- Materials
-
- Multiplex Ligation Sequencing Kit XL (SQK-MLK111.96-XL)
- 1200 ng gDNA per sample
- Consumables
-
- NEB Blunt/TA Ligase Master Mix (NEB, M0367)
- NEBNext® Quick Ligation Reaction Buffer (NEB, B6058)
- NEBNext FFPE Repair Mix (NEB, M6630)
- NEBNext Ultra II End repair/dA-tailing Module (NEB, E7546)
- NEBNext Quick Ligation Module (NEB, E6056)
- Nuclease-free water (e.g. ThermoFisher, AM9937)
- Freshly prepared 80% ethanol in nuclease-free water
- Qubit dsDNA HS Assay Kit (Invitrogen, Q32851)
- 1.5 ml Eppendorf DNA LoBind tubes
- 2.0 ml Eppendorf DNA LoBind tubes
- Agencourt AMPure XP beads (Beckman Coulter™ cat # A63881)
- Qubit™ Assay Tubes (Invitrogen, Q32856)
- Hamilton 50 µl CO-RE tips with filter (Cat# 235948)
- Hamilton 300 µl CO-RE tips with filter (Cat# 235903)
- Hamilton 1000 µl CO-RE tips with filter (Cat# 235905)
- Hamilton PCR ComfortLid (Cat# 814300)
- Bio-Rad Hard-Shell® 96-Well PCR Plates (Cat# HSP9601)
- Hamilton 60 ml Reagent Reservoir, Self-Standing with Lid (Cat# 56694-01)
- Equipment
-
- P1000 pipette and tips
- P200 pipette and tips
- P20 pipette and tips
- Ice bucket with ice
- Timer
- Qubit fluorometer (or equivalent)
- Hamilton NGS STAR 96 (NGS STAR with Multi-Probe Head 96)
- Hamilton On-Deck Thermal Cycler (ODTC)
- Hamilton MTP landscape carrier (cat# 182365)
- Hamilton Ambion magnet adapter (cat# 10107866)
- Optional equipment
-
- Agilent Bioanalyzer (or equivalent)
- Eppendorf 5424 centrifuge (or equivalent)
-
For this protocol, you will need 1200 ng gDNA per sample for R9.4.1 flow cells.
We recommend using 80 ng/µl per sample with a minimum of 15 µl per well when preparing the input plate.
-
Input DNA
How to QC your input DNA
It is important that the input DNA meets the quantity and quality requirements. Using too little or too much DNA, or DNA of poor quality (e.g. highly fragmented or containing RNA or chemical contaminants) can affect your library preparation.
For instructions on how to perform quality control of your DNA sample, please read the Input DNA/RNA QC protocol.
Chemical contaminants
Depending on how the DNA is extracted from the raw sample, certain chemical contaminants may remain in the purified DNA, which can affect library preparation efficiency and sequencing quality. Read more about contaminants on the Contaminants page of the Community.
-
Input workfile
Input workfiles are required prior to running this protocol on the Hamilton NGS STAR 96.
Oxford Nanopore Technologies will provide a template of the input workfiles in an Excel file.
