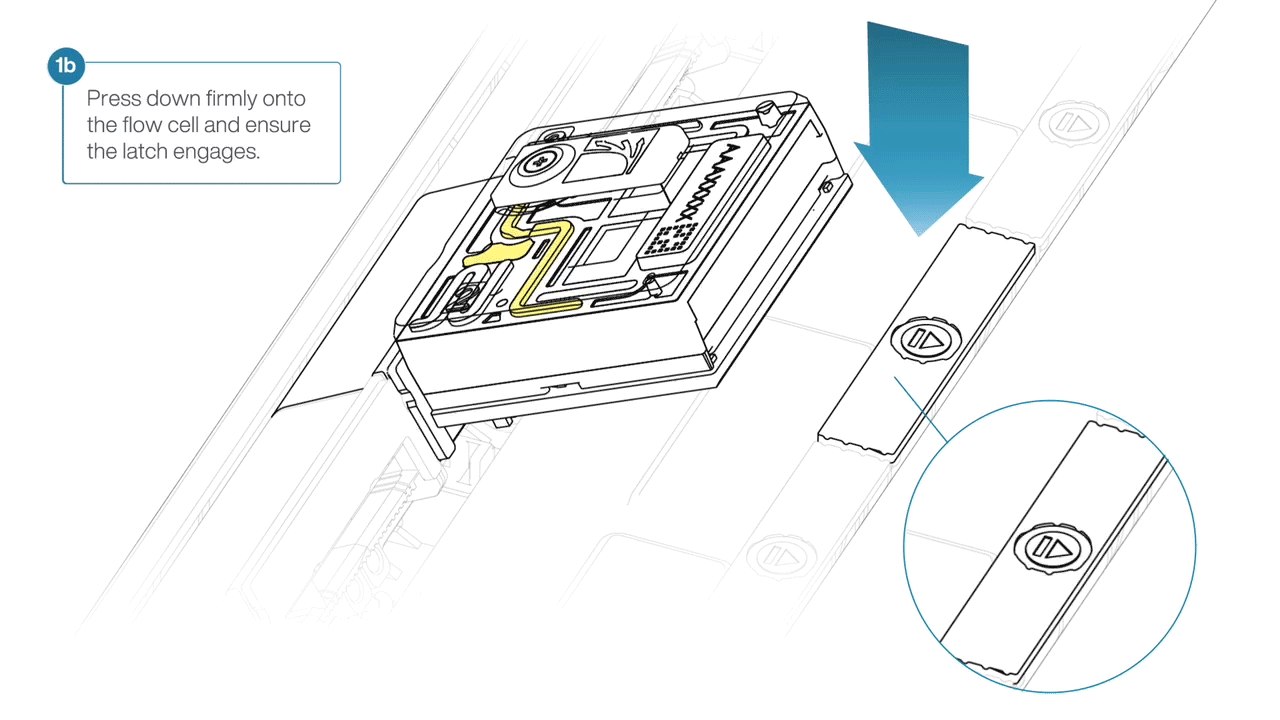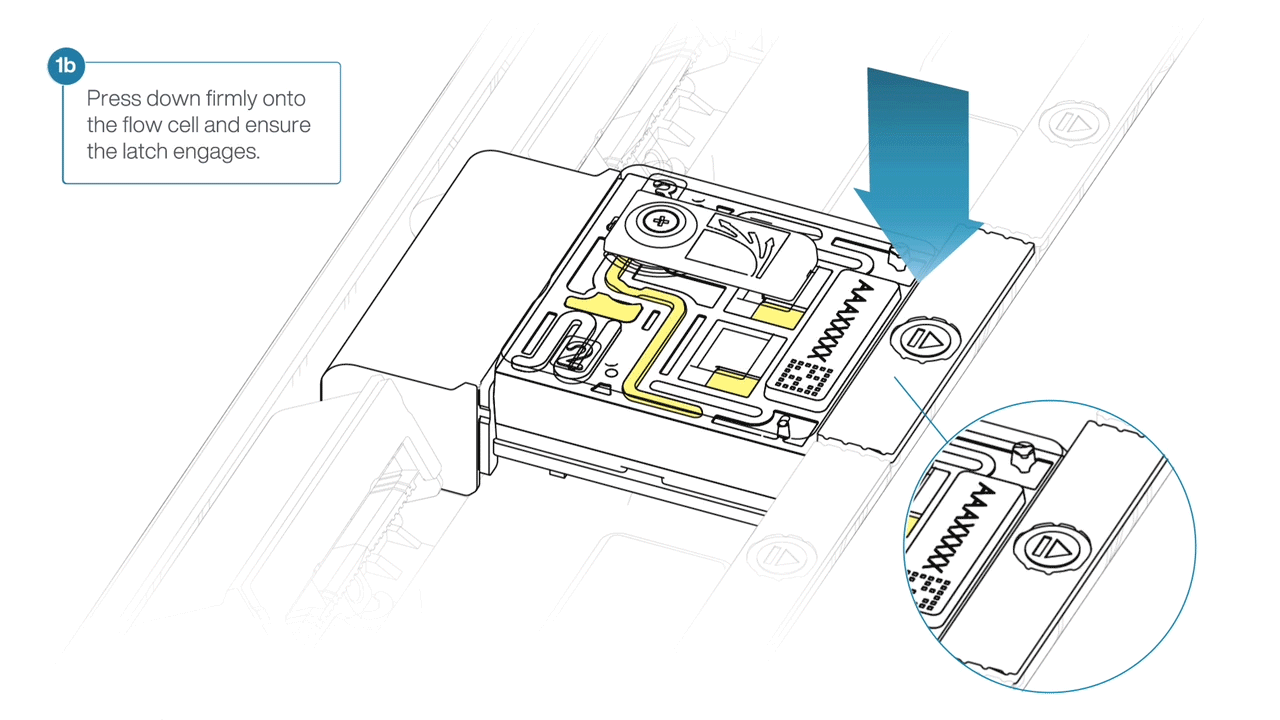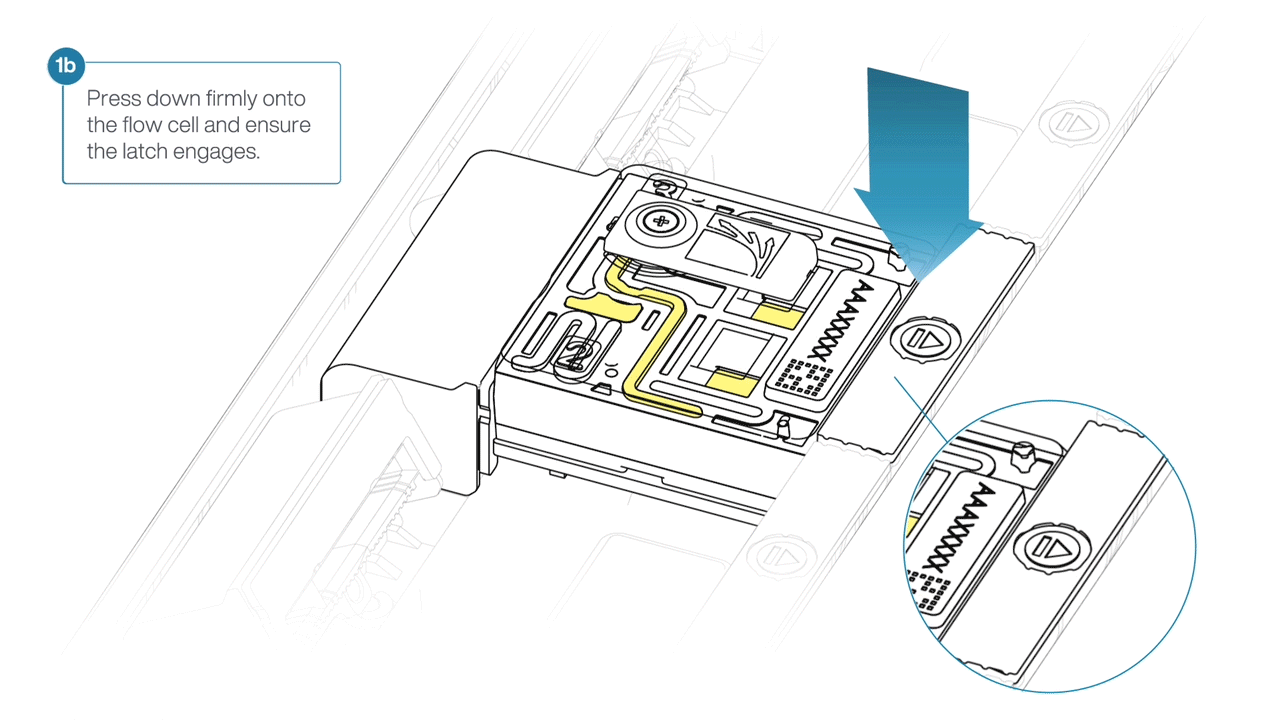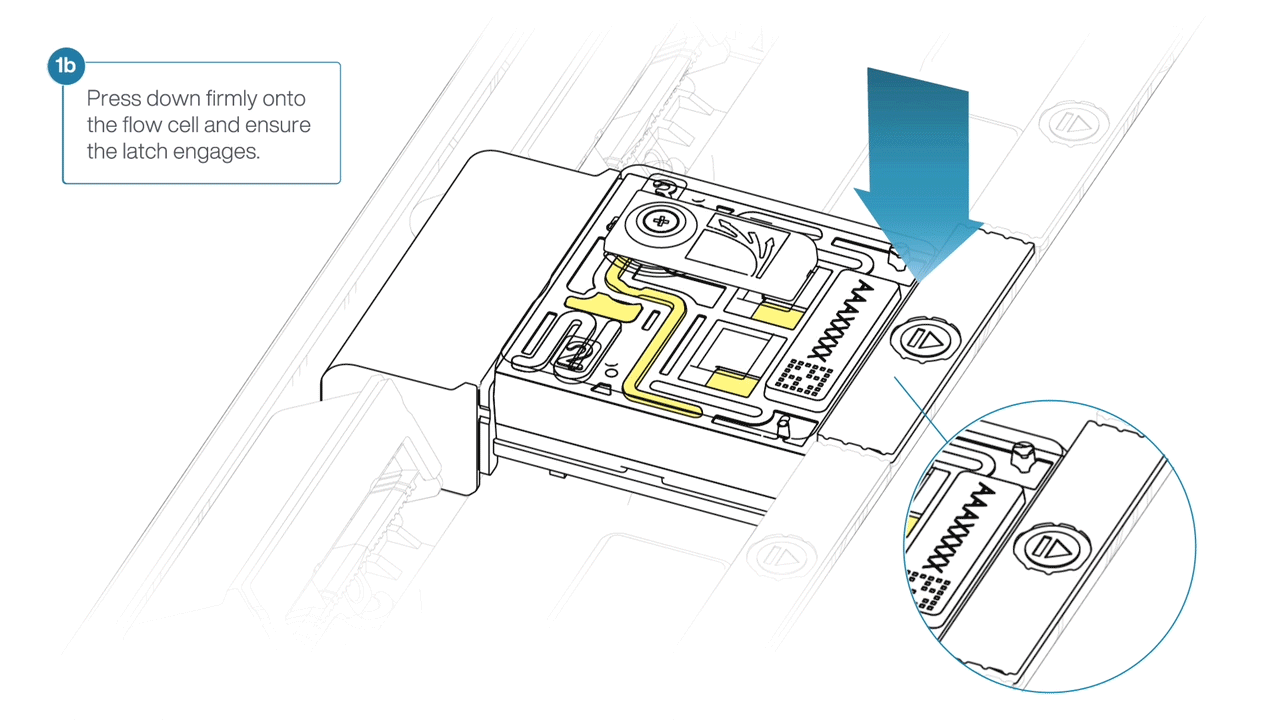
-
If not already completed, perform a flow cell check on all flow cells.
