- Materials
-
- For Kit 14 libraries, Flow Cell Flush (FCF), Flow Cell Tether (FCT), Library Solution (LIS), Library Beads (LIB), and Sequencing Beads (SB)
- For Kit 9, 10 and 11 libraries, Flush Buffer (FB), Flush Tether (FLT) Loading Beads/II (LB/LBII), Loading Solution (LS), and Sequencing Buffer/II (SQB/SBII),
- AMPure XP Beads (AXP)
- Elution Buffer (EB)
- Short Fragment Buffer (SFB)
- Consumables
-
- MinION Flow Cell (FLO-MIN106 or FLO-MIN114)
- Bovine Serum Albumin (BSA) (50 mg/ml) (e.g Invitrogen™ UltraPure™ BSA 50 mg/ml, AM2616)
- 1.5 ml Eppendorf DNA LoBind tubes
- Qubit™ Assay Tubes (Invitrogen, Q32856)
- Qubit dsDNA HS Assay Kit (Invitrogen, Q32851)
- Equipment
-
- MinION or GridION device
- MinION and GridION Flow Cell Light Shield
- P1000 pipette and tips
- P200 pipette and tips
- Vortex mixer
- Hula mixer (gentle rotator mixer)
- Microfuge
- Magnetic rack
- Heating block
- Qubit fluorometer (or equivalent)
-
Preparation to clean up a library before transfer to a second flow cell
- The aim is to recover and clean up your library before priming the second flow cell for loading the recovered library at a later date.
- The cleaned up library can be stored at 4°C for short term storage or repeated use, for example, re-loading flow cells between washes.
- Data acquisition in MinKNOW should be stopped during the recovery and transfer procedure.
-
Thaw the kit components at room temperature and prepare as indicated by the table below:
