- Materials
-
- 1200 ng gDNA per sample
- Multiplex Ligation Sequencing Kit XL (SQK-MLK111.96-XL)
- Consumables
-
- NEB Blunt/TA Ligase Master Mix (NEB, M0367)
- NEBNext® Quick Ligation Reaction Buffer (NEB, B6058)
- NEBNext FFPE Repair Mix (NEB, M6630)
- NEBNext® Ultra™ II End Repair/dA-Tailing Module (E7546)
- NEBNext Quick Ligation Module (NEB, E6056)
- Nuclease-free water (e.g. ThermoFisher, cat #AM9937)
- Freshly prepared 80% ethanol in nuclease-free water
- 1.5 ml Eppendorf DNA LoBind tubes
- Agencourt AMPure XP beads (Beckman Coulter™ cat # A63881)
- Qubit™ Assay Tubes (Invitrogen, Q32856)
- Qubit dsDNA HS Assay Kit (ThermoFisher, cat # Q32851)
- Hamilton 50 µl CO-RE tips with filter (Cat# 235948)
- Hamilton 300 µl CO-RE tips with filter (Cat# 235903)
- Hamilton 1000 µl CO-RE tips with filter (Cat# 235905)
- Hamilton PCR ComfortLid (Cat# 814300)
- Hamilton 60 ml Reagent Reservoir, Self-Standing with Lid (Cat# 56694-01)
- Bio-Rad Hard-Shell® 96-Well PCR Plates (Cat# HSP9601)
- Equipment
-
- P100 pipette and tips
- Qubit fluorometer (or equivalent)
- Ice bucket with ice
-
Consumables and equipment quantities:
Consumables No. of consumables for all conditions Hamilton 50 µl CO-RE tips with filter 960 Hamilton 300 µl CO-RE tips with filter 960 Hamilton 1000 µ CO-RE tips with filter 96 Bio-Rad Hard-Shell® 96-well PCR Plate 8 Note: We recommend using Hamilton tips for efficient liquid handling.
-
Reagent quantities:
Full method
Reagents x96 samples 80% ethanol 80 ml AMPure XP Beads 15 ml Nuclease-free water 10 ml Long Fragment Buffer (LFB) 2 bottles Elution Buffer (EB) 1 bottle EDTA 1 vial Native Barcode plate 1 plate Adapter Mix F (AMII F) 1 vial NEBNext FFRE DNA Repair Buffer 130 µl NEBNext FFPE DNA Repair Mix 90 µl Ultra II End Prep Reaction Buffer 130 µl Ultra II End Prep Enzyme Mix 120 µl Blunt/TA Ligase Master Mix 1210 µl NEBNext Quick Ligation Reaction Buffer (5x) 590 µl Quick T4 DNA Ligase 310 µl -
Prepare the reagents as follows and store on ice.
Reagent 1. Thaw at room temperature 2. Briefly spin down NEBNext FFPE DNA Repair Buffer ✓ - NEBNext FFPE DNA Repair Mix ✓ Note: Do no vortex NEBNext Ultra II End-prep repair buffer ✓ - NEBNext Ultra II End-prep enzyme mix ✓ Note: Do not vortex NEBNext Quick Ligation Reaction Buffer (5x) ✓ - NEBNext Quick T4 DNA Ligase ✓ Note: Do not vortex Native barcode plate ✓ ✓ Elution Buffer (EB) ✓ - Adapter Mix II F (AMII-F) ✓ ✓ Long Fragment Buffer (LFB) ✓ - -
In a clean hard shell PCR plate, prepare the sample input plate as follows:
- Dispense 1200 ng DNA into each sample well. Note: We suggest aliquoting your DNA at 80 ng/µl per sample.
- Make up the volume of each well containing DNA samples to at least 15 µl.
-
Switch on the Hamilton NGS STAR 96 robot and open the method from the desktop shortcut.
-
When the method is loaded, click 'Start'.
To find further information, click 'About MLK111.96-XL' to view the automation section of the Community in the default web browser.

-
Click 'MLK111.96-XL' to proceed to the method parameter selection.
-
Optional actionBefore starting, a user ID can be entered for traceability purposes.
