-
Open the EPI2ME Agent using the desktop shortcut.
-
If prompted, select a folder containing FASTQ files to be analysed.
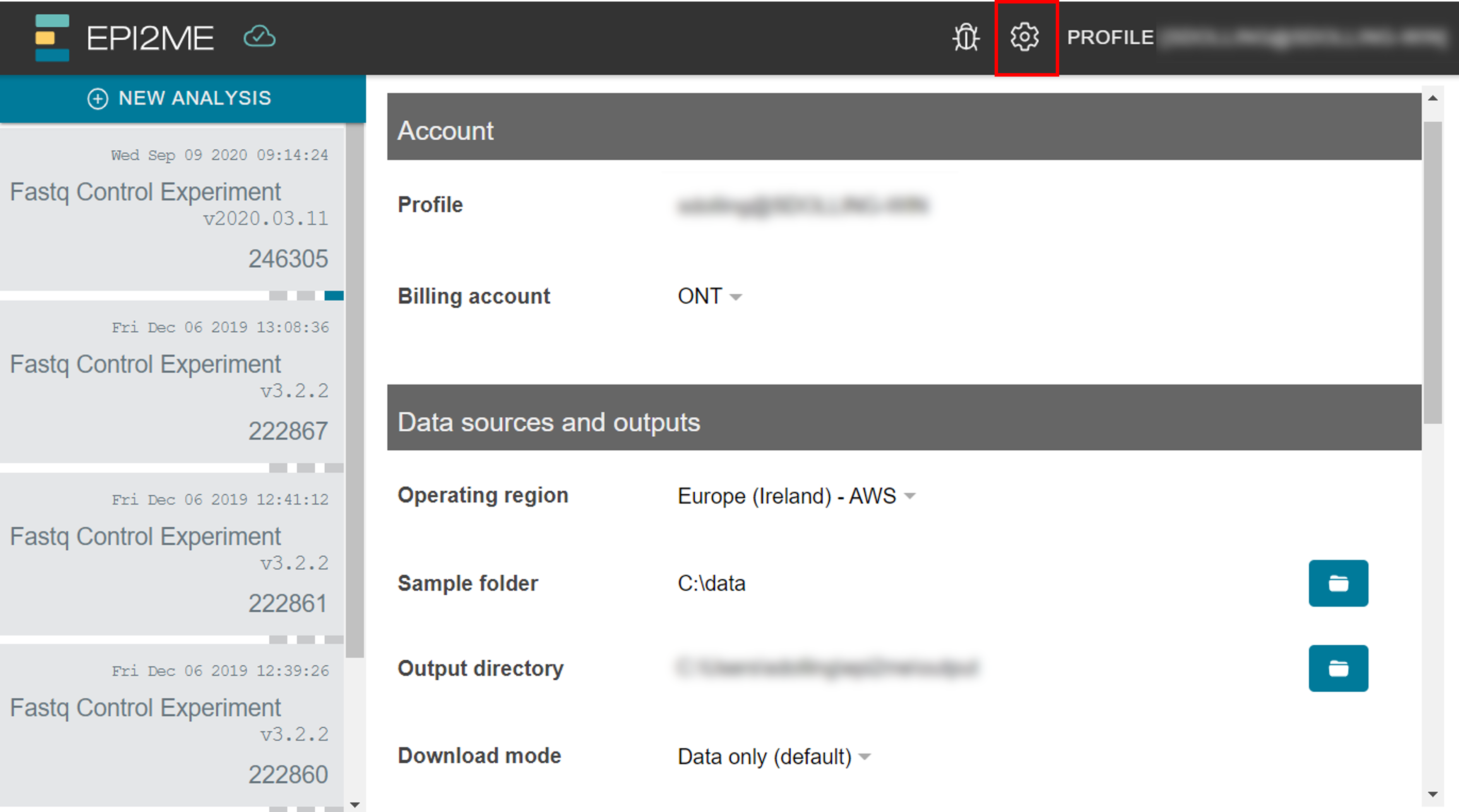
If a MinKNOW-generated folder structure is detected in the default input location, this will be shown in the Experiments & samples tab. You can change the default input location in the Settings page (cog icon) under Data sources. The root of the Output folder can be changed and any output from the analysis will be stored beneath that path as folders (named by instance ID). The root output location can be changed in the Settings page or when adjusting the parameters for the individual analysis.
Alternatively, you will be directed to the Folders tab, where you can select your input folder.