Reagent 1. Thaw at room temperature 2. Mix well by vortexing 3. Briefly spin down 4. Keep on ice AMPure XP Beads (AXP) ✓ ✓ X X Keep at room temperature Short Fragment Buffer (SFB) ✓ ✓ ✓ ✓ Elution Buffer (EB) ✓ ✓ ✓ ✓ AMPure XP Beads from an Oxford Nanopore sequencing kit require thawing as they are stored with the kit at -20°C.
-
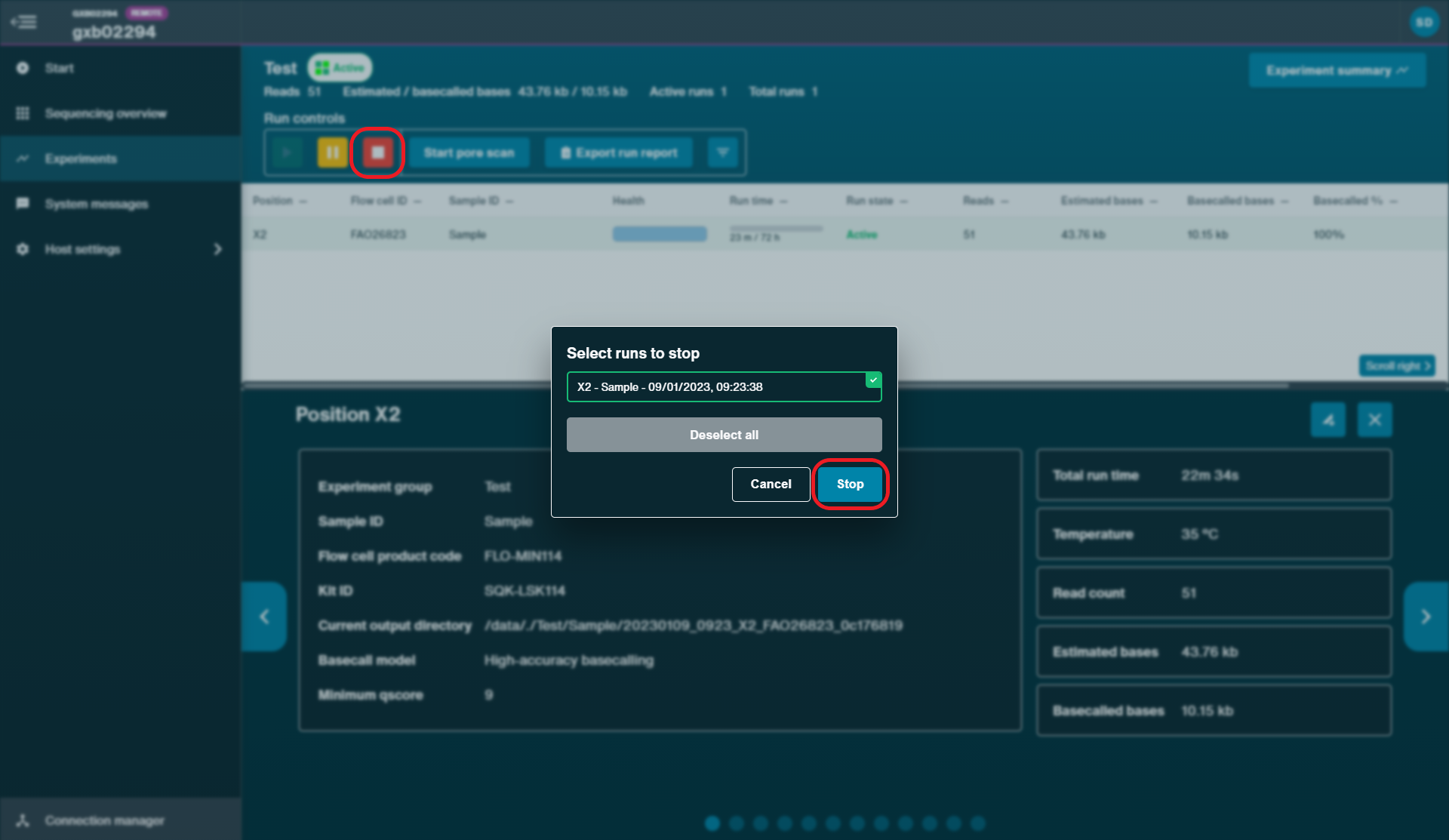
Stop the sequencing run for the original flow cell on MinKNOW by clicking 'Stop'.
-
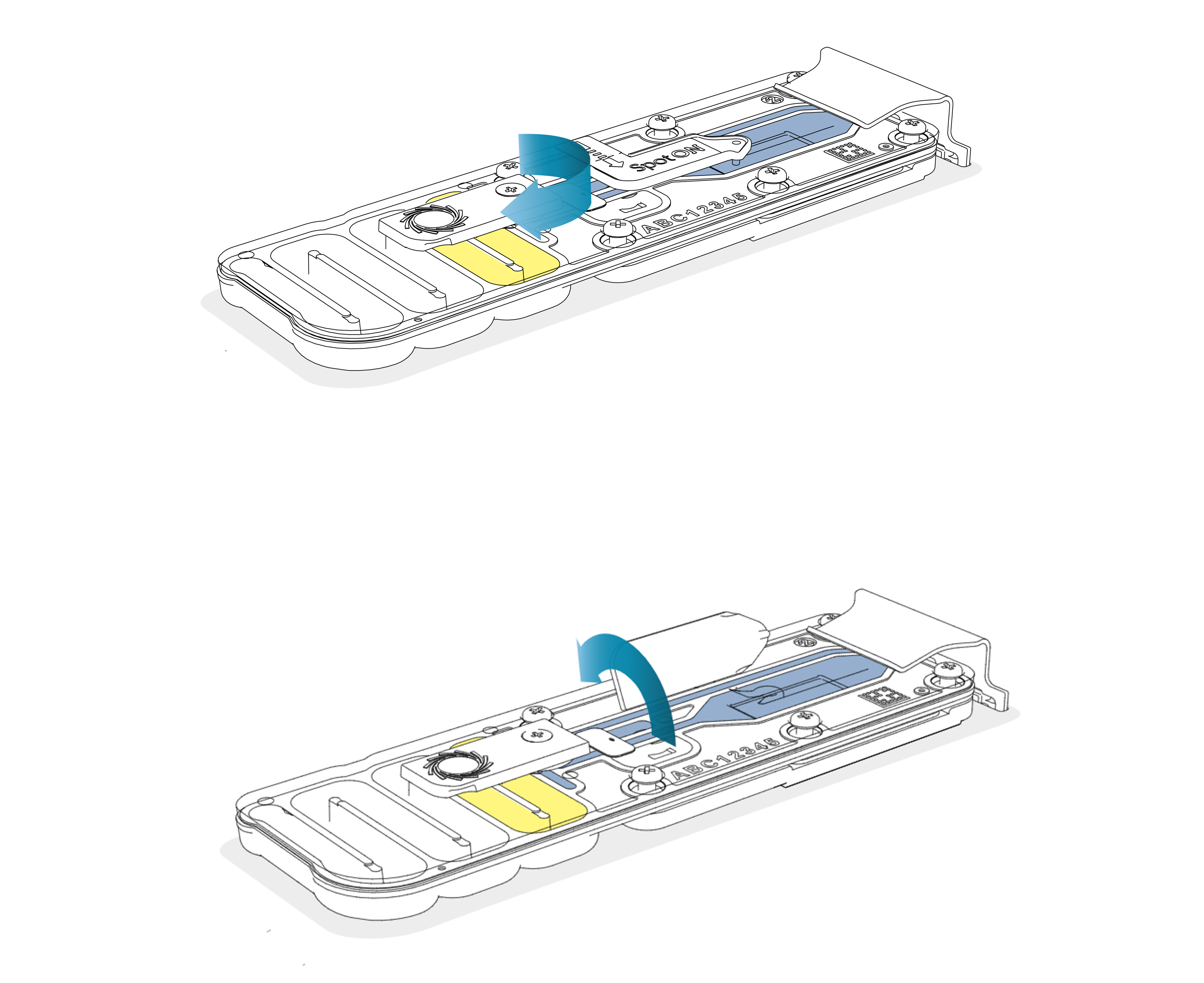
To prepare the original flow cell for library recovery, slide open the priming port cover and lift open the SpotON sample port cover.
-
Set a pipette to 75 µl and fully depress the pipette before inserting the tip into the SpotON port of the original flow cell. Slowly aspirate to recover the DNA library from the flow cell.
Note: Insert the pipette tip to the point where it is touching the liquid in the flow cell. Do not insert the tip too far into the port as this will impact removal.
The recovered library may appear slightly yellow and not all the library loading beads (LB, LBII, LIB) will be fully recovered but this will not impact library recovery and transfer.
-
Transfer the recovered library to a fresh 1.5 ml Eppendorf DNA LoBind tube and store on ice.
-
The original flow cell can be removed from the MinION or GridION device by sliding the flow cell from under the clip.
-
The original flow cell can be flushed with deionised water and returned to Oxford Nanopore.