Note: Any format of user ID can be used.
-
Enter the barcode of the input plate containing the samples and the output plate which will contain the prepared DNA libraries.
-
Optional actionSelect where to start on a previously used native barcode plate when using 48 or fewer samples.
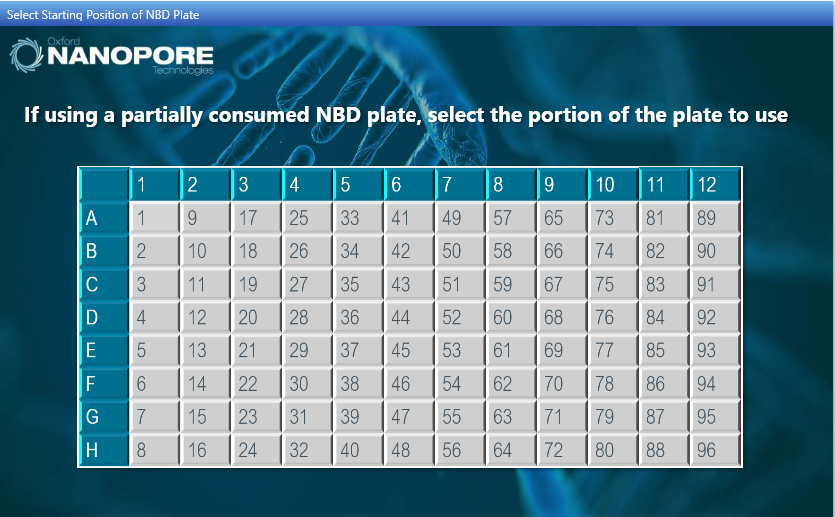
-
Click "Full method" to run the entire automated protocol.
Note: We recommend using the "Select steps" option if sample quantification is required after each step. Please see the "Select steps in the automated library preparation" step.

-
Optional actionFor traceability purposes, the lot barcodes of the reagents can be entered for the Oxford Nanopore Technologies (ONT) reagents, the native barcode plate (NBD) and the NEB reagents used.
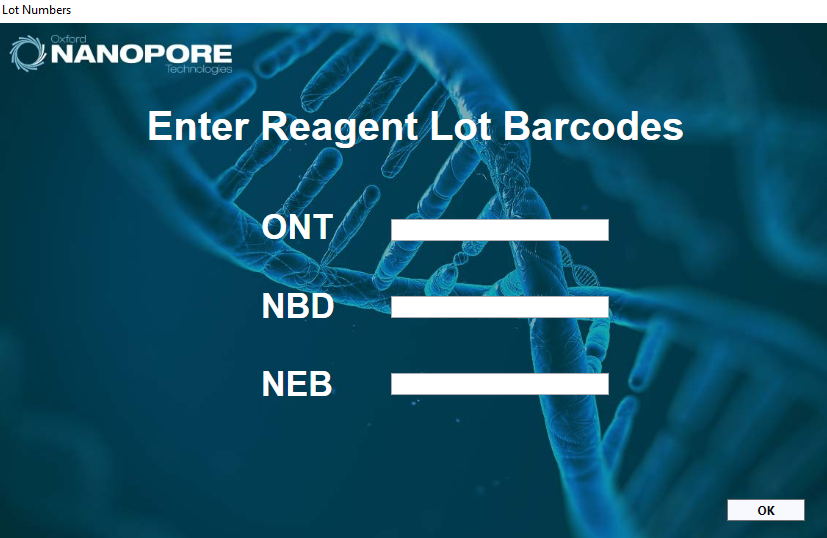

-
Once settings for the run have been selected, there will be a series of dialogues illustrating how to load the deck depending on steps selected.
Note: The following screenshots are an example of performing the full method for x96 samples.
-
Place the Hamilton Comfort PCR lid on the PCR lid position.

-
Load 50 µl tips as indicated on screen.