- In the DATA tab of the file, there will be the template for the library preparation data
- In the INFO tab, there will be the template for the identification data
Library preparation data example:
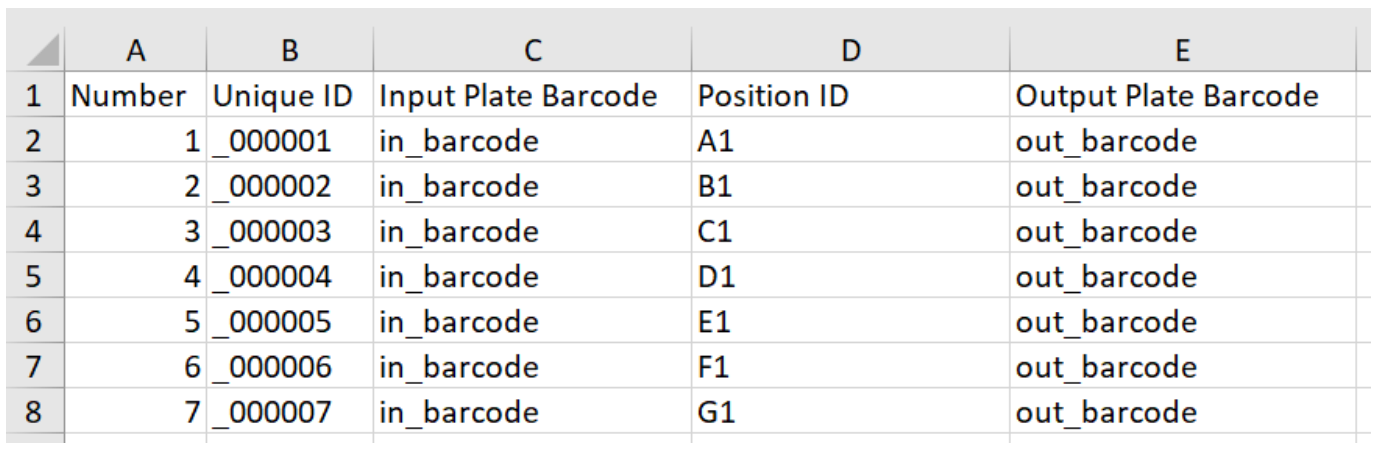
Identification data example:
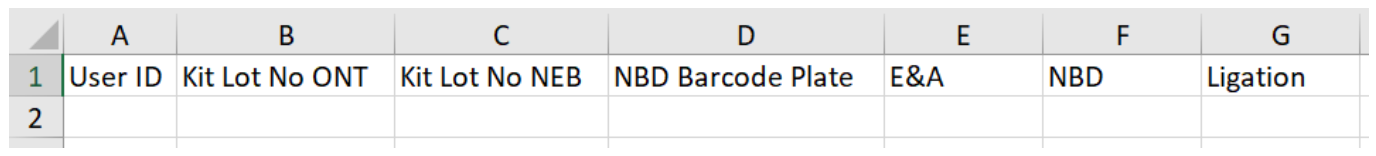
Output workfiles will be generated. Below are examples of the output workfiles after a run through of the method.
Library preparation data example:
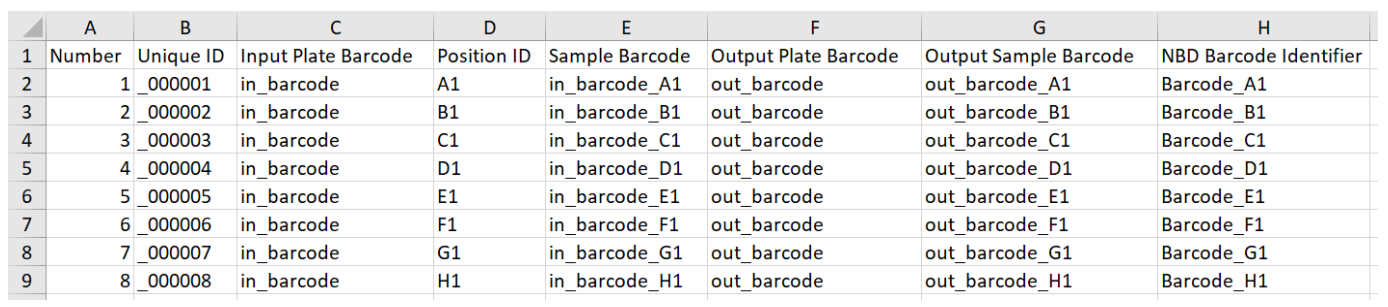
Identification data example:
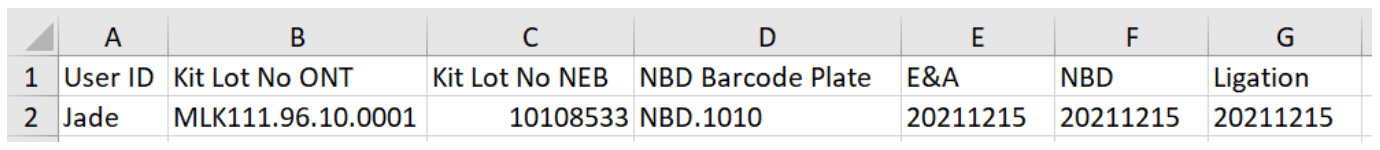

-
Hamilton NGS STAR 96 and deck layout
This method has been tested and validated using the Hamilton NGS STAR 96 (with Multi-Probe Head 96), Hamilton Ambion magnet adapter, Hamilton MTP landscape carrier and Hamilton On-Deck Thermal Cycler (ODTC).
