-
Introduction to the 16S Barcoding Kit 24 V14
This protocol describes how to carry out rapid barcoding of 16S amplicons using the 16S Barcoding Kit 24 V14 (SQK-16S114.24). Due to the presence of both highly conserved (adequate for universal primers and phylogenetic signal) and highly variant regions (different across species), the 16S rRNA gene is often used for sequence-based bacterial identification.
The 16S Barcoding Kit 24 V14 enables access to rapid 16S sequencing for organism identification. By narrowing down to a specific region of interest, you can see all the organisms in the sample without sequencing unnecessary regions of the genome, making the test quicker and more economical. There are 24 unique barcodes, allowing you to pool up to 24 different samples in one sequencing experiment.
After sequencing, you can perform downstream analysis using the EPI2ME 16S workflow (wf-16s) to classify 16S amplicons from your samples.
Steps in the sequencing workflow:
Prepare for your experiment
You will need to:
- Extract your gDNA, and check its length, quantity and purity using the Input DNA/RNA QC protocol. The quality checks performed during the protocol are essential in ensuring experimental success
- Ensure you have your sequencing kit, the correct equipment and third-party reagents
- Download the software for acquiring and analysing your data
- Check your flow cell to ensure it has enough pores for a good sequencing run
Library preparation
The table below is an overview of the steps required in the library preparation, including timings and stopping points.
Library preparation step Process Time Stop option 16S barcoded PCR amplification Amplify the 16S gene using barcodes supplied in the kit 10 minutes + PCR 4°C overnight Barcoded sample pooling and bead clean-up Quantify and pool the barcoded samples and perform a library clean-up using beads 15 minutes 4°C short-term storage or for repeated use, such as re-loading your flow cell.
-80°C for single-use long-term storage.Adapter ligation Attach the rapid sequencing adapters to the to the DNA ends. 5 minutes We strongly recommend sequencing your library as soon as it is adapted. Priming and loading the flow cell Prime the flow cell and load the prepared DNA library for sequencing 5 minutes 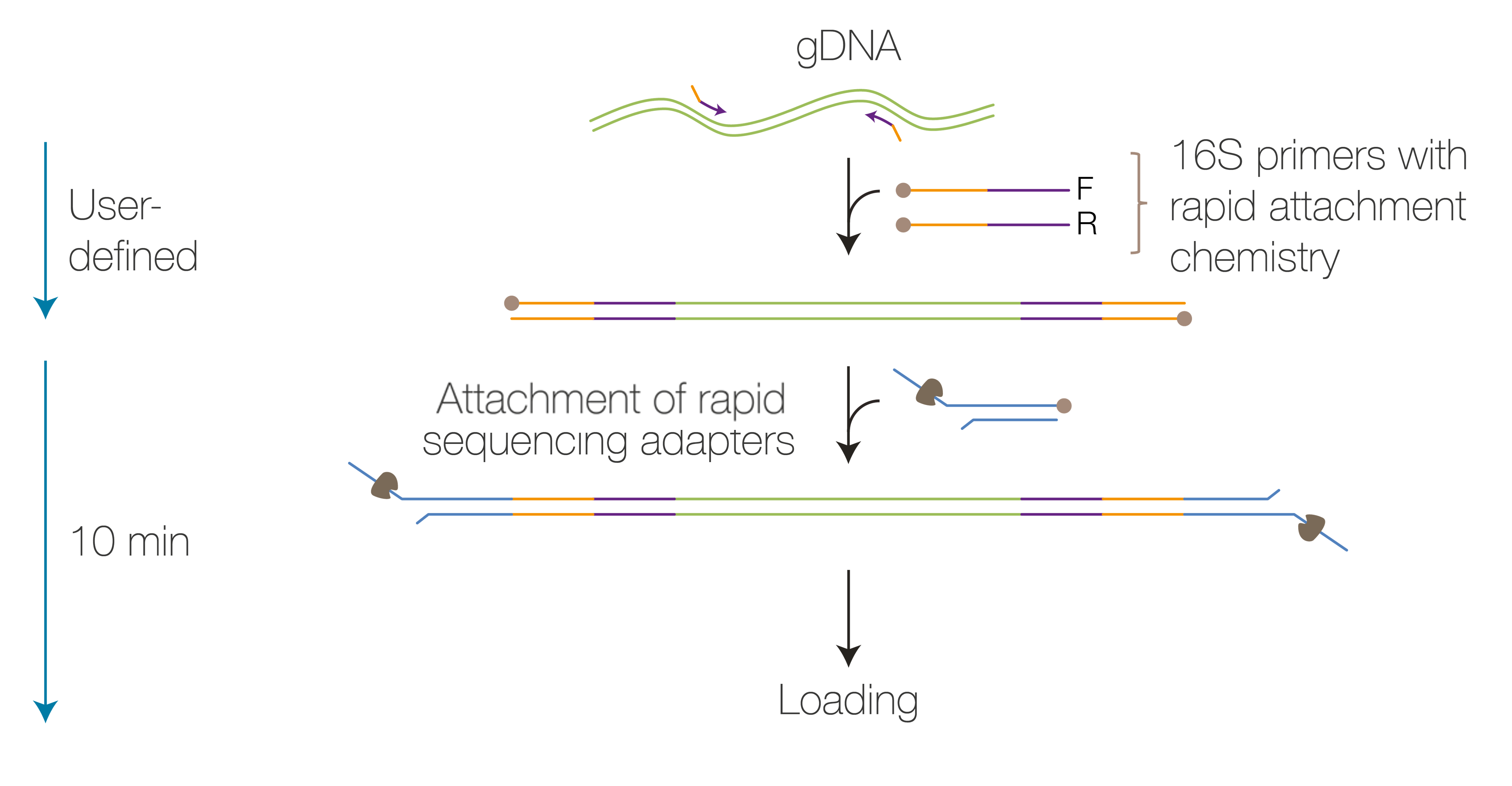
Sequencing and analysis
You will need to:
- Start a sequencing run using the MinKNOW software, which will collect raw data from the device and convert it into basecalled reads
- Optional: Start the EPI2ME software and select the wf-16S workflow