Please refer to the Flow Cell Check protocol for further information.
-
Slide the inlet port cover clockwise to open.
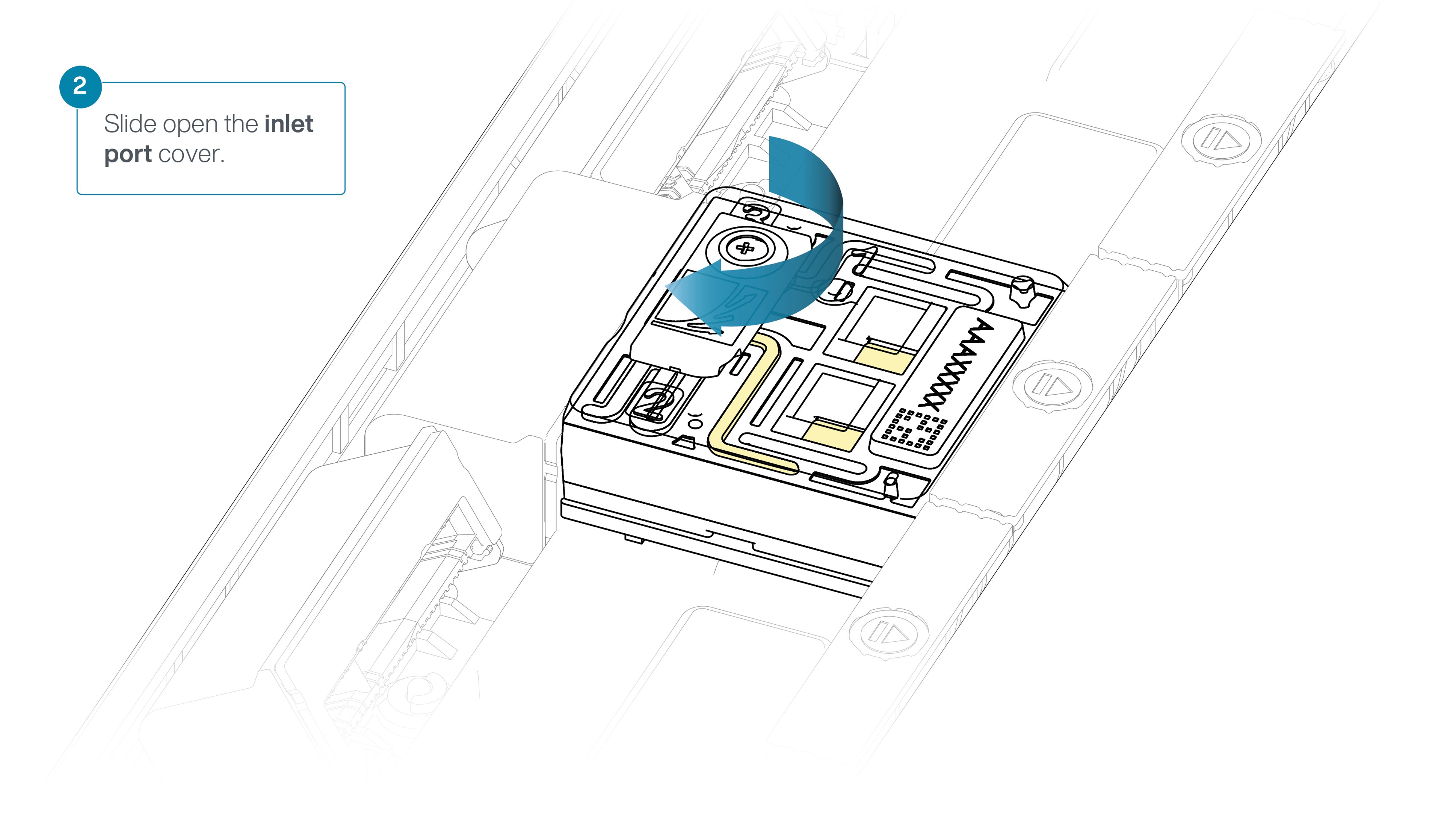
-
After opening the inlet port, draw back a small volume to remove any air bubbles:
- Set a P1000 pipette tip to 200 µl.
- Insert the tip into the inlet port.
- Turn the wheel until the dial shows 220-230 µl, or until you see a small volume of buffer entering the pipette tip.
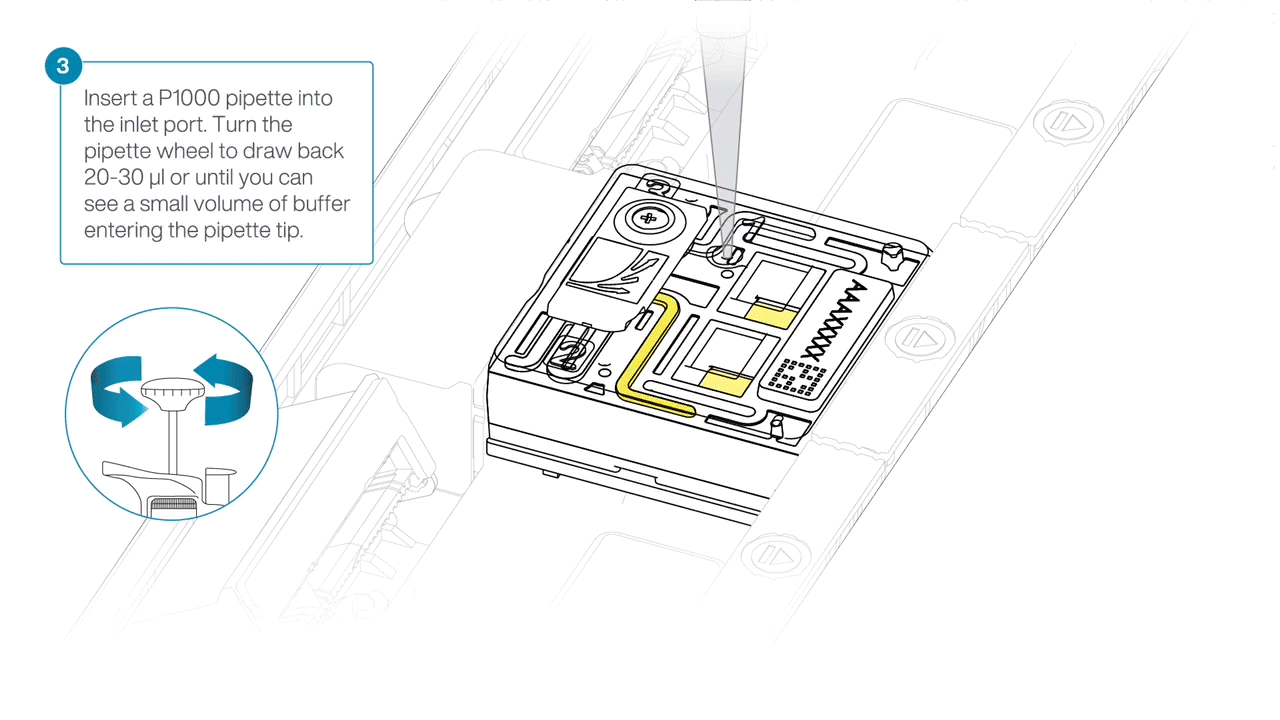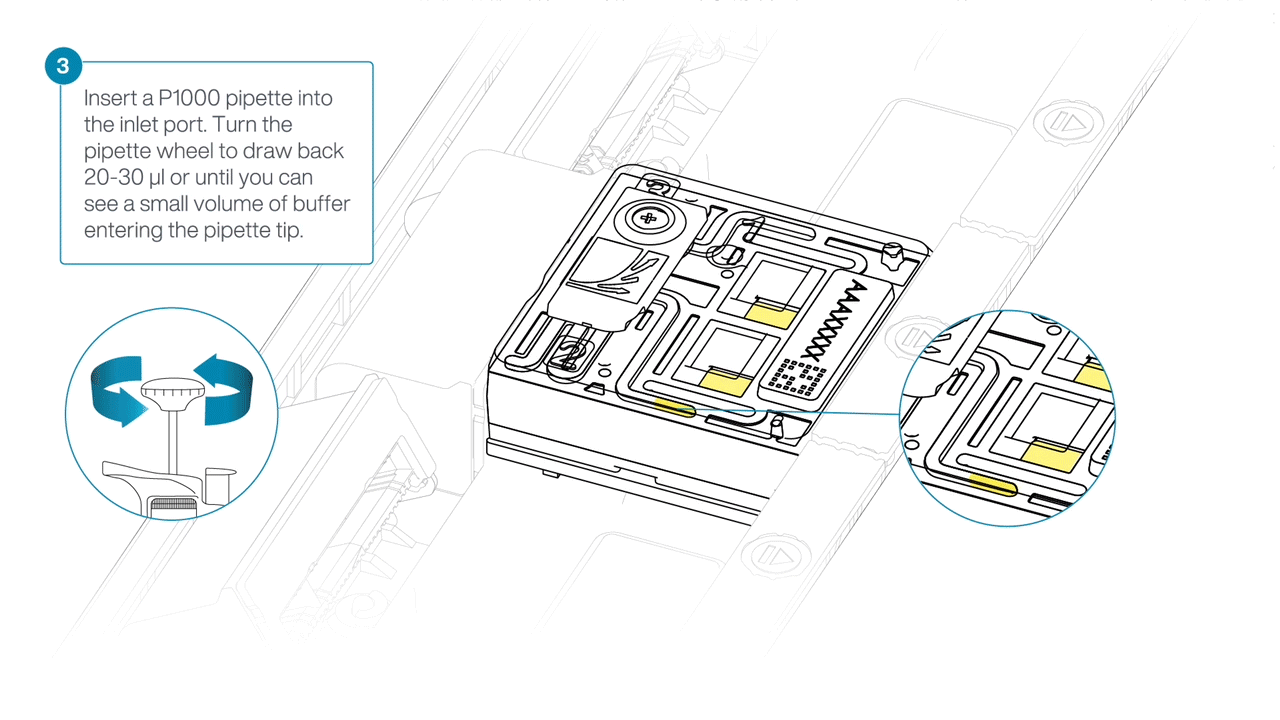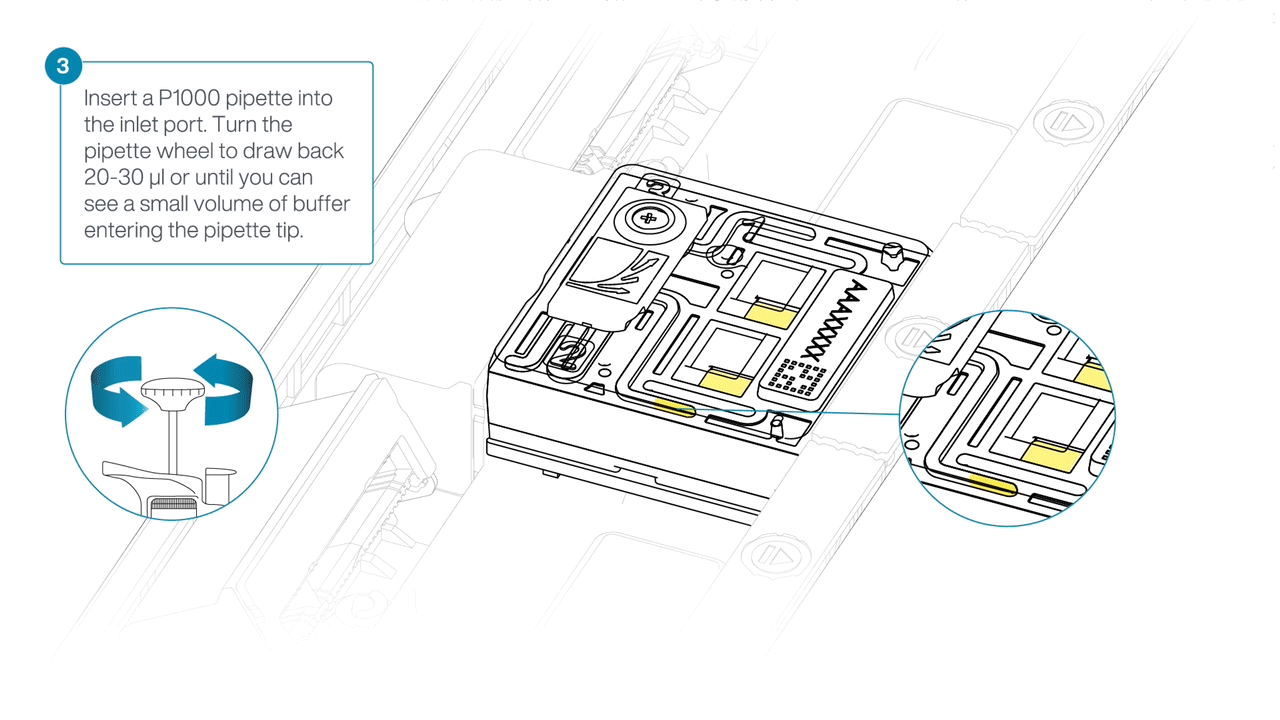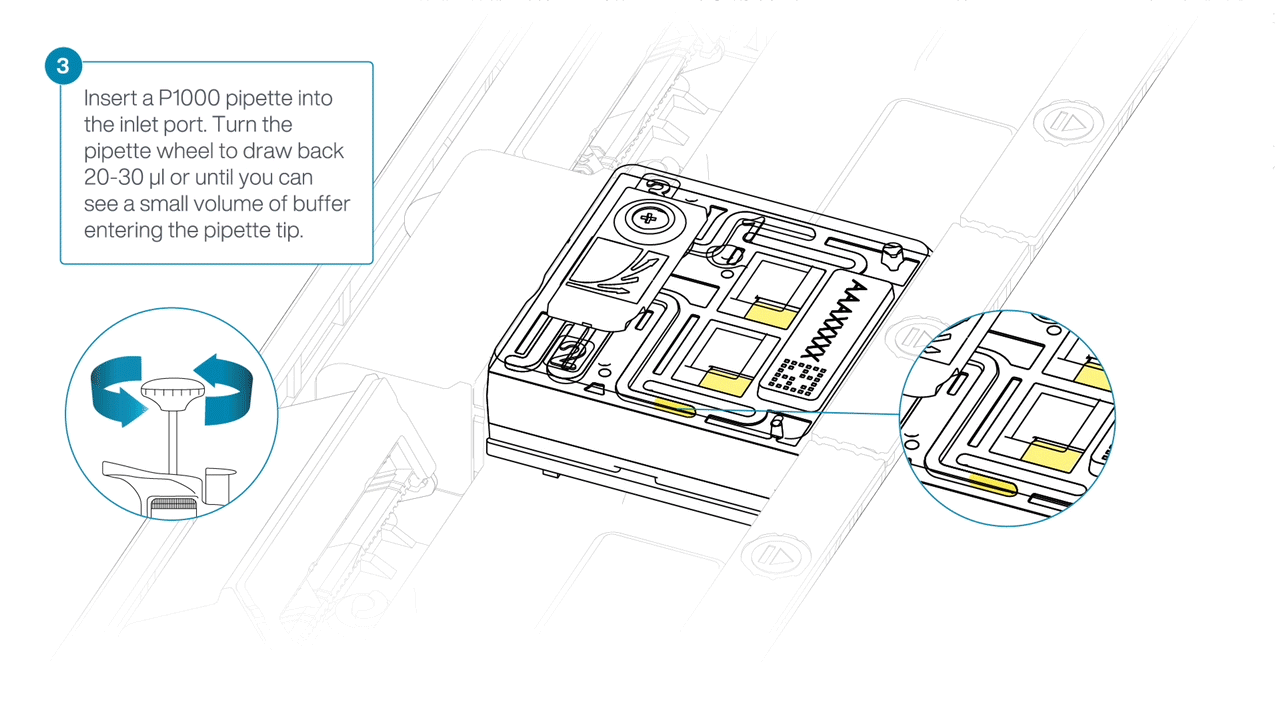
-