Deck layout
The deck layout has been updated from the standard layout. Please see the "Prepare the deck" step of this protocol for more details.
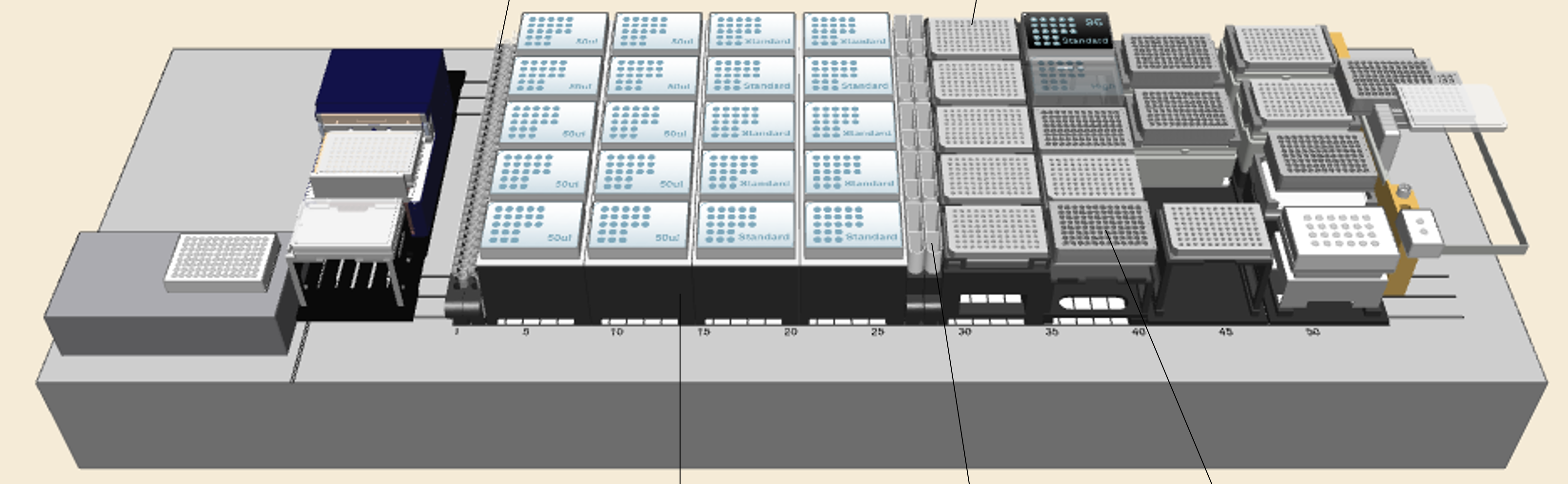

-
Data tracking
For data tracking purposes, we have included the option to add user ID before starting any process. This can be filled using any method the user prefers.
We recommend using barcode stickers to track the input and output plates for data tracking. These can be tracked on the workfile and entered on the UI alongside the reagent lot barcodes when prompted.
-
Convenient reagent kits are available on request from NEB for the Multiplex Ligation Sequencing Kit XL.