-
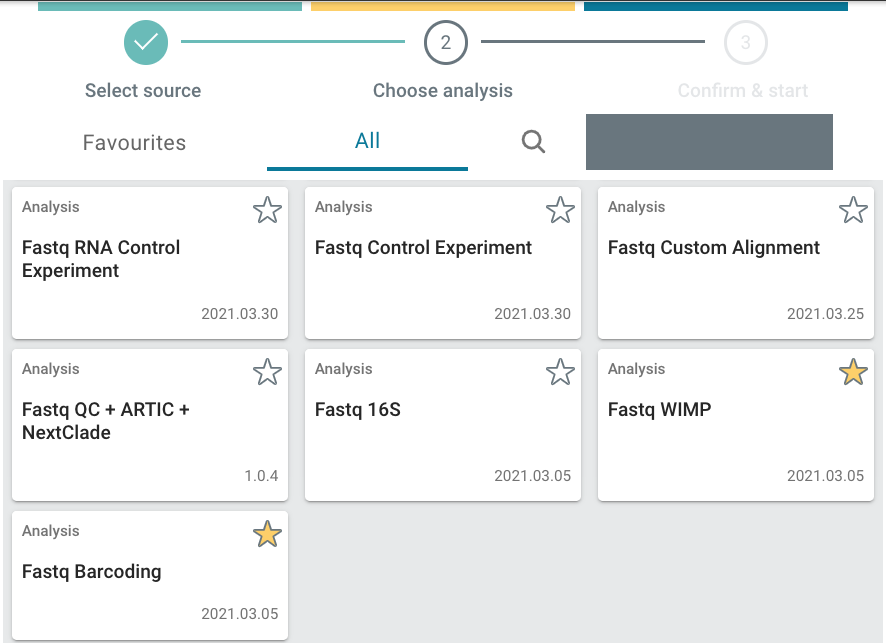
Select the analysis you want to perform.
You can select from the full list of analyses (All tab), or from the Favourites tab. Different user profiles can have different favourite analyses.
-
Adjust the run-time parameters of your analysis.
The parameters available to fine-tune will depend on the type of analysis selected.
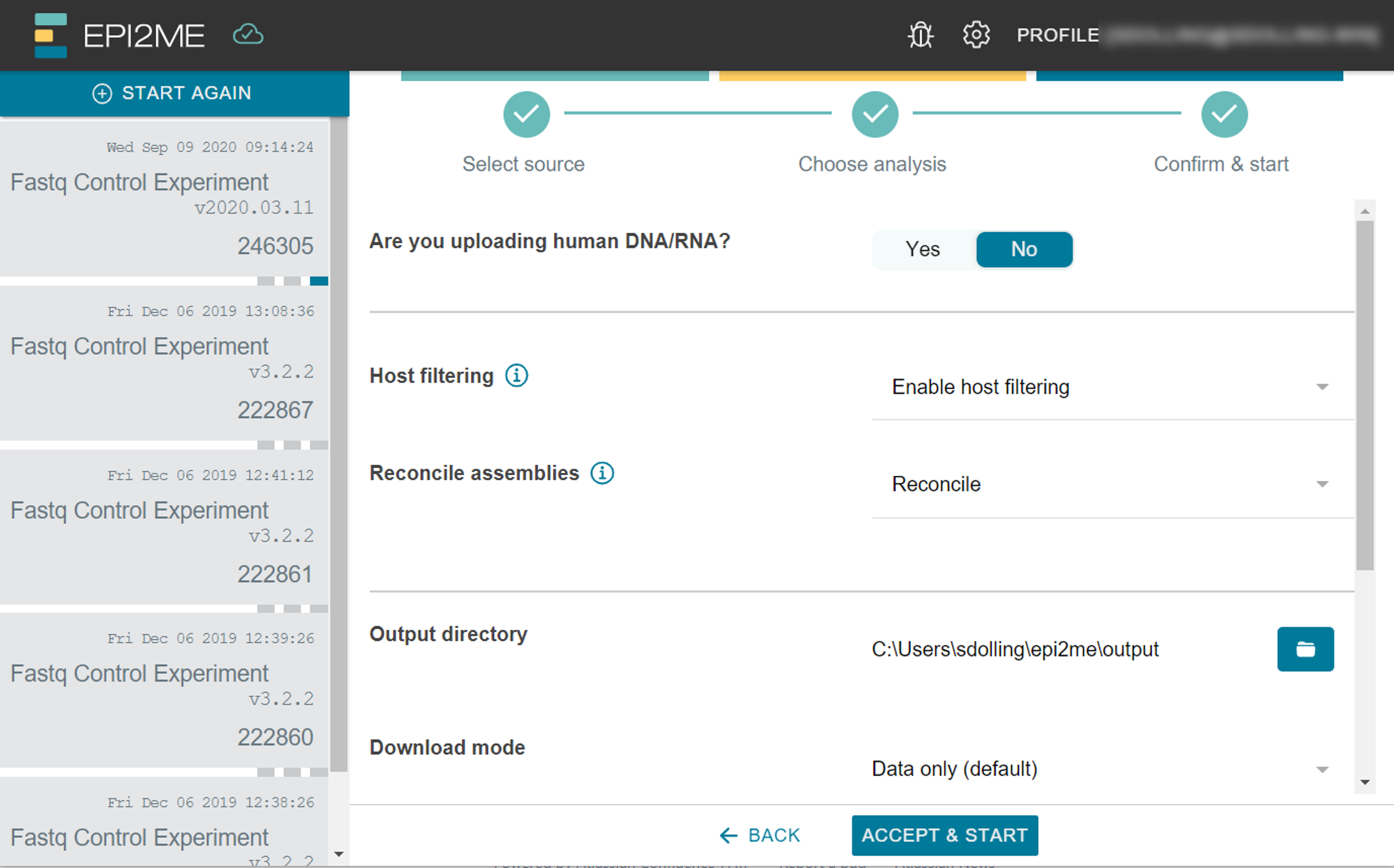
Please note that the input and output folders/locations cannot be the same.