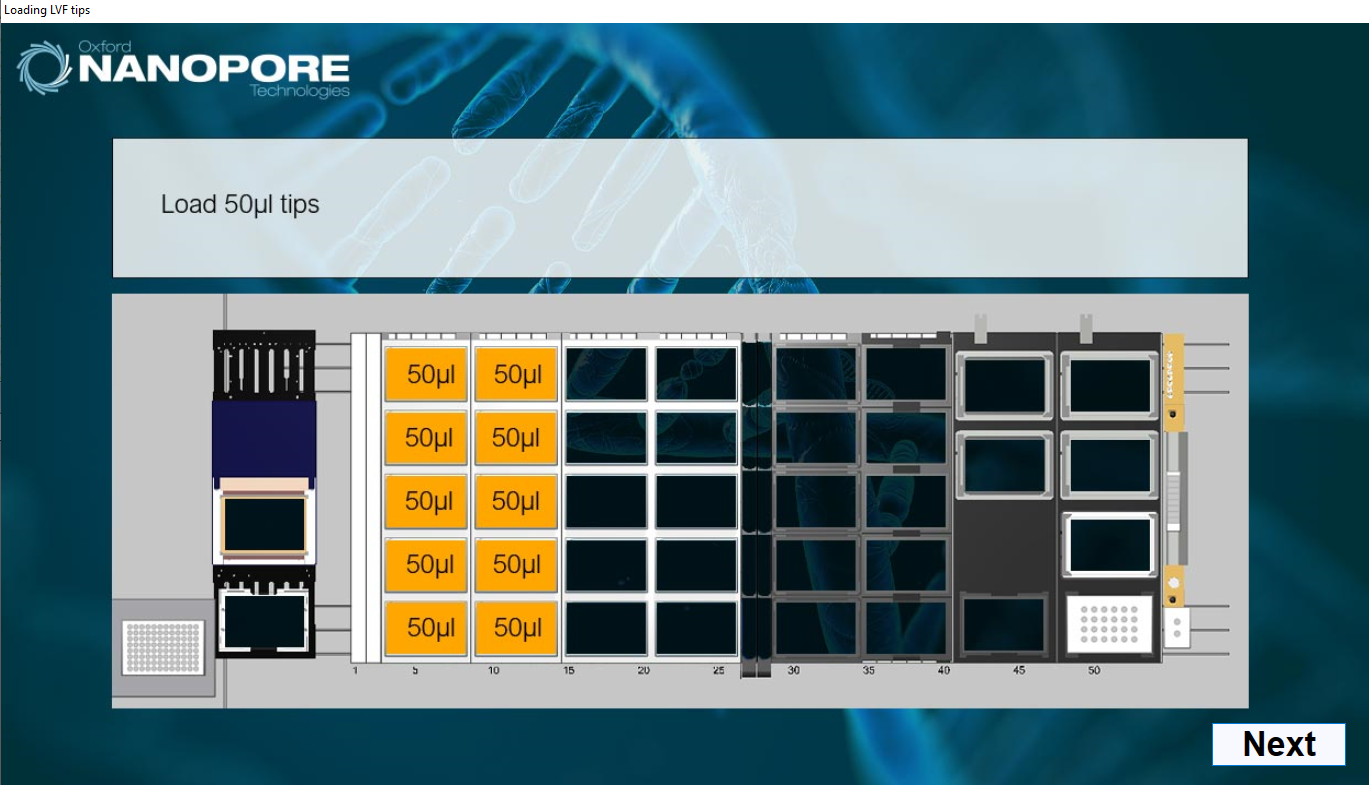
-
Load 300 µl tips as indicated on screen.

-
Mix by inverting and prepare the following reagents in troughs:
For X96 samples:
Reagent Volume per trough No. of troughs Total volume Freshly prepared 80% ethanol 40 ml 2 80 ml Agencourt AMPure XP beads 15 ml 1 15 ml Nuclease-free water 10 ml 1 10 ml Long Fragment Buffer (LFB) 2 bottles 1 2 bottles Elution Buffer (EB) 1 bottle 1 1 bottle -
Load the reagent troughs as indicated on screen.