This will contain the appropriate NEB reagents and the required volumes for the protocol on the Hamilton NGS STAR 96. For more information from NEB, please see "Find Products for Nanopore Sequencing".
-
Multiplex Ligation Sequencing Kit XL (SQK-MLK111.96-XL) contents
Name Acronym Cap colour Number of vials Fill volume per vial (µl) Adapter Mix II T AMII T Green 1 320 Sequencing Buffer II SBII Red 4 1,500 Loading Beads II LBII Pink 4 1,500 Loading Solution LS White cap, pink sticker 4 1,500 EDTA EDTA Clear 1 700 Elution Buffer EB 15 ml bottle 1 10,000 Long Fragment Buffer LFB 30 ml bottle 1 20,000 Flush Buffer XL FB 30 ml bottle 6 15,500 Flush Tether FLT White cap, purple sticker 2 1,600 Native Barcodes NB01-96 N/a 1 plate 8 µl per well -
Consumables and equipment quantities:
Consumables No. of consumables for all conditions Hamilton 50 µl CO-RE tips with filter 960 Hamilton 300 µl CO-RE tips with filter 960 Hamilton 1000 µ CO-RE tips with filter 96 Bio-Rad Hard-Shell® 96-well PCR Plate 8 This protocol requires the tip decks to be completely filled before starting a run. Partially filled tip decks will cause an error with the liquid handling robot.
We recommend using Hamilton tips for efficient liquid handling.
-
Reagent quantities:
Note: Volumes for x48 samples will be available soon.
Full method
Reagents x96 samples 80% ethanol 80 ml AMPure XP Beads 15 ml Nuclease-free water 10 ml Long Fragment Buffer (LFB) 2 bottles Elution Buffer (EB) 1 bottle EDTA 1 vial Native Barcode plate 1 plate Adapter Mix F (AMII F) 1 vial NEBNext FFRE DNA Repair Buffer 130 µl NEBNext FFPE DNA Repair Mix 90 µl Ultra II End Prep Reaction Buffer 130 µl Ultra II End Prep Enzyme Mix 120 µl Blunt/TA Ligase Master Mix 1210 µl NEBNext Quick Ligation Reaction Buffer (5x) 590 µl Quick T4 DNA Ligase 310 µl
Multiple steps combined:End Repair and Adenylation step to Native Barcode Ligation step
Reagents x96 samples 80% ethanol 80 ml AMPure XP Beads 15 ml Nuclease-free water 10 ml NEBNext FFRE DNA Repair Buffer 130 µl NEBNext FFPE DNA Repair Mix 90 µl Ultra II End Prep Reaction Buffer 130 µl Ultra II End Prep Enzyme Mix 120 µl Blunt/TA Ligase Master Mix 1210 µl EDTA 1 vial Native Barcode plate 1 plate Native Barcode Ligation step to Adapter Ligation step