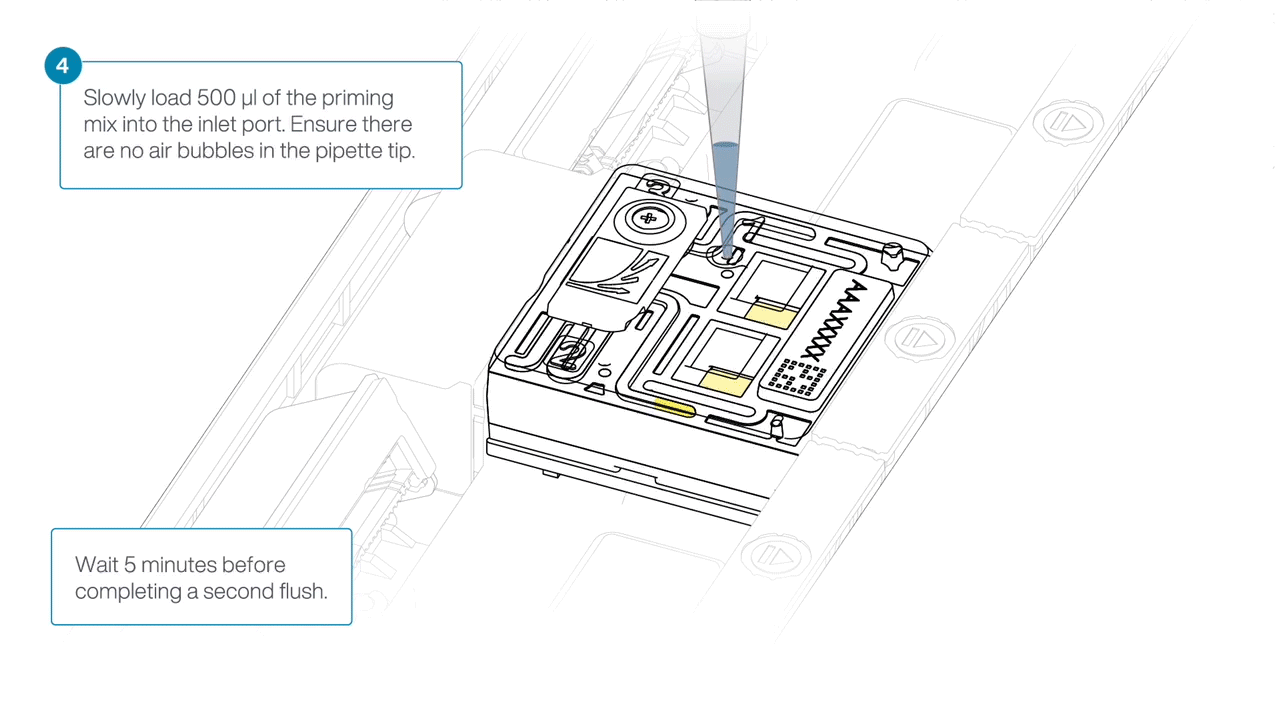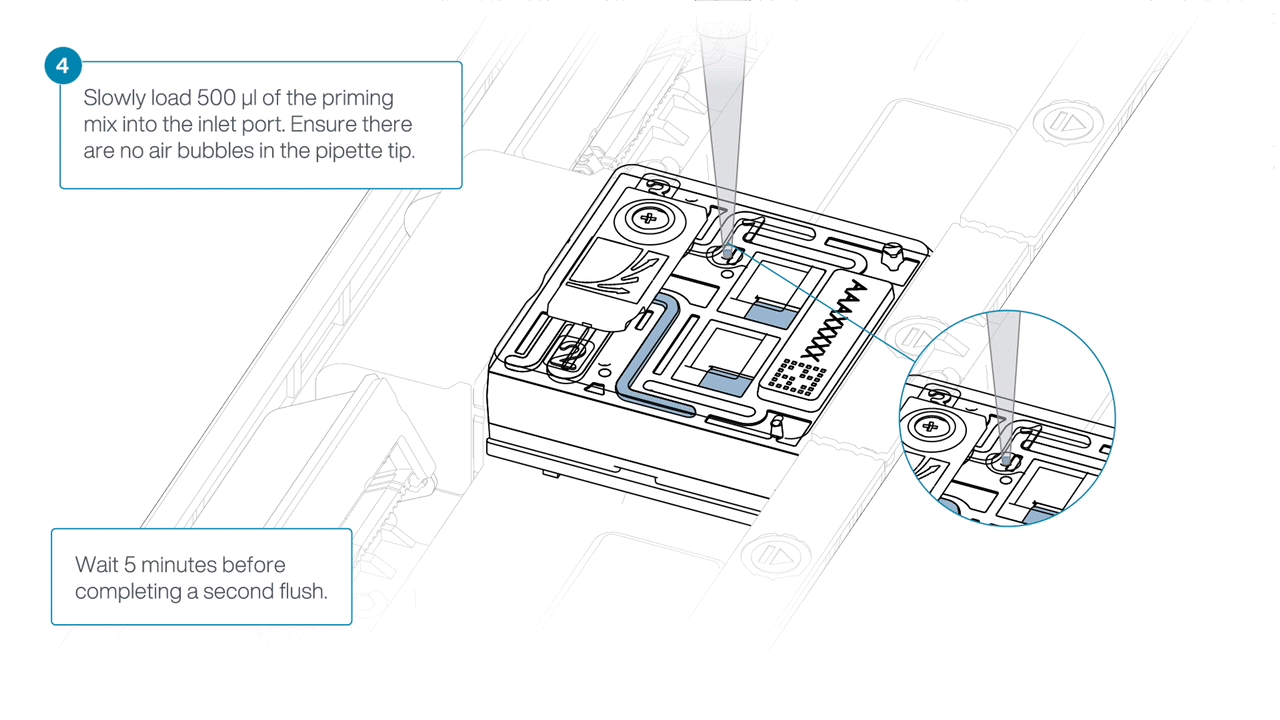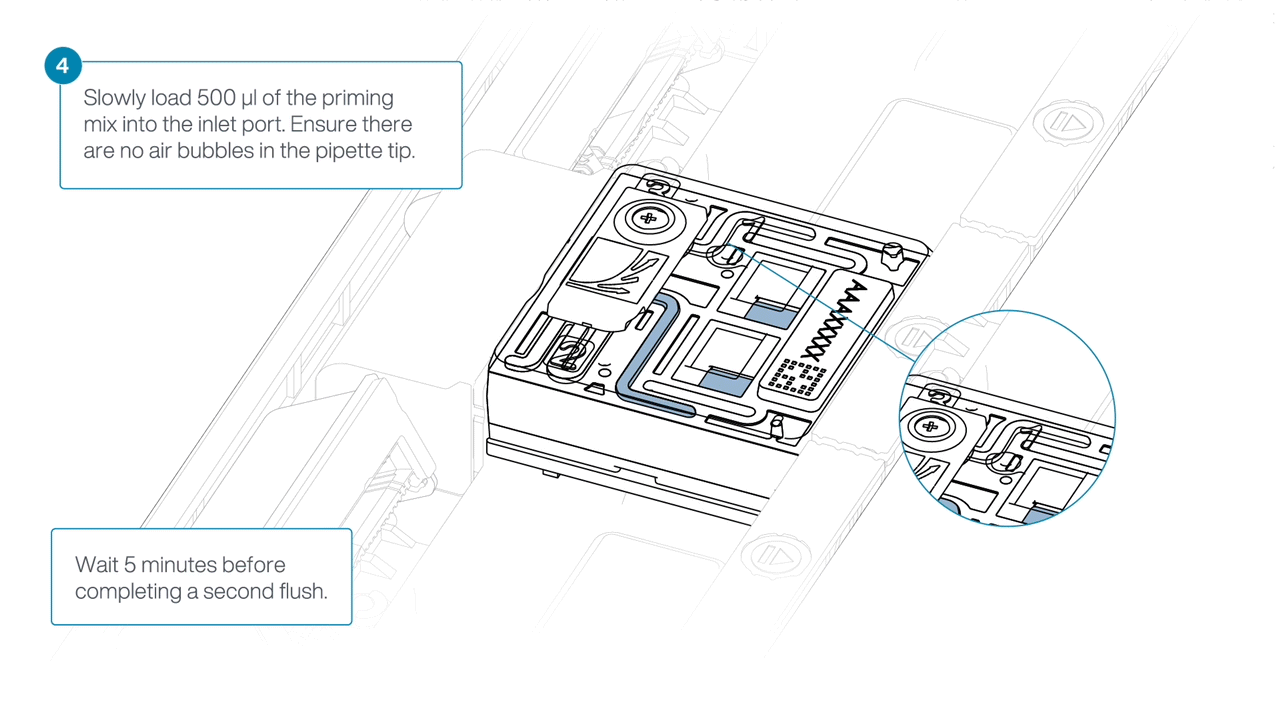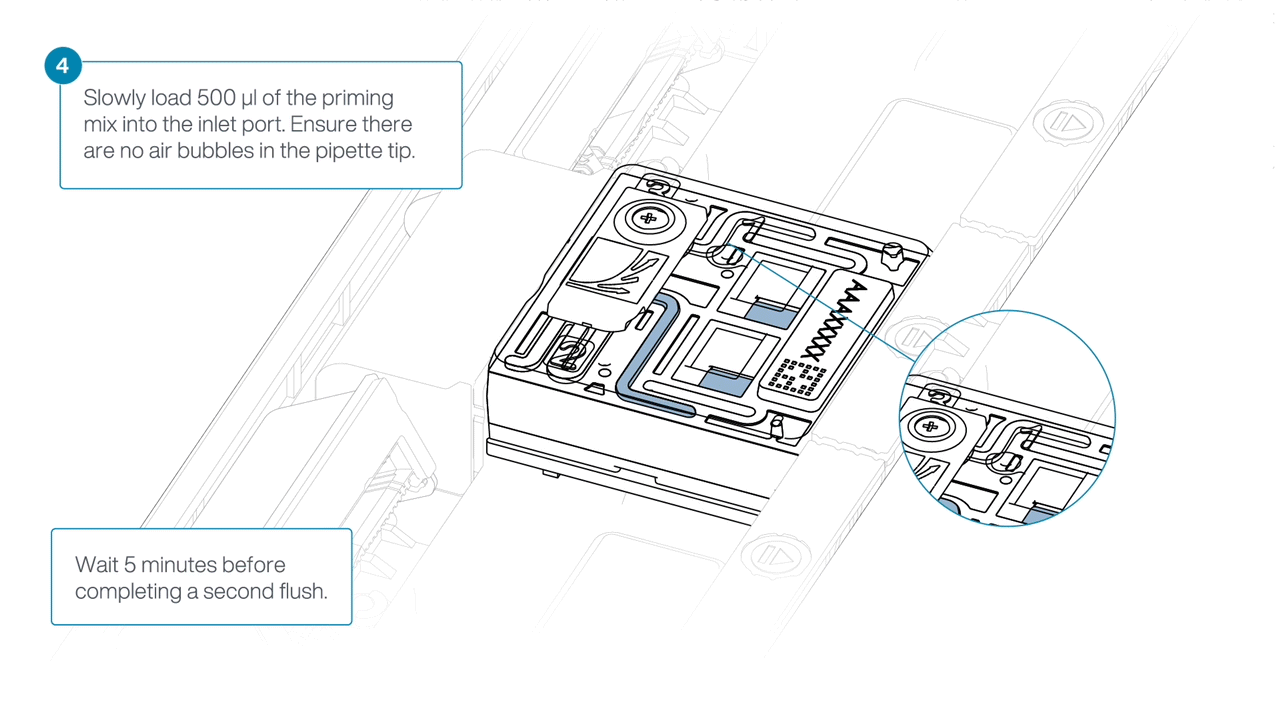
Load 500 µl of the priming mix into the flow cell via the inlet port, avoiding the introduction of air bubbles. Wait five minutes. During this time, prepare the library for loading using the next steps in the protocol.
-
Thoroughly mix the contents of the Loading Beads II (LBII) by pipetting.
-
In a new tube, prepare the library for loading as follows:
Reagent Volume per flow cell Sequencing Buffer II (SBII) 75 µl Loading Beads II (LBII) thoroughly mixed before use, or Loading Solution (LS), if using 51 µl DNA library 24 µl Total 150 µl -
Optional actionThe Multiplex Ligation Kit is designed for users running multiple flow cells. When handling multiple DNA libraries, the Sequencing Buffer (SBII) and Loading Beads II (LBII) can be combined in a master mix:
- Mix the Sequencing Buffer II (SBII) and Loading Beads II (LBII) as described above, scaling up the final volume for the appropriate number of samples and adding up to 20% excess of each reagent.
- Mix the master mix by pipetting immediately before adding to the DNA samples
- Pipette 126 µl of the master mix into each DNA sample-containing tube.
- Mix the samples by pipetting.
-
Complete the flow cell priming by slowly loading 500 µl of the priming mix into the inlet port.
