-
Features of using Cas9 targeted sequencing
We recommend CRISPR/Cas targeted sequencing if the user:
- Wishes to sequence multiple human gene targets (up to 100 in a single panel) to high coverage (>100x) on a single MinION Mk1B flow cell
- Wishes to sequence up to a 50 kb Region of Interest (ROI), using up to 100 target sites, in a single assay*
- Has 1-10 µg of available gDNA
- Wishes to gain insight into methylation patterns or other modified bases
- Has gene targets that are highly repetitive, or wishes to evaluate the number of repeats in an expansion, where traditional amplification methods or sequencing-by-synthesis methods could yield a biased result
- Wishes to sequence long gene targets in a single pass that are not amenable to long-range PCR (> 30 kb)
- Optionally wishes to run multiple barcoded samples on a single flow cell.
* Note: This is known as ‘tiling’ an ROI.
-
Excision/single cut vs tiling approach
This protocol can be used for any probe design method (details of which can be found in the Targeted, amplification-free DNA sequencing using CRISPR/Cas info sheet). The recommended ‘excision approach’ and ‘single cut and read out’ method can follow the main flow of this protocol (shown in Figure 1). The ‘tiling’ approach requires the alterations to the protocol described in the Important (orange) boxes in the library preparation section. The main difference with the tiling approach is that both pools of probes need to be prepared separately (RNP complex formation, cleavage and dA-tailing and adapter ligation performed in separate tubes) then pooled during the final AMPure XP bead purification (shown in Figure 2).