Reagents x96 samples 80% ethanol 40 ml AMPure XP Beads 15 ml Nuclease-free water 10 ml Long Fragment Buffer (LFB) 2 bottles Elution Buffer (EB) 1 bottle Blunt/TA Ligase Master Mix 1210 µl EDTA 1 vial Native Barcode plate 1 plate Adapter Mix F (AMII F) 1 vial NEBNext Quick Ligation Reaction Buffer (5x) 590 µl Quick T4 DNA Ligase 310 µl
Individual steps:End Repair step
Reagents x96 samples 80% ethanol 40 ml AMPure XP Beads 15 ml Nuclease-free water 10 ml NEBNext FFRE DNA Repair Buffer 130 µl NEBNext FFPE DNA Repair Mix 90 µl Ultra II End Prep Reaction Buffer 130 µl Ultra II End Prep Enzyme Mix 120 µl Native Barcode Ligation step
Reagents x96 samples 80% ethanol 40 ml AMPure XP Beads 15 ml Nuclease-free water 10 ml Blunt/TA Ligase Master Mix 1210 µl EDTA 1 vial Native Barcode plate 1 plate Adapter Ligation step
Reagents x96 samples AMPure XP Beads 15 ml Long Fragment Buffer (LFB) 2 bottles Elution Buffer (EB) 1 bottle NEBNext Quick Ligation Reaction Buffer (5x) 590 µl Quick T4 DNA Ligase 310 µl -
Native barcode sequences
Component Forward sequence Reverse sequence NB01 CACAAAGACACCGACAACTTTCTT AAGAAAGTTGTCGGTGTCTTTGTG NB02 ACAGACGACTACAAACGGAATCGA TCGATTCCGTTTGTAGTCGTCTGT NB03 CCTGGTAACTGGGACACAAGACTC GAGTCTTGTGTCCCAGTTACCAGG NB04 TAGGGAAACACGATAGAATCCGAA TTCGGATTCTATCGTGTTTCCCTA NB05 AAGGTTACACAAACCCTGGACAAG CTTGTCCAGGGTTTGTGTAACCTT NB06 GACTACTTTCTGCCTTTGCGAGAA TTCTCGCAAAGGCAGAAAGTAGTC NB07 AAGGATTCATTCCCACGGTAACAC GTGTTACCGTGGGAATGAATCCTT NB08 ACGTAACTTGGTTTGTTCCCTGAA TTCAGGGAACAAACCAAGTTACGT NB09 AACCAAGACTCGCTGTGCCTAGTT AACTAGGCACAGCGAGTCTTGGTT NB10 GAGAGGACAAAGGTTTCAACGCTT AAGCGTTGAAACCTTTGTCCTCTC NB11 TCCATTCCCTCCGATAGATGAAAC GTTTCATCTATCGGAGGGAATGGA NB12 TCCGATTCTGCTTCTTTCTACCTG CAGGTAGAAAGAAGCAGAATCGGA NB13 AGAACGACTTCCATACTCGTGTGA TCACACGAGTATGGAAGTCGTTCT NB14 AACGAGTCTCTTGGGACCCATAGA TCTATGGGTCCCAAGAGACTCGTT NB15 AGGTCTACCTCGCTAACACCACTG CAGTGGTGTTAGCGAGGTAGACCT NB16 CGTCAACTGACAGTGGTTCGTACT AGTACGAACCACTGTCAGTTGACG NB17 ACCCTCCAGGAAAGTACCTCTGAT ATCAGAGGTACTTTCCTGGAGGGT NB18 CCAAACCCAACAACCTAGATAGGC