Instructions for returning flow cells can be found here.
-
Resuspend the AMPure XP Beads (AXP) by vortexing.
-
Add 300 µl of resuspended AMPure XP Beads (AXP) to the recovered library and mix by flicking.
-
Incubate on a Hula mixer (rotator mixer) for 10 minutes at room temperature.
-
Spin down the sample and pellet on a magnet until supernatant is clear and colourless. Keep the tube on the magnet, and pipette off the supernatant.
-
Wash the beads by adding 150 µl of Short Fragment Buffer (SFB). Flick the beads to resuspend, spin down, then return the tube to the magnetic rack and allow the beads to pellet. Remove the supernatant using a pipette and discard.
-
Spin down and place the tube back on the magnet. Pipette off any residual supernatant.
-
Remove the tube from the magnetic rack and resuspend the pellet in 13 µl of Elution Buffer (EB).
-
Spin down and incubate for 10 minutes at room temperature. For high molecular weight DNA, incubating at 37°C can improve recovery of long fragments.
-
Pellet the beads on a magnet until the eluate is clear and colourless, for at least 1 minute.
-
Remove and retain 13 µl of eluate containing the DNA library into a clean 1.5 ml Eppendorf DNA LoBind tube.
Dispose of the pelleted beads
-
Quantify 1 µl of eluted sample using a Qubit fluorometer. If the recovered library is below the detection level of the Qubit dsDNA HS Assay, we do not recommend continuing to load the flow cell.