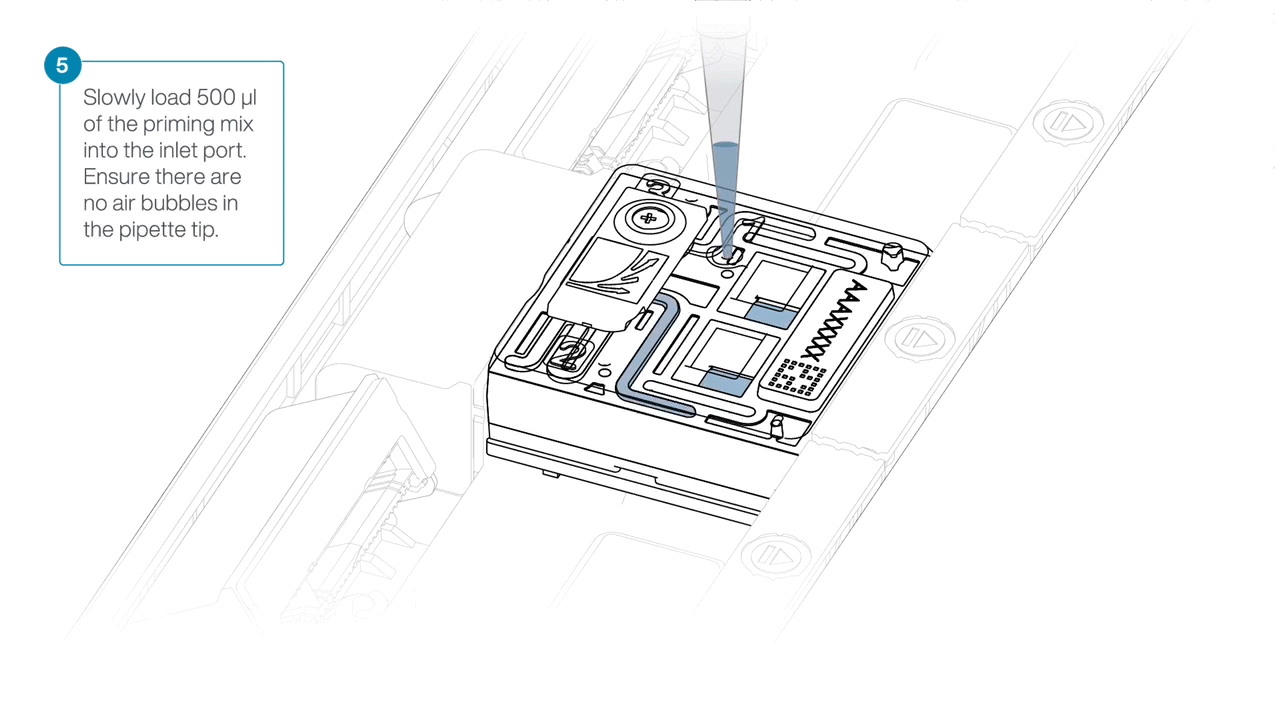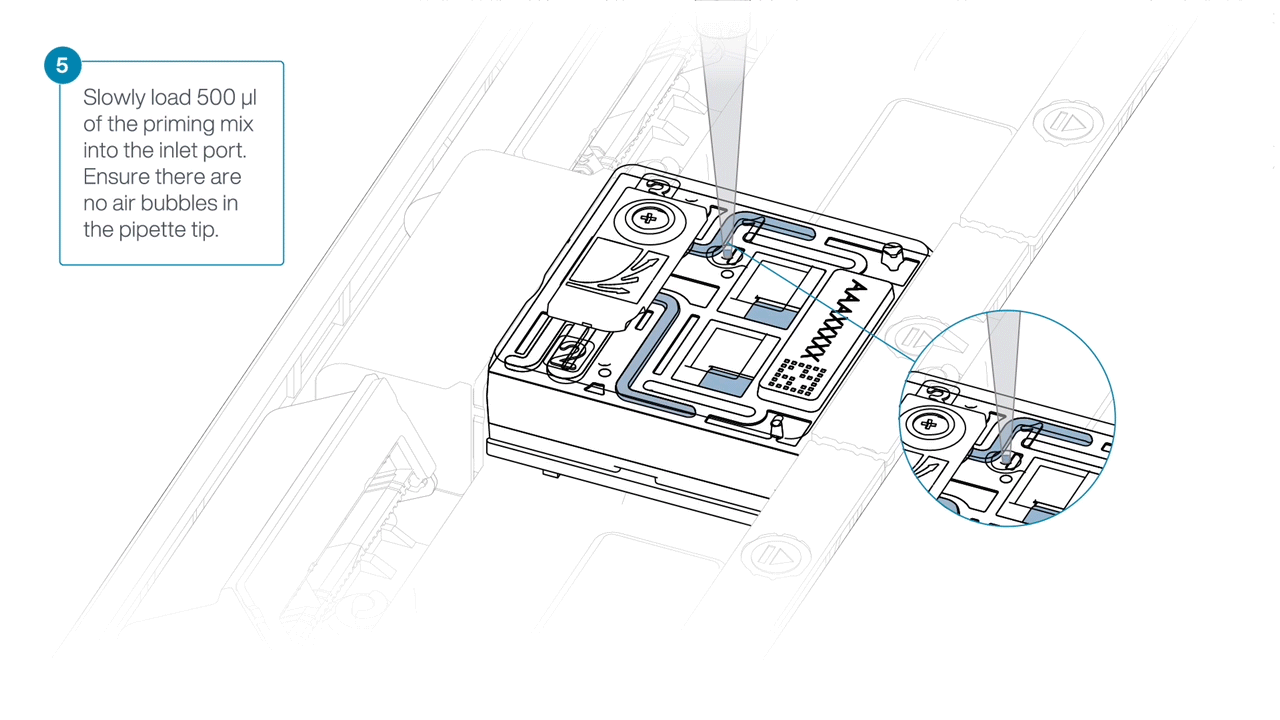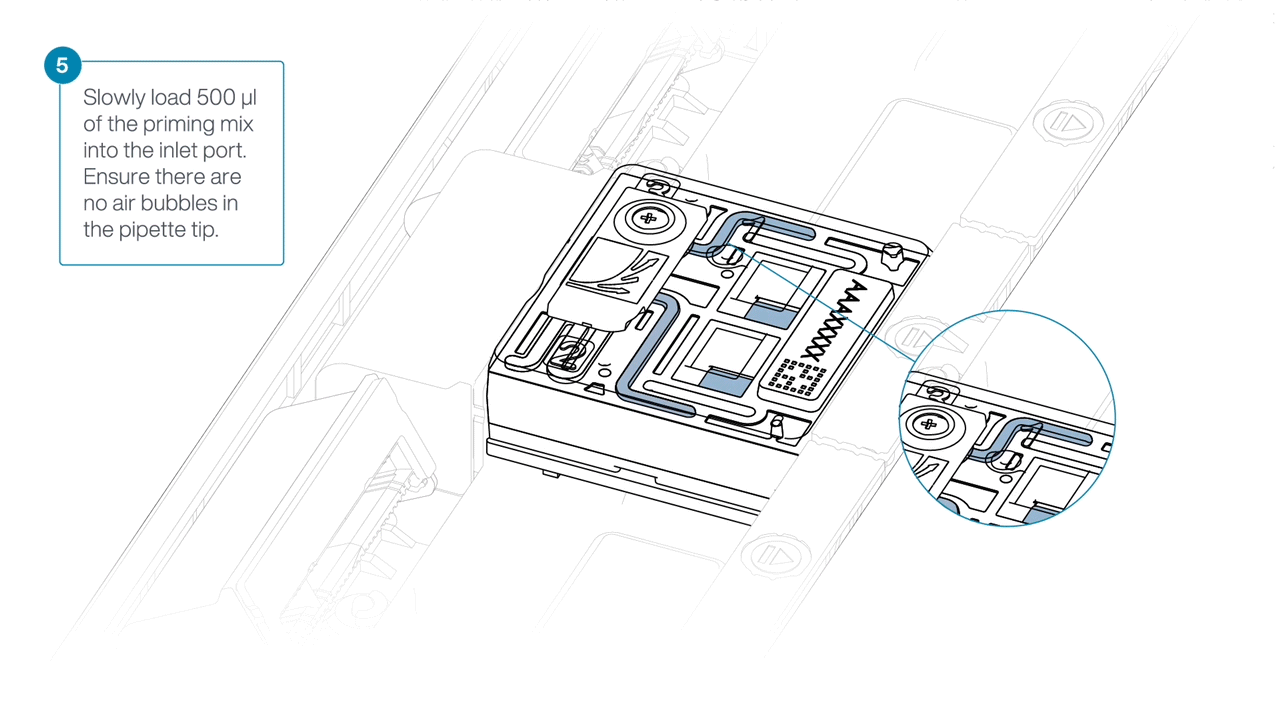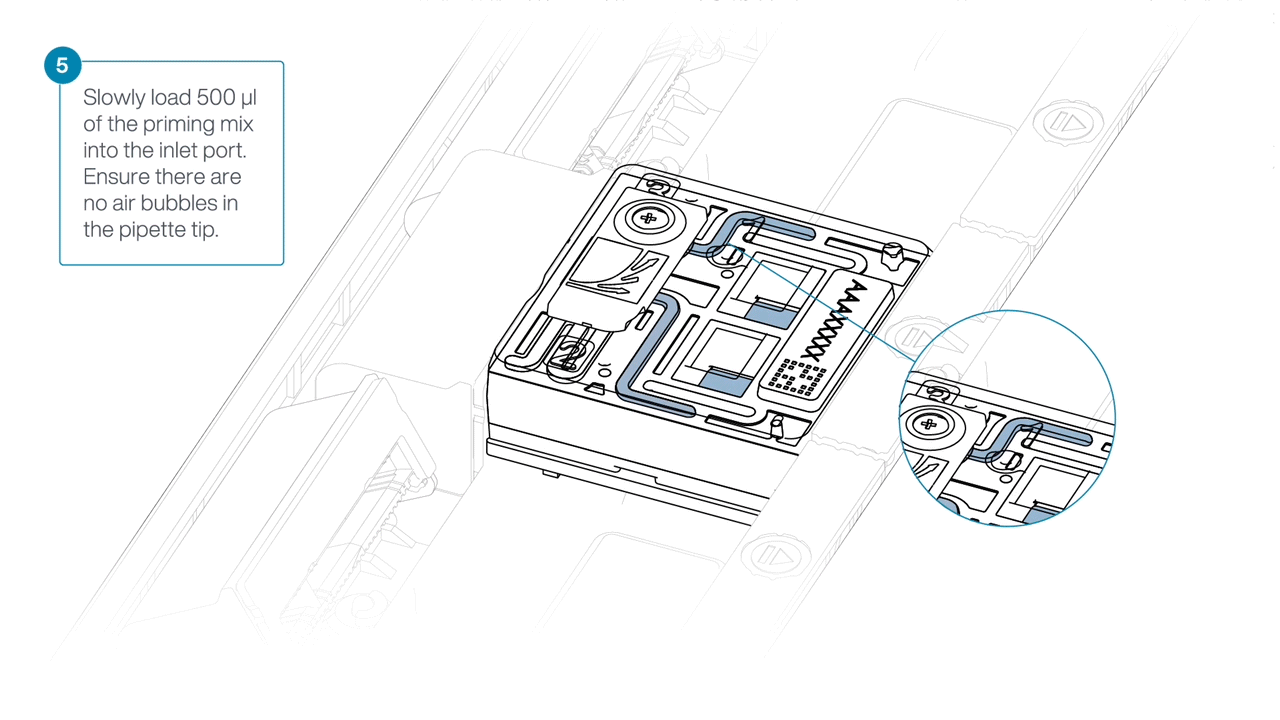
-
Mix the prepared library gently by pipetting up and down just prior to loading.
-
Using a P1000, insert the pipette tip into the inlet port and add 150 µl of library.
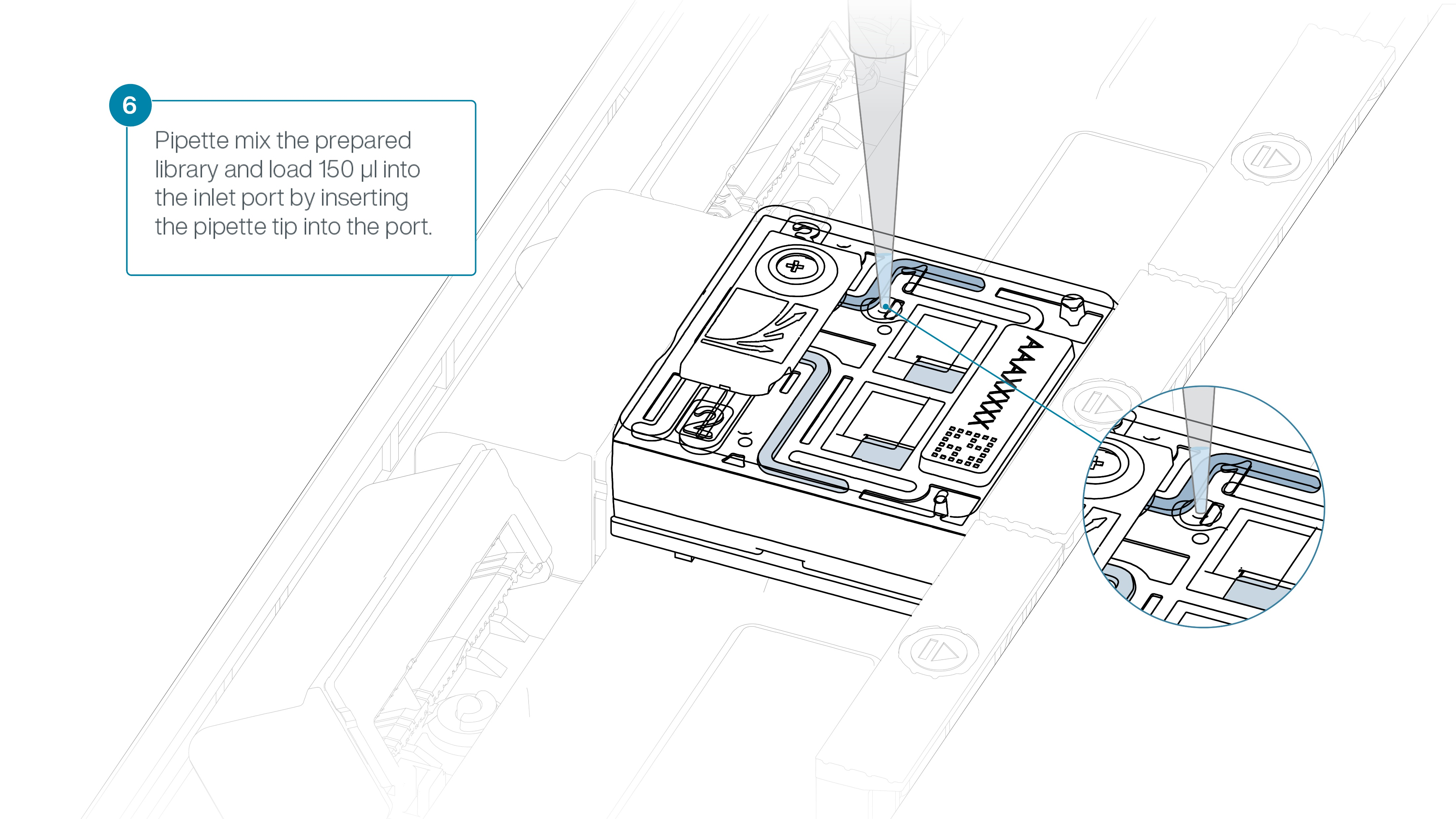
-
Close the valve to seal the inlet port.