GCCTATCTAGGTTGTTGGGTTTGG NB19 GTTCCTCGTGCAGTGTCAAGAGAT ATCTCTTGACACTGCACGAGGAAC NB20 TTGCGTCCTGTTACGAGAACTCAT ATGAGTTCTCGTAACAGGACGCAA NB21 GAGCCTCTCATTGTCCGTTCTCTA TAGAGAACGGACAATGAGAGGCTC NB22 ACCACTGCCATGTATCAAAGTACG CGTACTTTGATACATGGCAGTGGT NB23 CTTACTACCCAGTGAACCTCCTCG CGAGGAGGTTCACTGGGTAGTAAG NB24 GCATAGTTCTGCATGATGGGTTAG CTAACCCATCATGCAGAACTATGC NB25 GTAAGTTGGGTATGCAACGCAATG CATTGCGTTGCATACCCAACTTAC NB26 CATACAGCGACTACGCATTCTCAT ATGAGAATGCGTAGTCGCTGTATG NB27 CGACGGTTAGATTCACCTCTTACA TGTAAGAGGTGAATCTAACCGTCG NB28 TGAAACCTAAGAAGGCACCGTATC GATACGGTGCCTTCTTAGGTTTCA NB29 CTAGACACCTTGGGTTGACAGACC GGTCTGTCAACCCAAGGTGTCTAG NB30 TCAGTGAGGATCTACTTCGACCCA TGGGTCGAAGTAGATCCTCACTGA NB31 TGCGTACAGCAATCAGTTACATTG CAATGTAACTGATTGCTGTACGCA NB32 CCAGTAGAAGTCCGACAACGTCAT ATGACGTTGTCGGACTTCTACTGG NB33 CAGACTTGGTACGGTTGGGTAACT AGTTACCCAACCGTACCAAGTCTG NB34 GGACGAAGAACTCAAGTCAAAGGC GCCTTTGACTTGAGTTCTTCGTCC NB35 CTACTTACGAAGCTGAGGGACTGC GCAGTCCCTCAGCTTCGTAAGTAG NB36 ATGTCCCAGTTAGAGGAGGAAACA TGTTTCCTCCTCTAACTGGGACAT NB37 GCTTGCGATTGATGCTTAGTATCA TGATACTAAGCATCAATCGCAAGC NB38 ACCACAGGAGGACGATACAGAGAA TTCTCTGTATCGTCCTCCTGTGGT NB39 CCACAGTGTCAACTAGAGCCTCTC GAGAGGCTCTAGTTGACACTGTGG NB40 TAGTTTGGATGACCAAGGATAGCC GGCTATCCTTGGTCATCCAAACTA NB41 GGAGTTCGTCCAGAGAAGTACACG CGTGTACTTCTCTGGACGAACTCC NB42 CTACGTGTAAGGCATACCTGCCAG CTGGCAGGTATGCCTTACACGTAG NB43 CTTTCGTTGTTGACTCGACGGTAG CTACCGTCGAGTCAACAACGAAAG NB44 AGTAGAAAGGGTTCCTTCCCACTC GAGTGGGAAGGAACCCTTTCTACT NB45 GATCCAACAGAGATGCCTTCAGTG CACTGAAGGCATCTCTGTTGGATC NB46 GCTGTGTTCCACTTCATTCTCCTG CAGGAGAATGAAGTGGAACACAGC NB47 GTGCAACTTTCCCACAGGTAGTTC GAACTACCTGTGGGAAAGTTGCAC NB48 CATCTGGAACGTGGTACACCTGTA TACAGGTGTACCACGTTCCAGATG NB49 ACTGGTGCAGCTTTGAACATCTAG CTAGATGTTCAAAGCTGCACCAGT NB50 ATGGACTTTGGTAACTTCCTGCGT ACGCAGGAAGTTACCAAAGTCCAT NB51 GTTGAATGAGCCTACTGGGTCCTC GAGGACCCAGTAGGCTCATTCAAC NB52 TGAGAGACAAGATTGTTCGTGGAC GTCCACGAACAATCTTGTCTCTCA NB53 AGATTCAGACCGTCTCATGCAAAG CTTTGCATGAGACGGTCTGAATCT NB54 CAAGAGCTTTGACTAAGGAGCATG CATGCTCCTTAGTCAAAGCTCTTG NB55 TGGAAGATGAGACCCTGATCTACG CGTAGATCAGGGTCTCATCTTCCA NB56 TCACTACTCAACAGGTGGCATGAA TTCATGCCACCTGTTGAGTAGTGA NB57 GCTAGGTCAATCTCCTTCGGAAGT ACTTCCGAAGGAGATTGACCTAGC