Note: Library concentration of a recovered library will not be as high as the initial library, but the best results will be achieved by loading as close to the requirements as possible.
-
The library can be stored at 4°C.
-
Thaw and prepare the flow cell priming mix according to the "Priming and loading the SpotON flow cell" section of the suitable protocol.
-
Open the MinION or GridION device lid and slide the second flow cell under the clip. Press down firmly on the flow cell to ensure correct thermal and electrical contact.
-
To prime the second flow cell, slide the priming port cover clockwise to open the priming port.
-
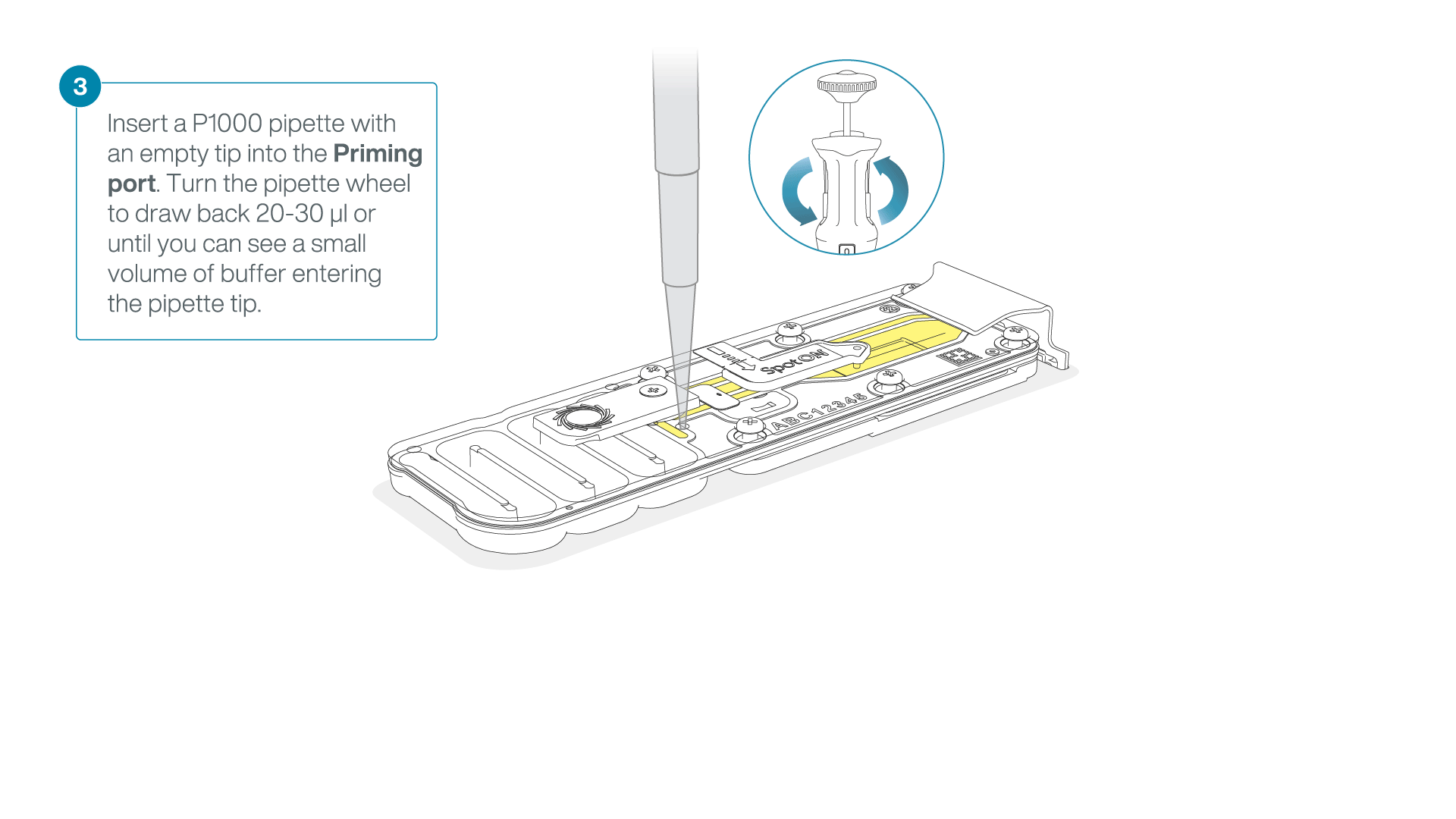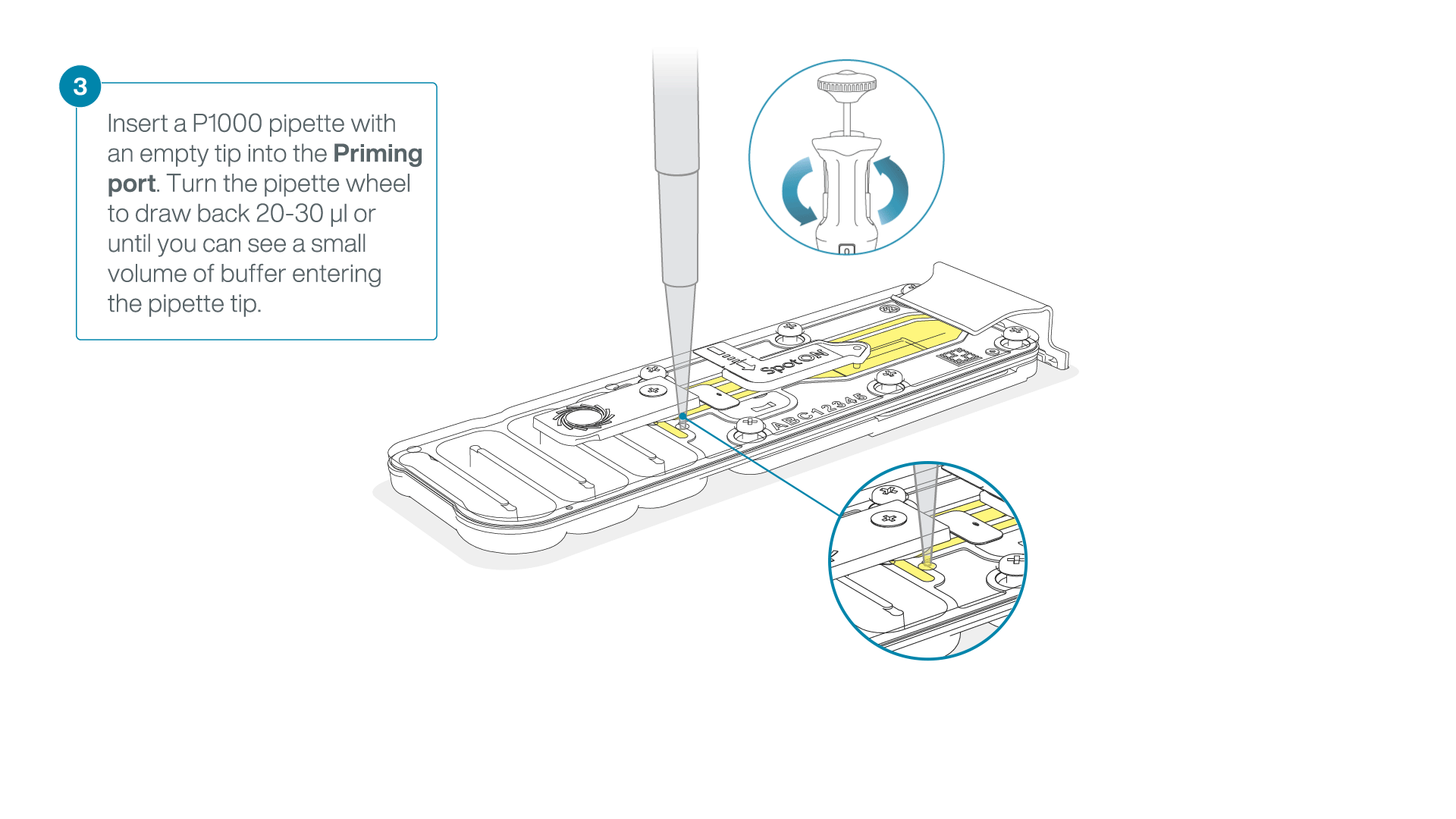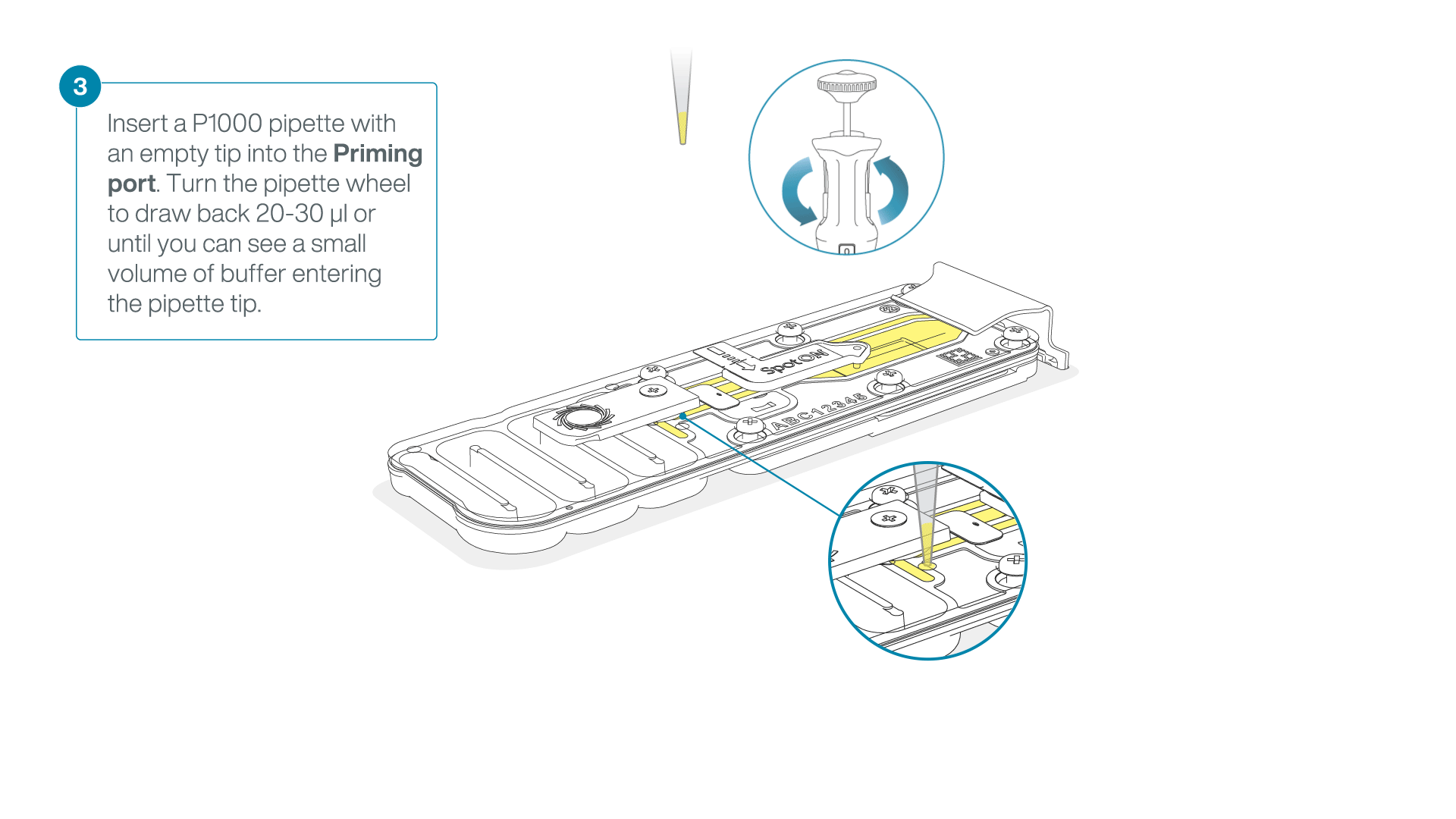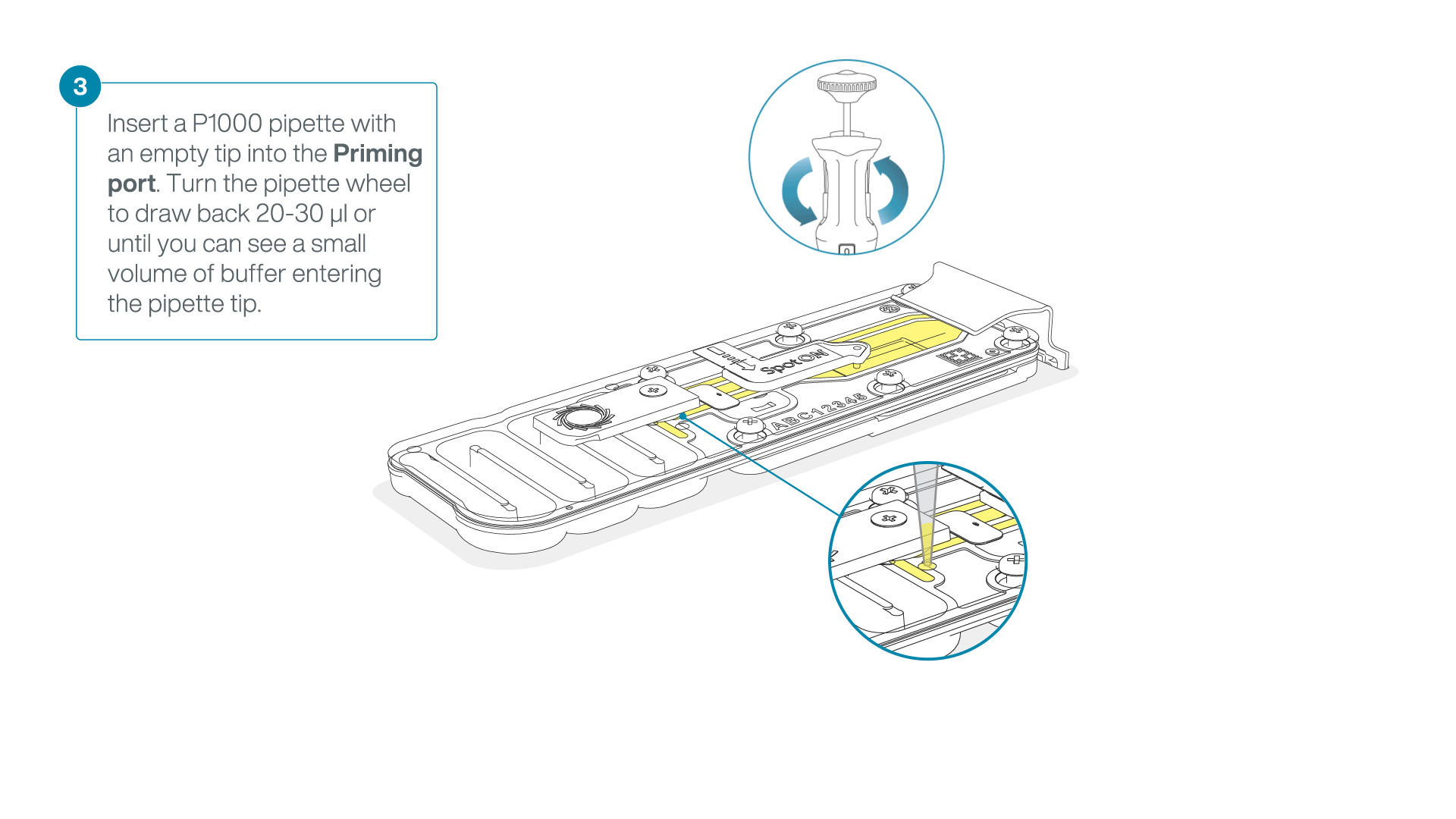
After opening the priming port, check for a small air bubble under the cover. Draw back a small volume to remove any bubbles:
- Set a P1000 pipette to 200 µl
- Insert the tip into the priming port
- Turn the wheel until the dial shows 220-230 µl, to draw back 20-30 µl, or until you can see a small volume of buffer entering the pipette tip
Note: Visually check that there is continuous buffer from the priming port across the sensor array.