The Download mode option allows you to select which files to download during the analysis:
- No files
- Data only - usually FASTQ and sometimes BAM files. This is set by default
- Telemetry only - metadata about the experiment
- Both data and telemetry
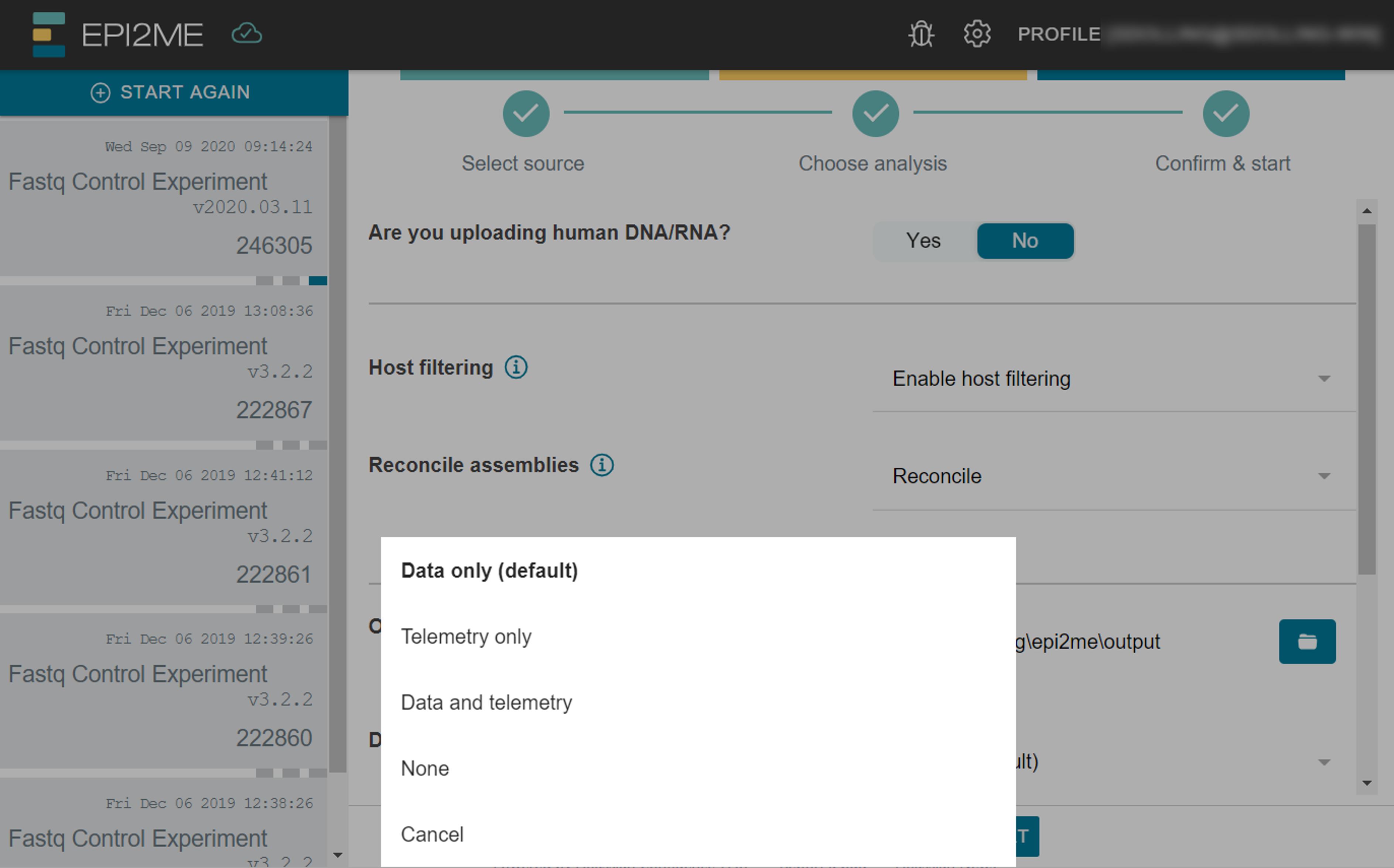
The default download mode can also be changed globally in the Settings page.