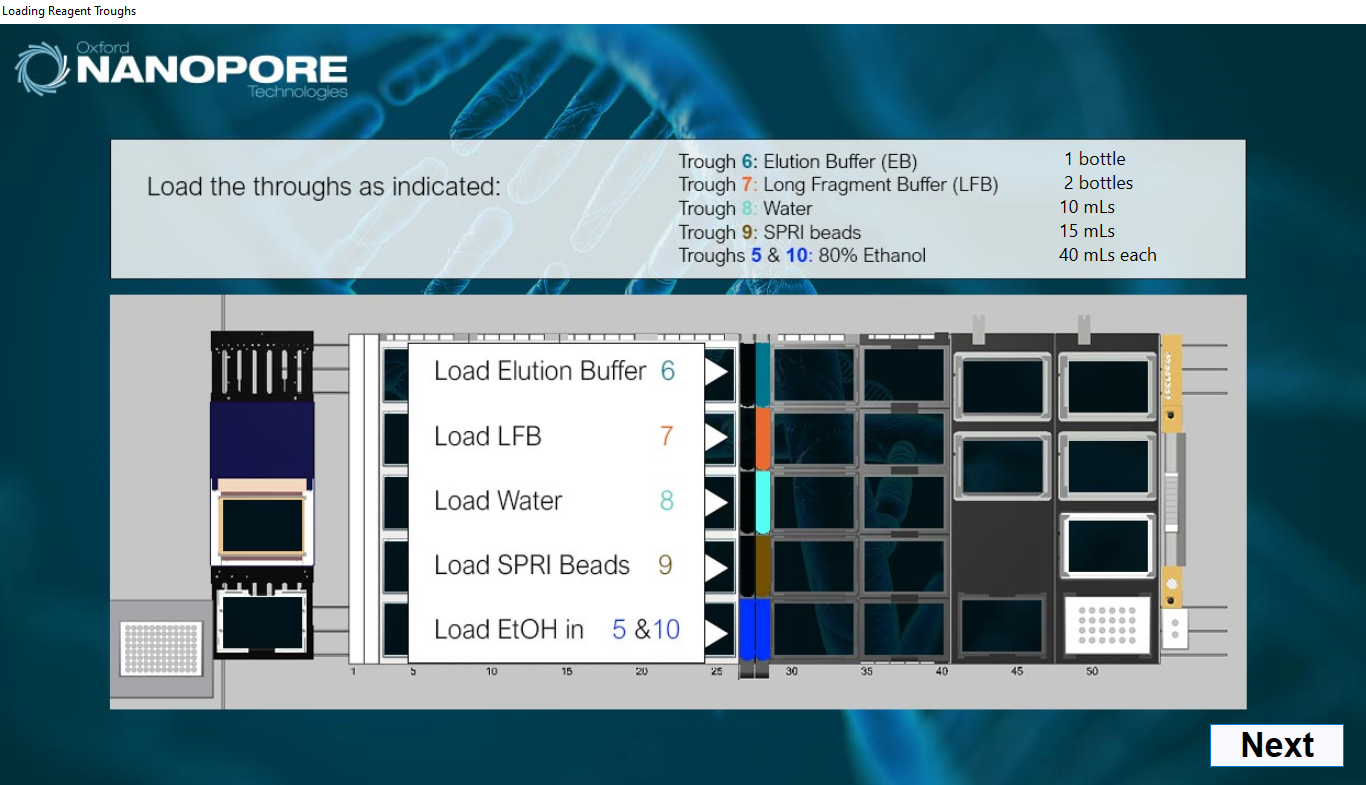
-
Ensure the foil seal is removed from the native barcode plate foil seal.
-
Load the native barcode plate, 3 fresh PCR plates and the input plate containing the DNA samples.
