-
Introduction to barcoding amplicons
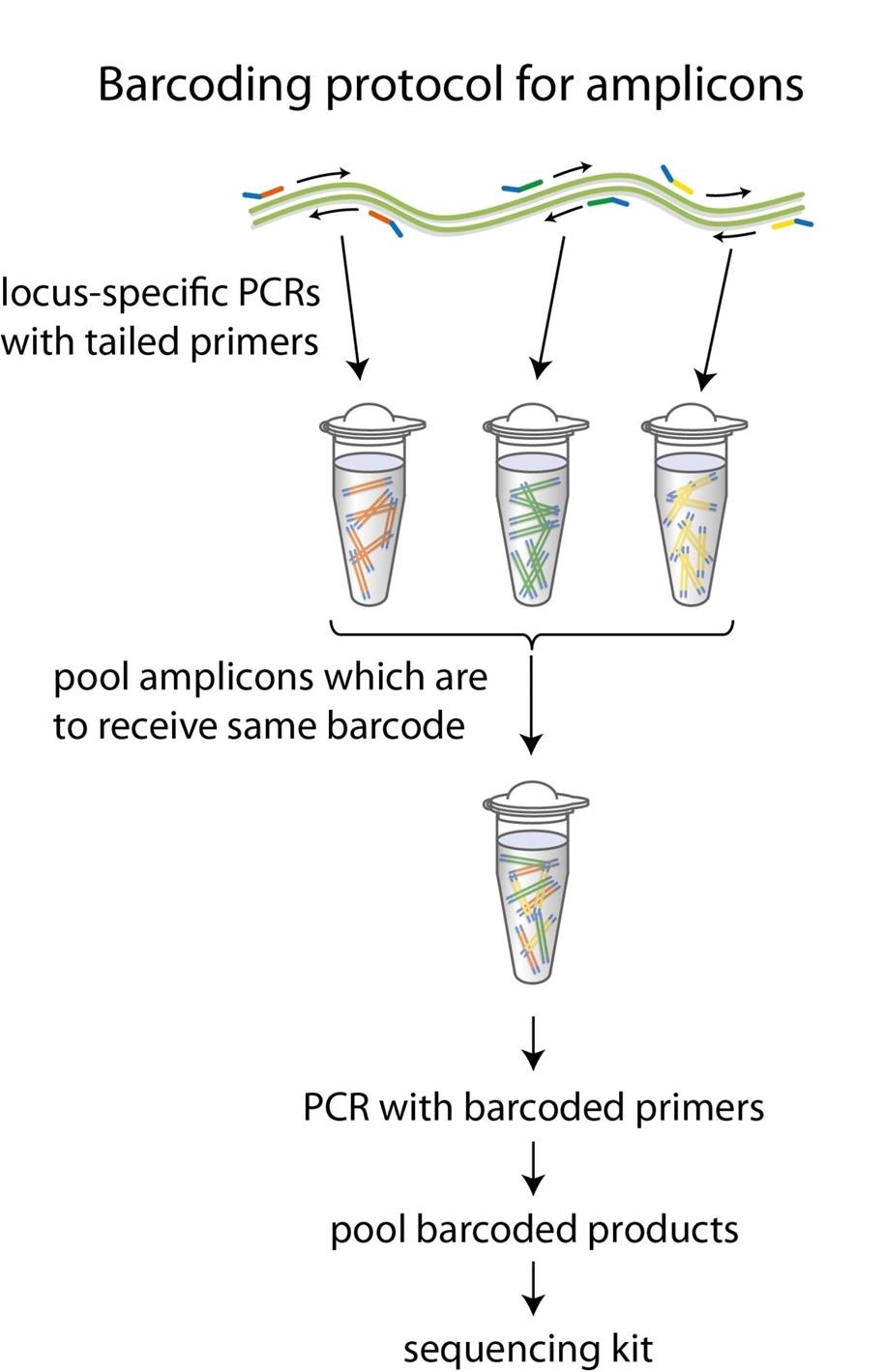
You will need to perform a first PCR using your specific primers which are tailed with the universal sequences given below, and then a second PCR to incorporate the Oxford Nanopore barcode sequences into your amplicons. Multiple first-round PCR products can be pooled together so that each amplicon sample in the pool receives the same barcode in the second PCR.
-
Universal sequences for tailing PCR primers
The first PCR amplification requires tailed primers to be used which carry these sequences:
5’ TTTCTGTTGGTGCTGATATTGC-[project-specific forward primer sequence] 3’
5’ ACTTGCCTGTCGCTCTATCTTC-[project-specific reverse primer sequence] 3’
-
We recommend a purification step at the end of this PCR reaction using standard methods which are suitable for the amplicon size, for example AMPure XP beads, to remove proteins, salts, dNTPs and primers.
-
All amplicon samples which are to receive the same barcode should be quantified and pooled in the desired ratios before the barcoding PCR is performed.
-
In the second PCR, a typical template concentration is 100-200 fmol, but the input mass and the number of PCR cycles can be adjusted as appropriate, depending upon the requirements of the experiment.
-
The Barcoding kit contains Oxford Nanopore primers with barcode sequences. These primers are added in a second amplification reaction.
-
The Oxford Nanopore Barcoding primers are supplied at 10 µM and should be used at a final concentration of 0.2 µM.