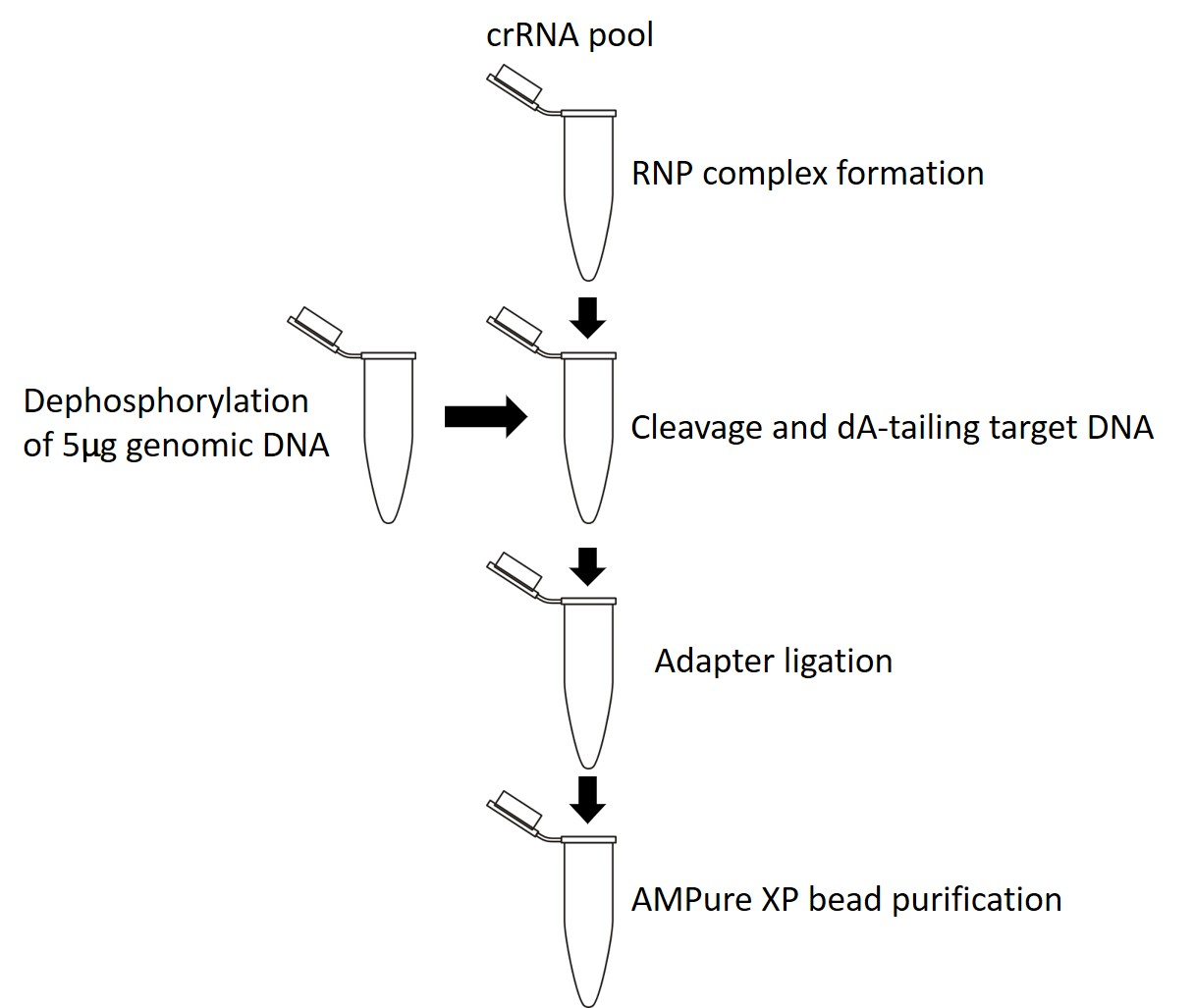
Figure 1. Cas9 targeted sequencing protocol using the 'excision approach' or 'single cut and read out'.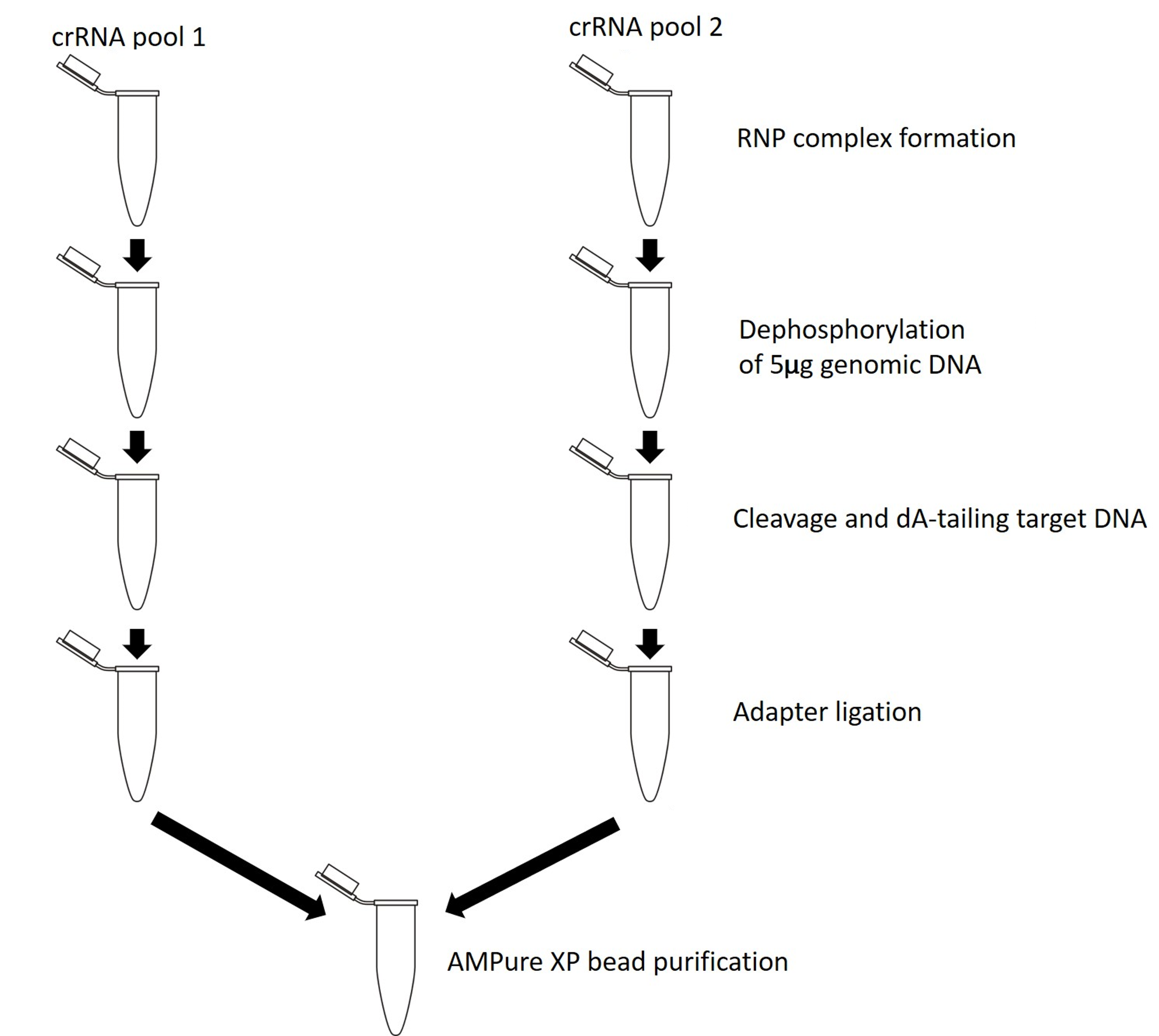
Figure 2. Cas9 targeted sequencing protocol using the 'tiling' approach -
Introduction to the CRISPR/Cas protocol
This protocol describes how to carry out sequencing of genomic DNA using the Cas9 Sequencing Kit (SQK-CS9109) with enrichment of specific genomic regions using CRISPR/Cas.
For users with no previous nanopore sequencing experience, we recommend that a Lambda control experiment is completed first to become familiar with the technology.
Prepare for your experiment
You will need to:
- Extract your DNA, and check its length, quantity and purity.
The quality checks performed during the protocol are essential in ensuring experimental success.
- Ensure you have your sequencing kit, the correct equipment and third-party reagents
- Download the software for acquiring and analysing your data
- Check your flow cell to ensure it has enough pores for a good sequencing runLibrary preparation
Figure 1. shows and explains the biochemical steps used to prepare your barcoded DNA library using a Cas9 Sequencing Kit (SQK-CS9109), plus several third-party reagents.Figure 1. Cas-mediated PCR-free enrichment library preparation for sequencing.
- After DNA extraction, 5’ ends are dephosphorylated to reduce ligation of sequencing adapters to non-target strands.