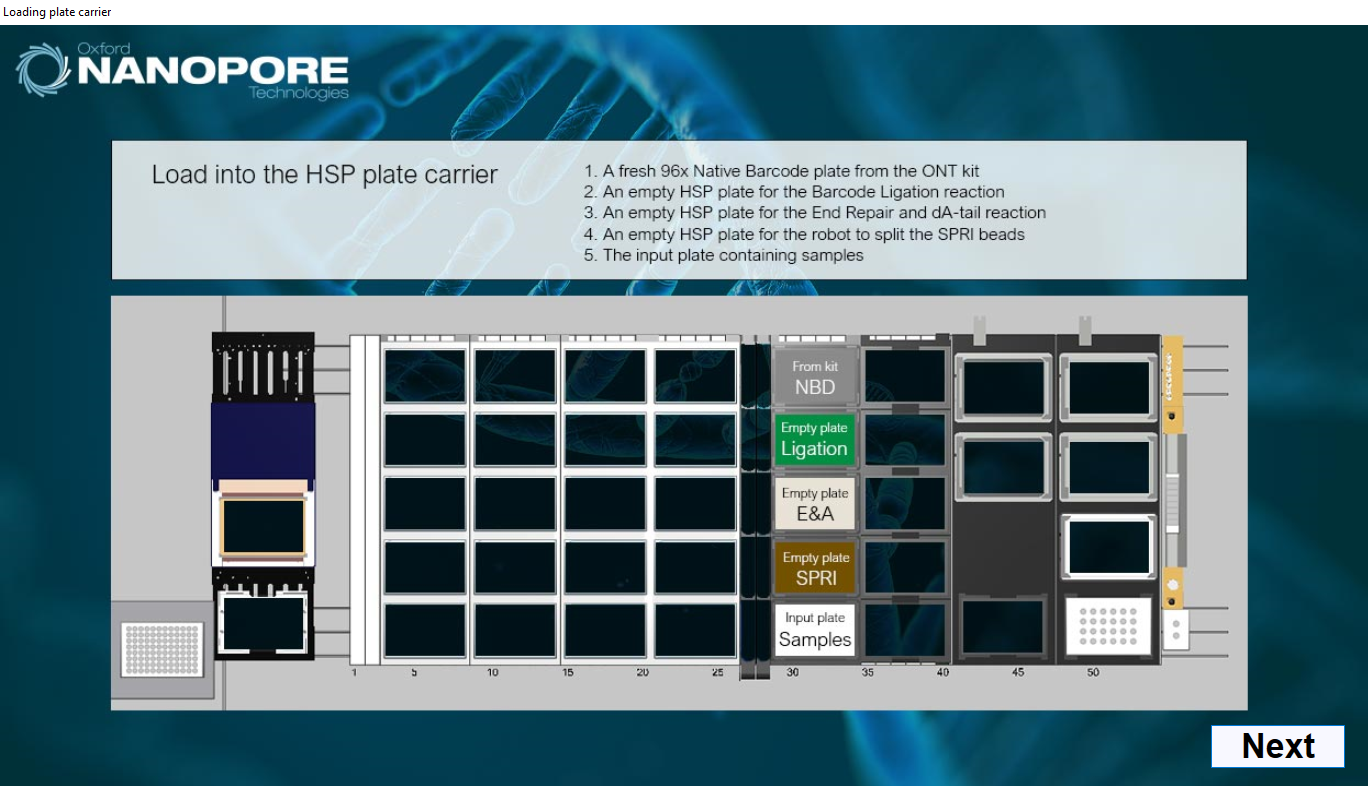

-
Load 1000 µl tips as indicated on screen and ensure the magnet is in place with the adapter for PCR plates.

-
Load the CPAC module as indicated on screen with the reagent tubes before loading on deck.
