-
Targeted sequencing
In some sequencing applications, the focus of study — a single gene, or a selection of genomic regions — comprises a small fraction of the genome/sample. In these cases, characterisation through whole-genome sequencing can be inefficient and costly. Targeted sequencing is a term used to describe strategies that reduce the time spent sequencing regions that are not of interest, thereby significantly reducing the amount of data required to achieve the desired depth of the regions of interest. As well as reducing sequencing cost, this reduces the data analysis burden and enables a quicker workflow. Targeted sequencing using nanopore technology can be achieved in several ways:
- amplicon sequencing
- pull-down
- Cas9-based enrichment
- adaptive sampling (AS)
-
Advantages of nanopore sequencing for a targeted approach
- Using methods such as Cas9-based enrichment and adaptive sampling allows enriching for regions inaccessible to traditional technologies, e.g. repetitive or GC-rich regions that cannot be amplified with PCR
- Characterisation of very large regions of the interest: it is possible to enrich and sequence targets spanning tens of kilobases or more in single reads
- Base modifications are retained without the need for any further library preparation
- Information-rich data: detection of single nucleotide variants (SNVs), structural variants, repeat expansions and base modifications in a single sequencing run
- On-demand sequencing, rapid access to results, simple end-to-end workflows and minimal start-up costs
-
Introduction to adaptive sampling
Adaptive sampling offers a fast and flexible method to enrich regions of interest by depleting off-target regions: target selection takes place during sequencing itself, with no requirement for upfront sample manipulation. The library is prepared and loaded as normal and “adaptive sampling” is selected in MinKNOW (typically a .bed file detailing target regions will need to be uploaded). Once the run commences, sequencing will begin and due to the real-time nature of nanopore sequencing, it is possible to identify whether or not the strand that is being sequenced is within the region of interest (ROI). If the read does not map to the ROI, MinKNOW will reverse the polarity of the applied potential, ejecting the strand from the pore so it is able to accept a new, different template strand. Off-target strands are continually rejected until a strand from the ROI is detected and sequencing is allowed to proceed.
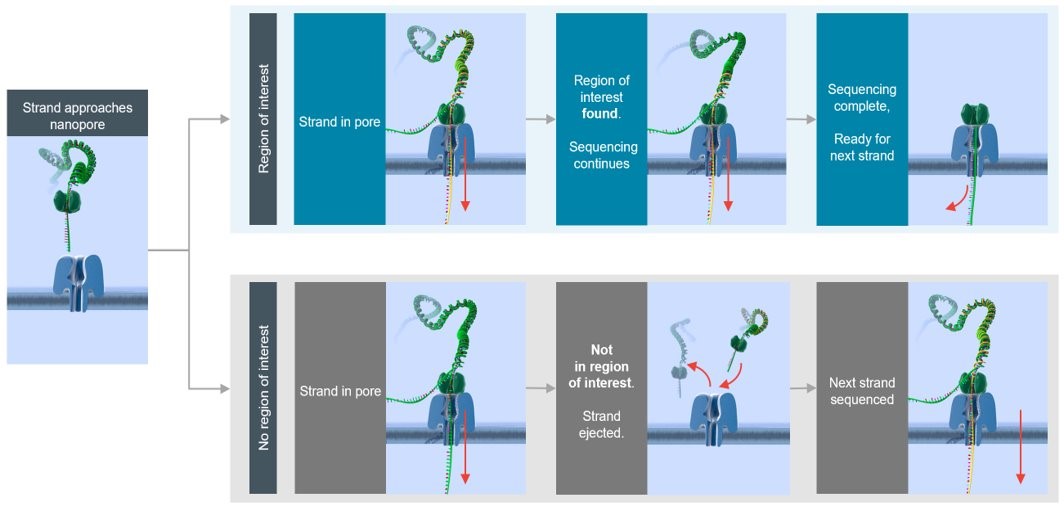
Adaptive sampling can run in two different modes: “enrichment” and “depletion”. In “enrichment”, ROIs are uploaded to MinKNOW and strands that fall outside of this are rejected. In “depletion” mode, targets that are not of interest (e.g. host DNA in a host:microbiome metagenomic analysis) are uploaded to MinKNOW and strands that fall within these regions are rejected. In this document, we discuss “enrichment” strategies only. We generally observe an enrichment for ROI of ~5-10-fold when targeting using AS, and the sections below outline our advice on how this can be achieved. For targeting regions within human genomes, we find this level of enrichment is robust as long as the total fraction that is being targeted is <10% of the total genome, and can enable users to obtain a mean depth >20-40x of ROI on a MinION flow cell.