- Cas9 ribonucleoprotein particles (RNPs), with bound crRNA and tracrRNA, are added to the genomic DNA, then bind and cleave the Region of Interest (ROI).
- dsDNA cleavage by Cas9 reveals blunt ends with ligatable 5’ phosphates.
- All of the DNA in the samples are dA-tailed, which prepares the blunt ends for barcode ligation.
- Sequencing adapters are ligated primarily to Cas9 cut sides, which are both 3’ dA-tailed and 5’ phosphorylated. The library preparation is cleaned up to remove excess adapters using AMPure XP beads and resuspended in Sequencing Buffer. Non-target molecules are not removed. The subsequent library preparation is added to the flow cell for sequencing.
Sequencing and analysis
You will need to:
- Start a sequencing run using the MinKNOW software, which will collect raw data from the device and convert it into basecalled reads
- Start the EPI2ME software and select a workflow for further analysis (this step is optional) -
Enrichment experiment steps and associated instructions
Step Instructions 1. Extract and prepare DNA Extraction methods 2. Design probes Targeted, amplification-free DNA sequencing using CRISPR/Cas - Probe design 3. QC input DNA Input DNA/RNA QC 4. Perform enrichment, and prepare sequencing library Cas Sequencing Kit protocol 5. Sequence on device Cas Sequencing Kit protocol 6. Take basecalled FASTQ files into analysis pipeline Targeted, amplification-free DNA sequencing using CRISPR/Cas - Evaluation of read-mapping characteristics from a Cas9 targeted sequencing experiment 7. Assess success of experiment and feed back into probe design and quality of input