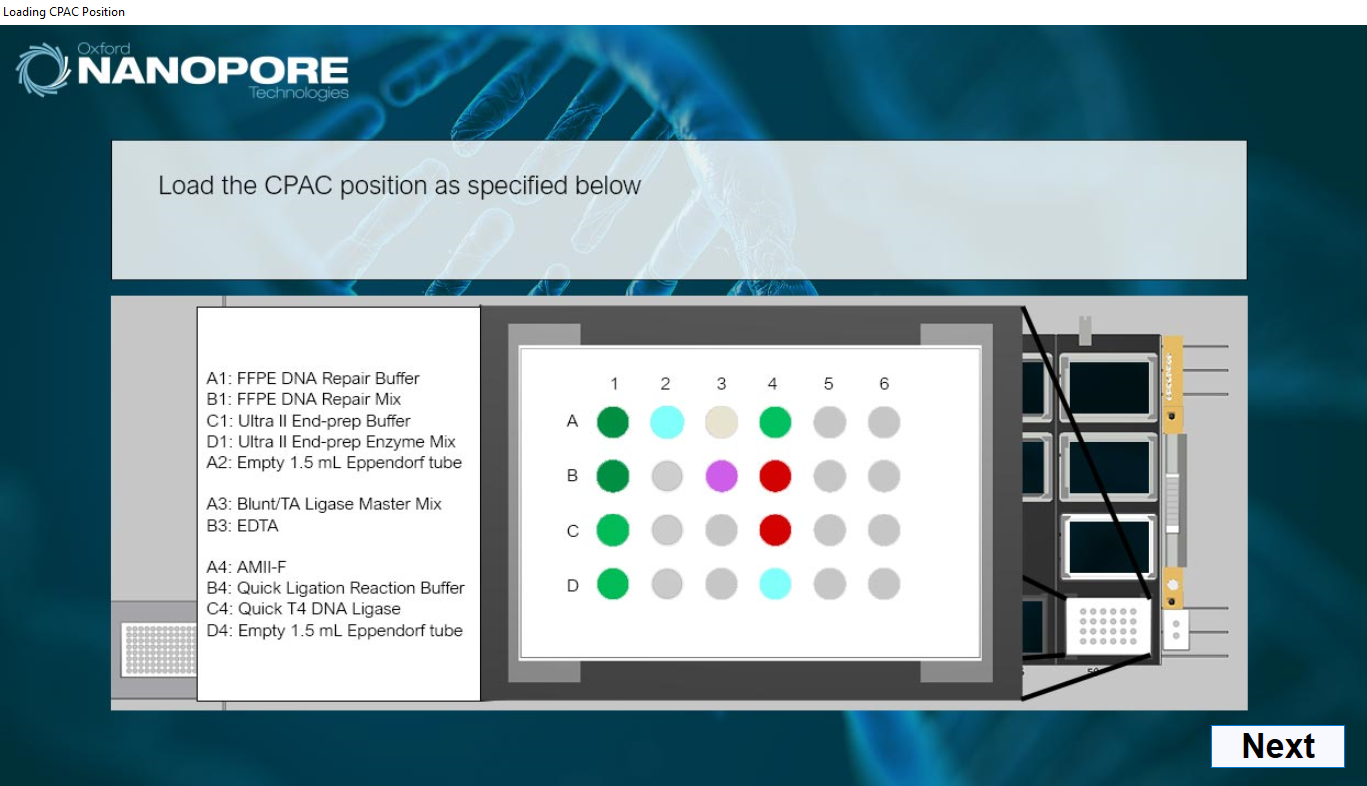

1 2 3 4 A FFPE DNA Repair Buffer Empty 1.5 ml Eppendorf LoBind tube Blunt/TA Ligase Master Mix AMII-F B FFPE DNA Repair Mix - EDTA Quick Ligation Reaction Buffer C Ultra II End-prep Buffer - - Quick T4 DNA Ligase D Ultra II End-prep Enzyme Mix - - Empty 1.5 ml Eppendorf LoBind tube Volumes required:
Reagent x96 samples
3 samples across 2 flow cellsx96 samples
2 samples per flow cellx48 samples
3 samples across 2 flow cellsx48 samples
2 samples per flow cellNEBNext FFPE DNA Repair buffer 130 µl 130 µl 72.45 µl 72.45 µl NEBNext FFPE DNA Repair Enzyme Mix 90 µl 90 µl 41.4 µl 41.4 µl Ultra II End Prep Reaction Buffer 130 µl 130 µl 72.45 µl 72.45 µl Ultra II End Prep Enzyme Mix 120 µl 120 µl 62.1 µl 62.1 µl Blunt/TA Ligase Master Mix 1210 µl 1210 µl 738 µl 738 µl AMII T 270.4 µl 310 µl 138 µl 202.8 µl NEBNext Quick Ligation Reaction Buffer (5x) 540.8 µl 590 µl 276 µl 405.6 µl Quick T4 DNA Ligase 270.4 µl 310 µl 138 µl 202.8 µl -
Once the deck is correctly loaded, click 'Begin method' to start with the parameters selected before loading.