NB58 CAGGTTACTCCTCCGTGAGTCTGA TCAGACTCACGGAGGAGTAACCTG NB59 TCAATCAAGAAGGGAAAGCAAGGT ACCTTGCTTTCCCTTCTTGATTGA NB60 CATGTTCAACCAAGGCTTCTATGG CCATAGAAGCCTTGGTTGAACATG NB61 AGAGGGTACTATGTGCCTCAGCAC GTGCTGAGGCACATAGTACCCTCT NB62 CACCCACACTTACTTCAGGACGTA TACGTCCTGAAGTAAGTGTGGGTG NB63 TTCTGAAGTTCCTGGGTCTTGAAC GTTCAAGACCCAGGAACTTCAGAA NB64 GACAGACACCGTTCATCGACTTTC GAAAGTCGATGAACGGTGTCTGTC NB65 TTCTCAGTCTTCCTCCAGACAAGG CCTTGTCTGGAGGAAGACTGAGAA NB66 CCGATCCTTGTGGCTTCTAACTTC GAAGTTAGAAGCCACAAGGATCGG NB67 GTTTGTCATACTCGTGTGCTCACC GGTGAGCACACGAGTATGACAAAC NB68 GAATCTAAGCAAACACGAAGGTGG CCACCTTCGTGTTTGCTTAGATTC NB69 TACAGTCCGAGCCTCATGTGATCT AGATCACATGAGGCTCGGACTGTA NB70 ACCGAGATCCTACGAATGGAGTGT ACACTCCATTCGTAGGATCTCGGT NB71 CCTGGGAGCATCAGGTAGTAACAG CTGTTACTACCTGATGCTCCCAGG NB72 TAGCTGACTGTCTTCCATACCGAC GTCGGTATGGAAGACAGTCAGCTA NB73 AAGAAACAGGATGACAGAACCCTC GAGGGTTCTGTCATCCTGTTTCTT NB74 TACAAGCATCCCAACACTTCCACT AGTGGAAGTGTTGGGATGCTTGTA NB75 GACCATTGTGATGAACCCTGTTGT ACAACAGGGTTCATCACAATGGTC NB76 ATGCTTGTTACATCAACCCTGGAC GTCCAGGGTTGATGTAACAAGCAT NB77 CGACCTGTTTCTCAGGGATACAAC GTTGTATCCCTGAGAAACAGGTCG NB78 AACAACCGAACCTTTGAATCAGAA TTCTGATTCAAAGGTTCGGTTGTT NB79 TCTCGGAGATAGTTCTCACTGCTG CAGCAGTGAGAACTATCTCCGAGA NB80 CGGATGAACATAGGATAGCGATTC GAATCGCTATCCTATGTTCATCCG NB81 CCTCATCTTGTGAAGTTGTTTCGG CCGAAACAACTTCACAAGATGAGG NB82 ACGGTATGTCGAGTTCCAGGACTA TAGTCCTGGAACTCGACATACCGT NB83 TGGCTTGATCTAGGTAAGGTCGAA TTCGACCTTACCTAGATCAAGCCA NB84 GTAGTGGACCTAGAACCTGTGCCA TGGCACAGGTTCTAGGTCCACTAC NB85 AACGGAGGAGTTAGTTGGATGATC GATCATCCAACTAACTCCTCCGTT NB86 AGGTGATCCCAACAAGCGTAAGTA TACTTACGCTTGTTGGGATCACCT NB87 TACATGCTCCTGTTGTTAGGGAGG CCTCCCTAACAACAGGAGCATGTA NB88 TCTTCTACTACCGATCCGAAGCAG CTGCTTCGGATCGGTAGTAGAAGA NB89 ACAGCATCAATGTTTGGCTAGTTG CAACTAGCCAAACATTGATGCTGT NB90 GATGTAGAGGGTACGGTTTGAGGC GCCTCAAACCGTACCCTCTACATC NB91 GGCTCCATAGGAACTCACGCTACT AGTAGCGTGAGTTCCTATGGAGCC NB92 TTGTGAGTGGAAAGATACAGGACC GGTCCTGTATCTTTCCACTCACAA NB93 AGTTTCCATCACTTCAGACTTGGG CCCAAGTCTGAAGTGATGGAAACT NB94 GATTGTCCTCAAACTGCCACCTAC GTAGGTGGCAGTTTGAGGACAATC NB95 CCTGTCTGGAAGAAGAATGGACTT AAGTCCATTCTTCTTCCAGACAGG NB96 CTGAACGGTCATAGAGTCCACCAT ATGGTGGACTCTATGACCGTTCAG