-
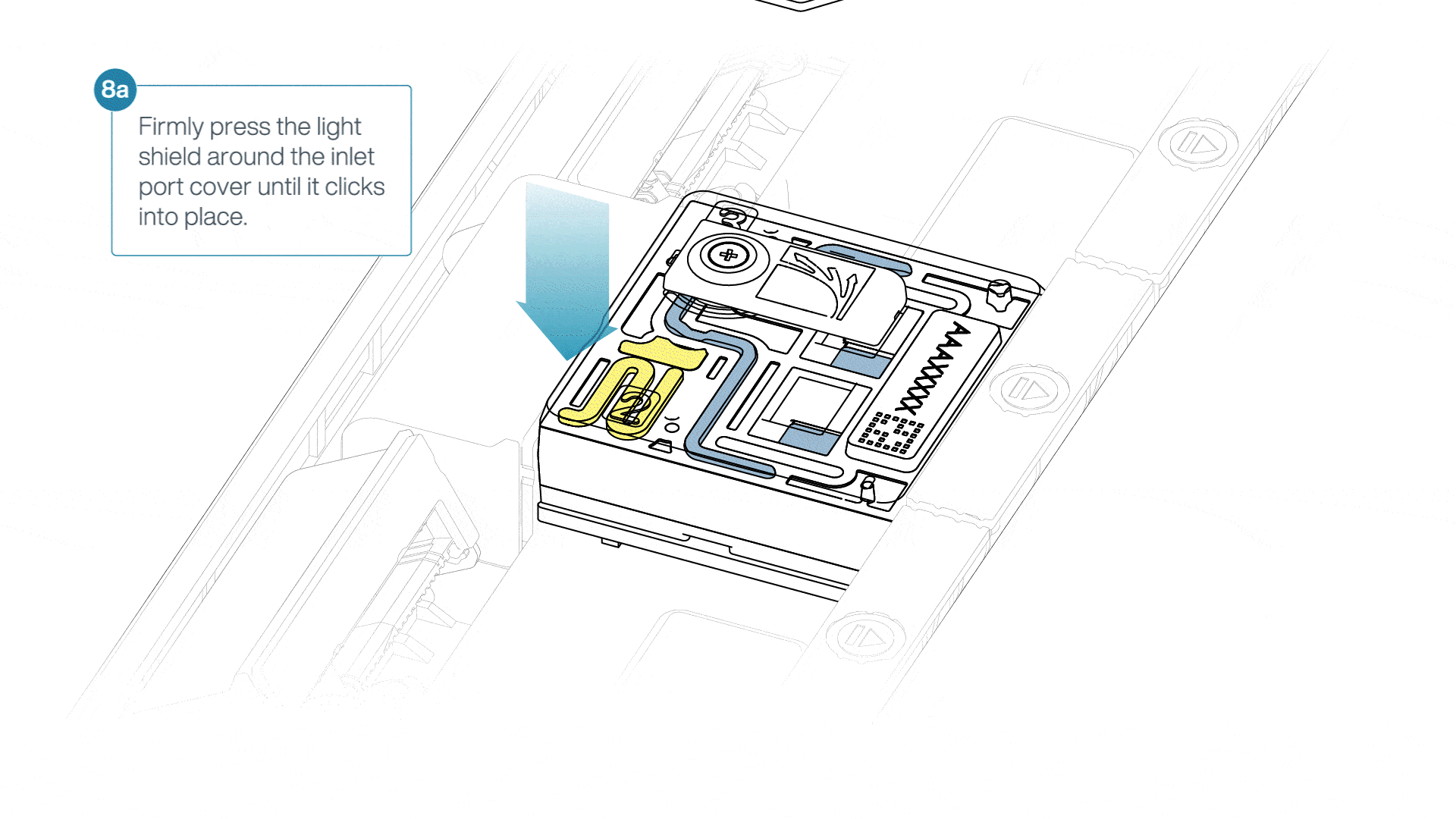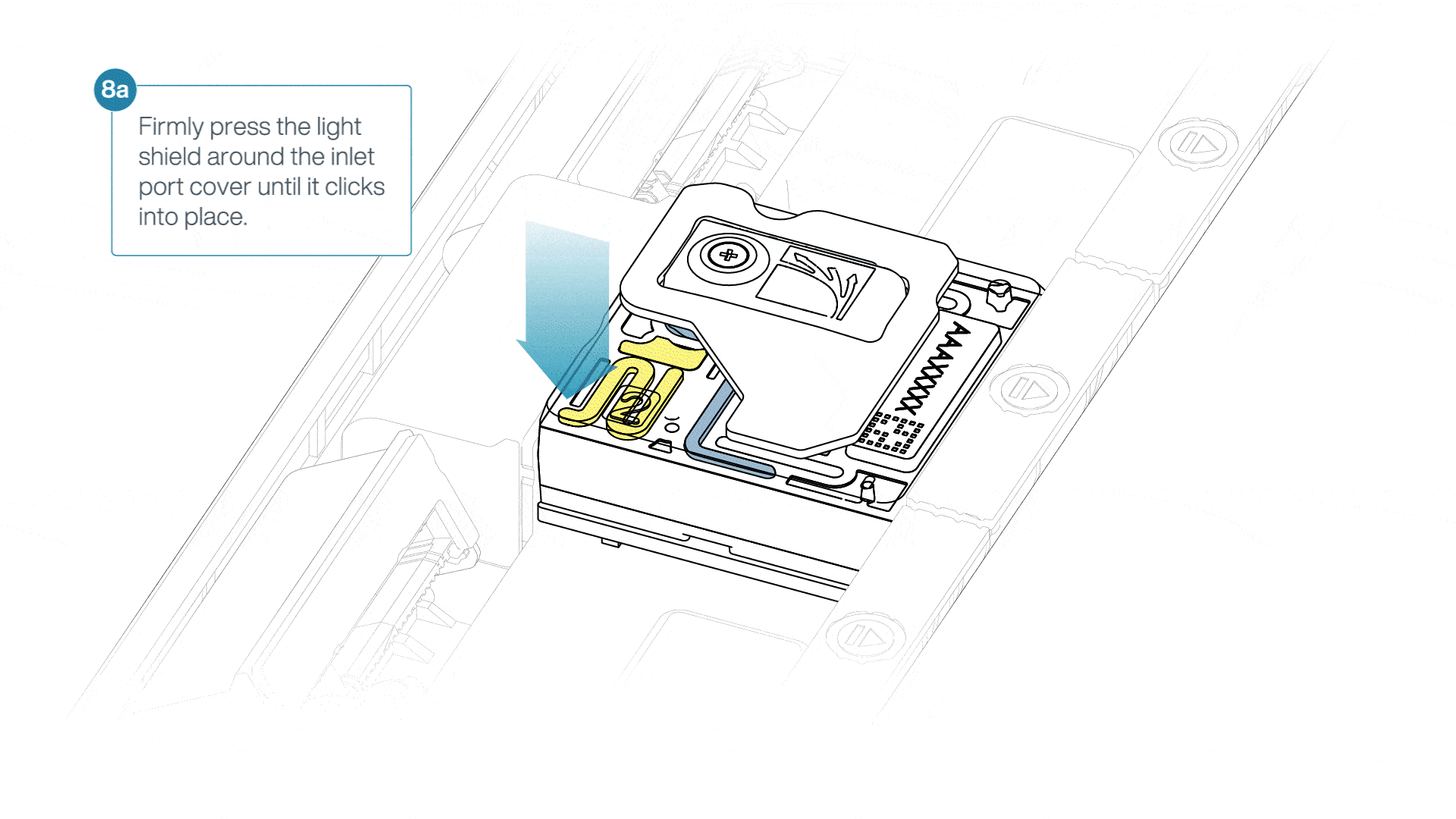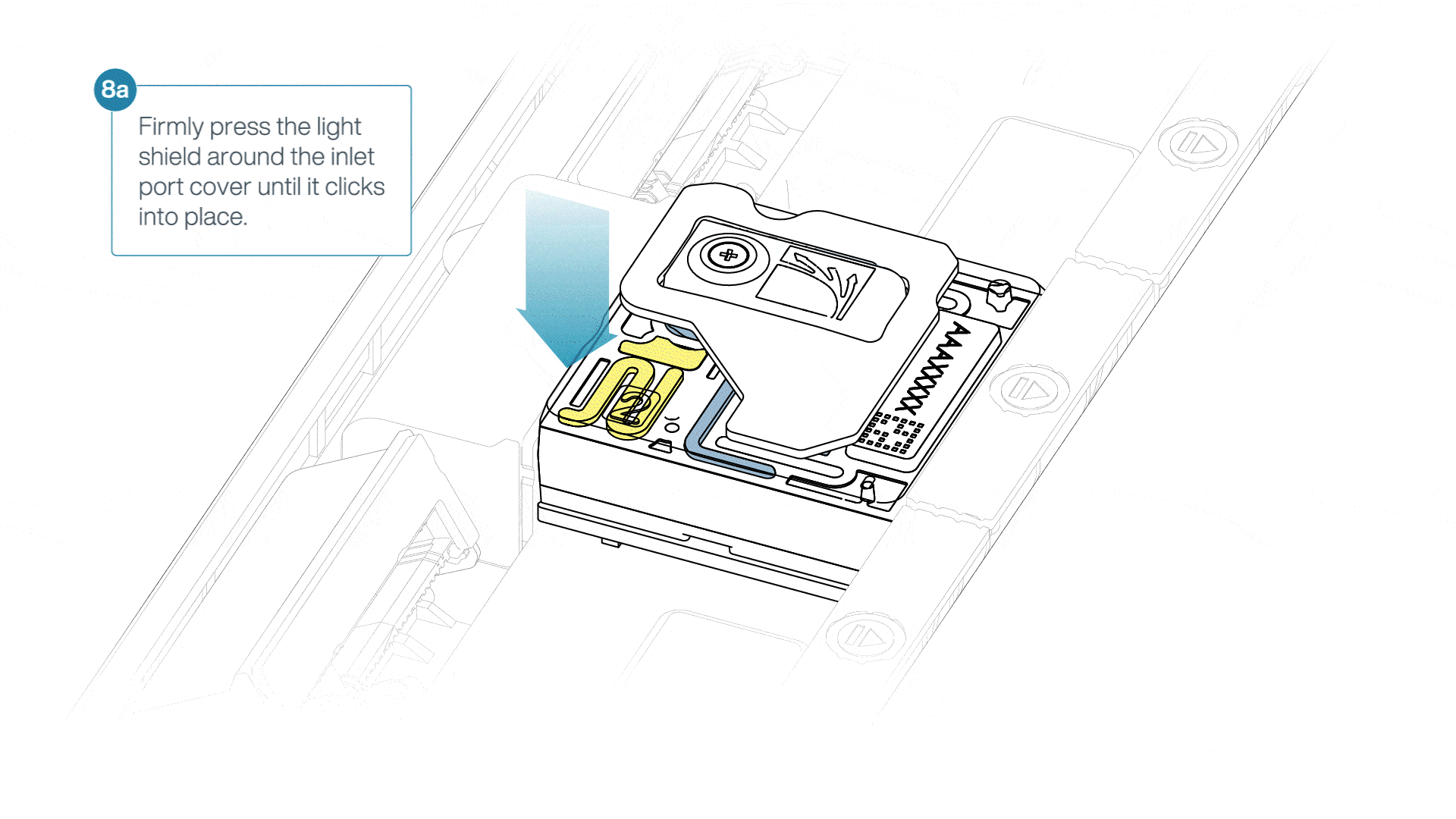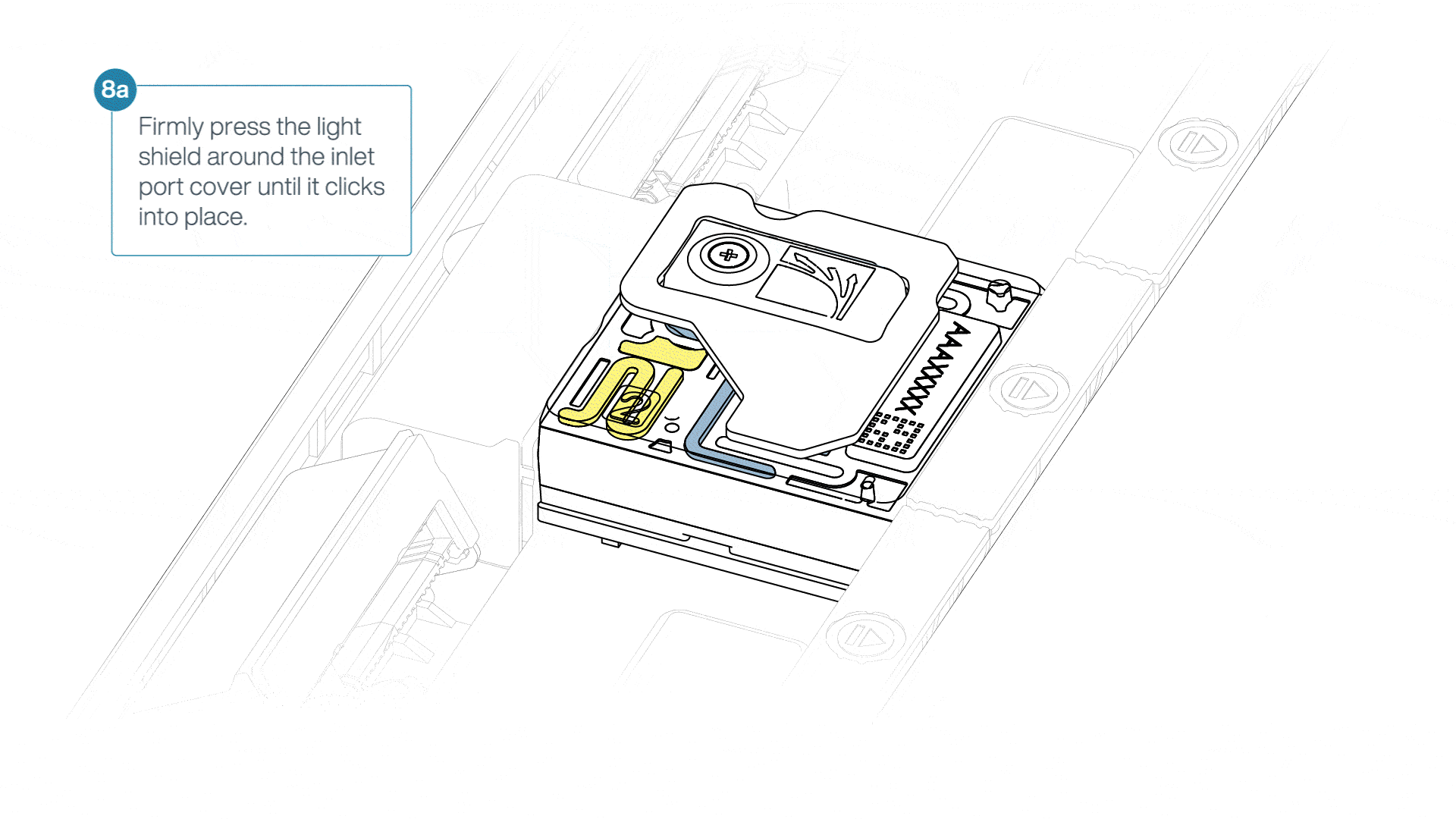
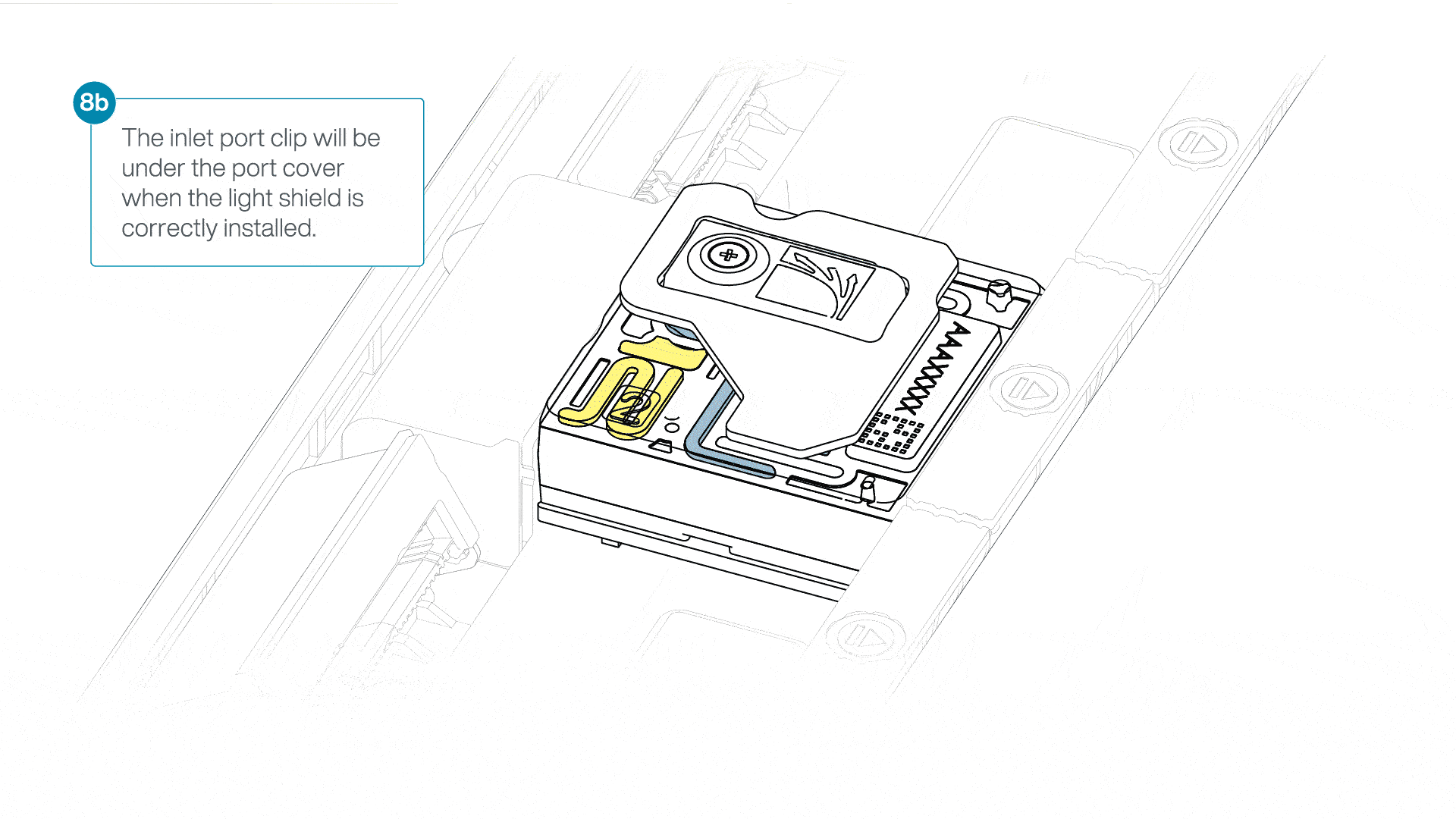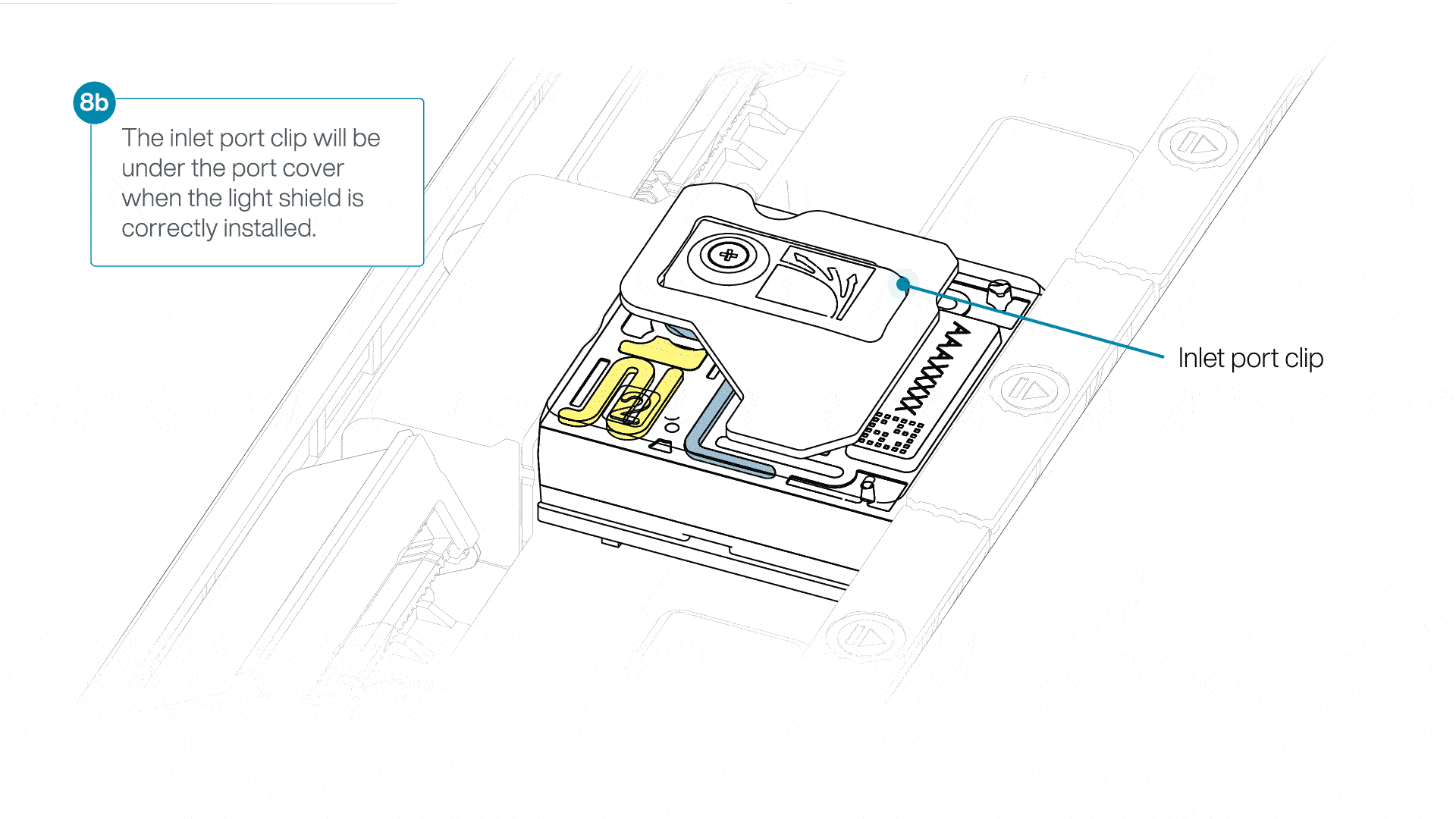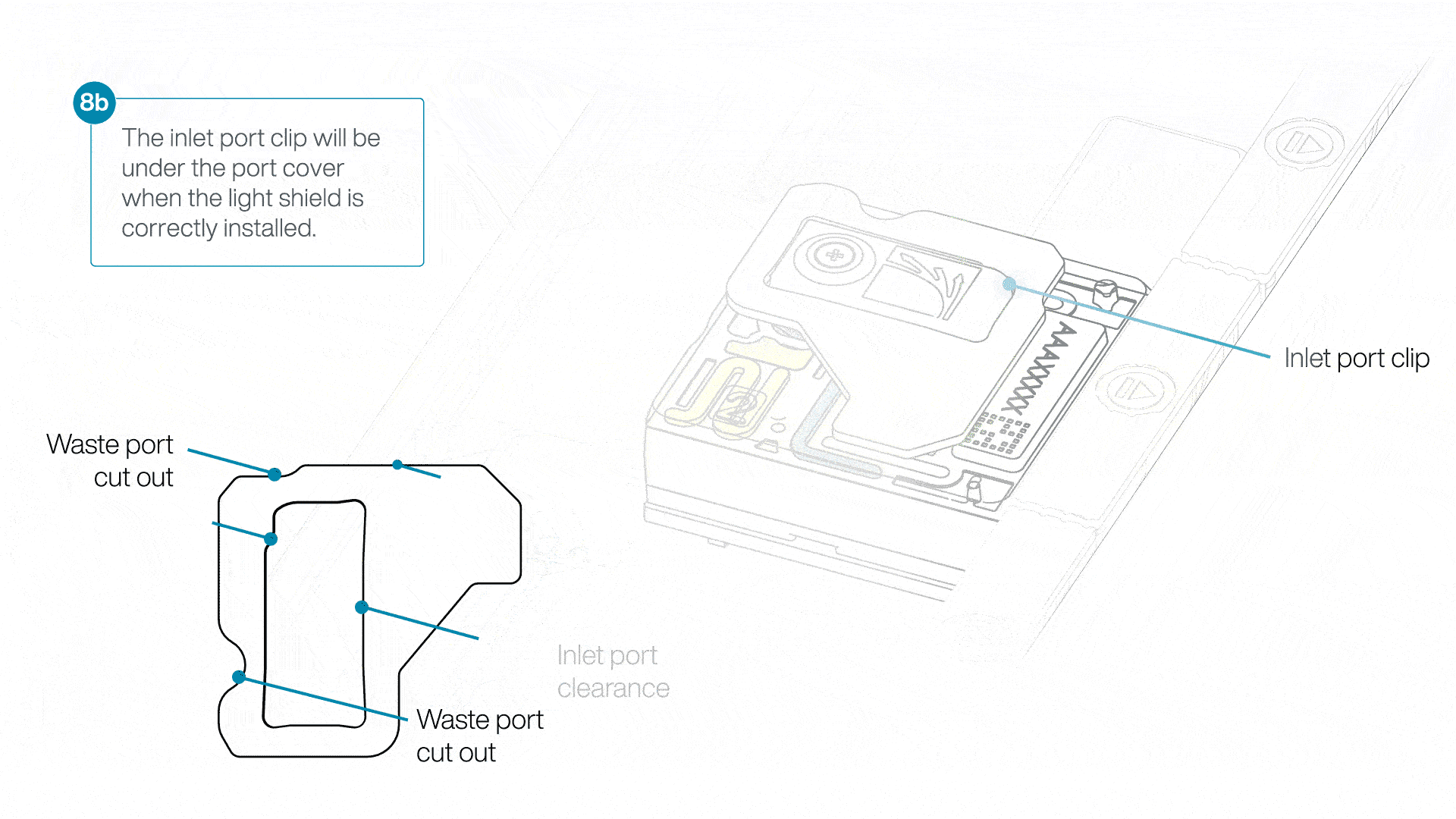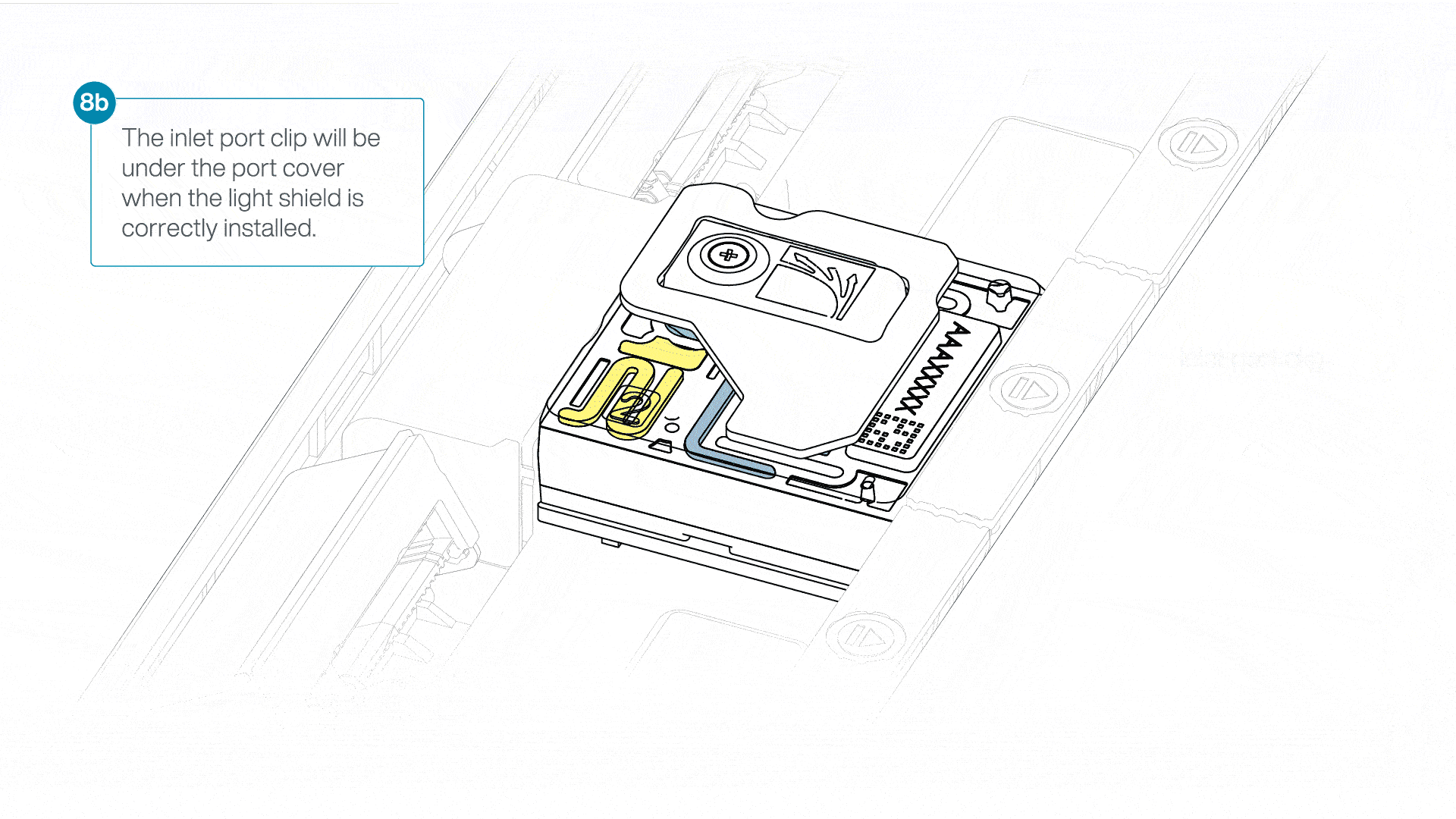
If the light shield has been removed from the flow cell, install the light shield as follows:
- Align the inlet port cut out of the light shield with the inlet port cover on the flow cell. The leading edge of the light shield should sit above the flow cell ID.
- Firmly press the light shield around the inlet port cover. The inlet port clip will click into place underneath the inlet port cover.
-
For multiple flow cell washing, use the same experiment name and identifying sample IDs for all runs to enable all flow cells to be paused simultaneously.