-
During the thermal cycle step for Step 1: End repair and adenylation thermal cycling reaction, the user will be prompted to remove the input plate with the input samples and to load 3 fresh PCR plates as indicated on screen.
This dialogue will prompt the user to remove their input plate:
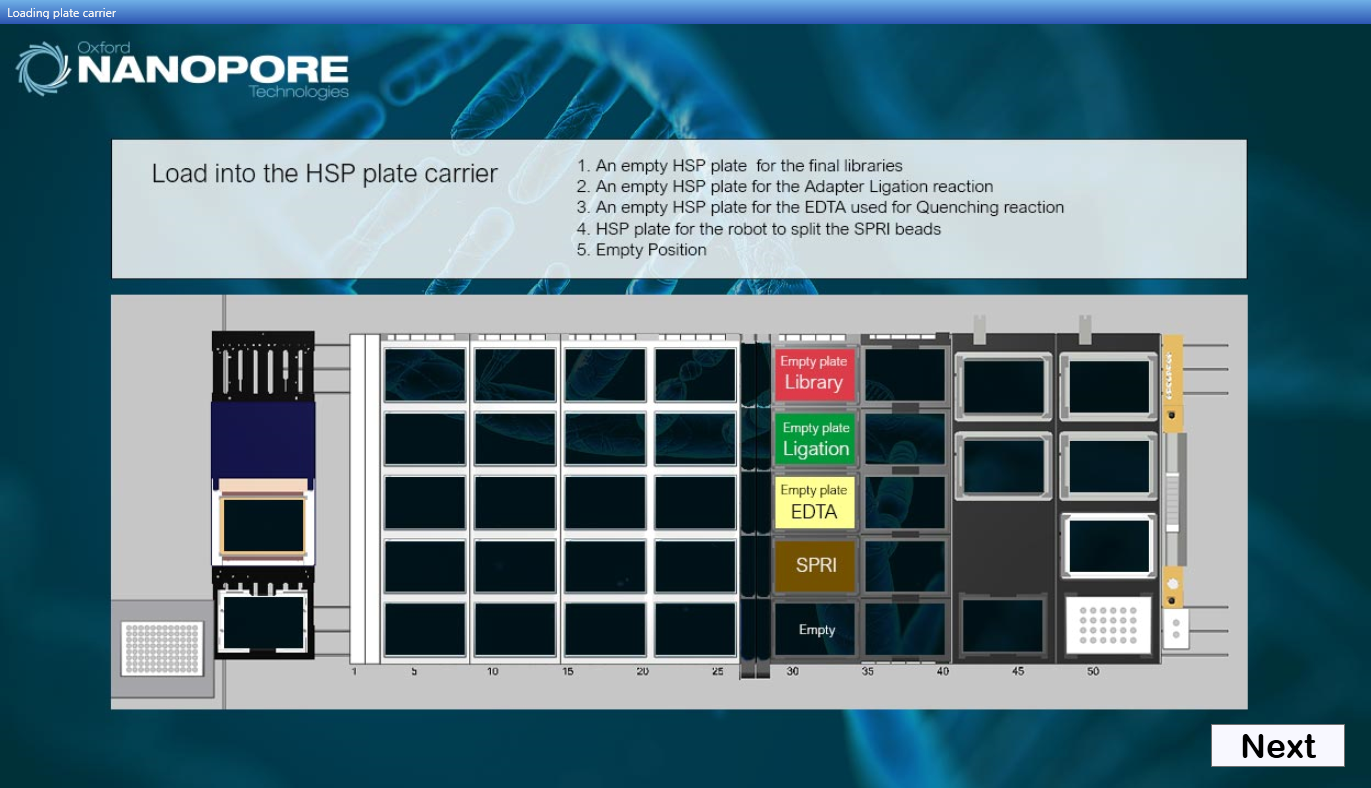

-
Optional actionQuantify 1 µl of eluted sample using a Qubit fluorometer.
-
Optional actionIf quantities allow, the libraries may be diluted in Elution Buffer (EB) for splittling across multiple flow cells.



