-
Once all the required fields are filled in, click "Accept & start" to start data analysis.
-
Follow the progression of upload and download of read files in the Agent.
-
Bug reports
If there is any problem with the EPI2ME Agent or data analysis, you can click the bug icon (shown in the red box) on the interface to send a report.
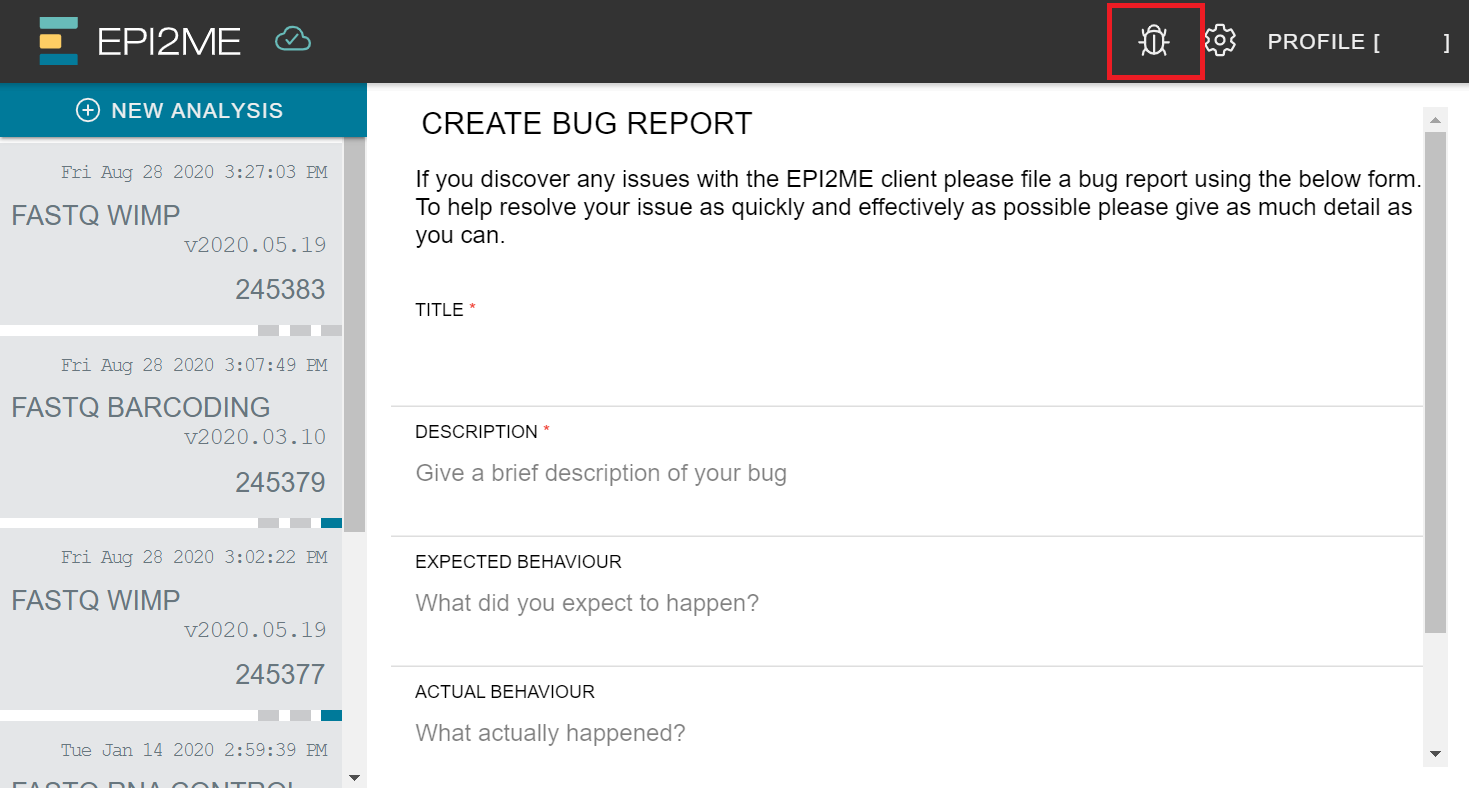
Alternatively, navigate to Report a bug in the EPI2MEAgent application menu. You can also access the log files from here.