-
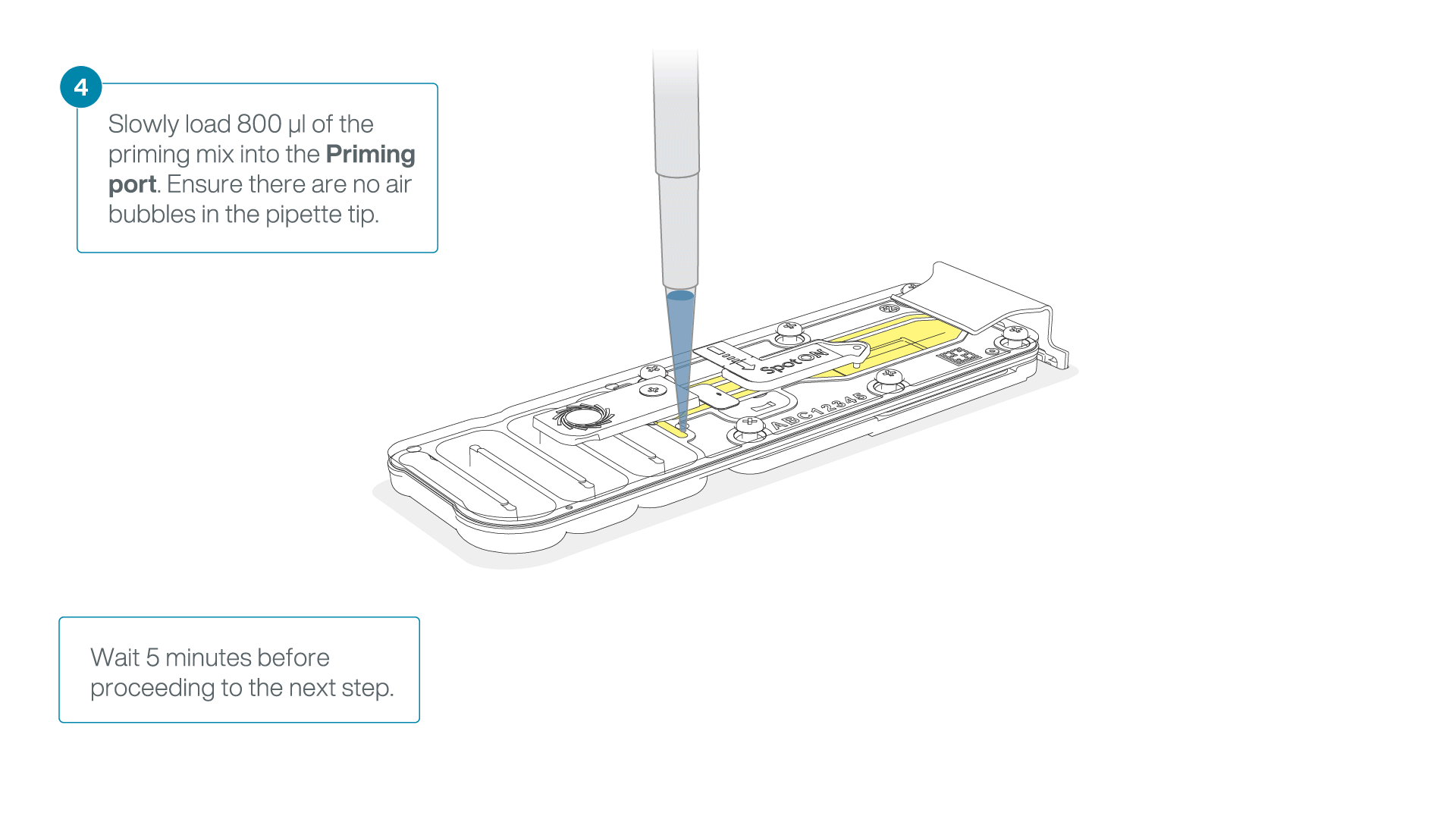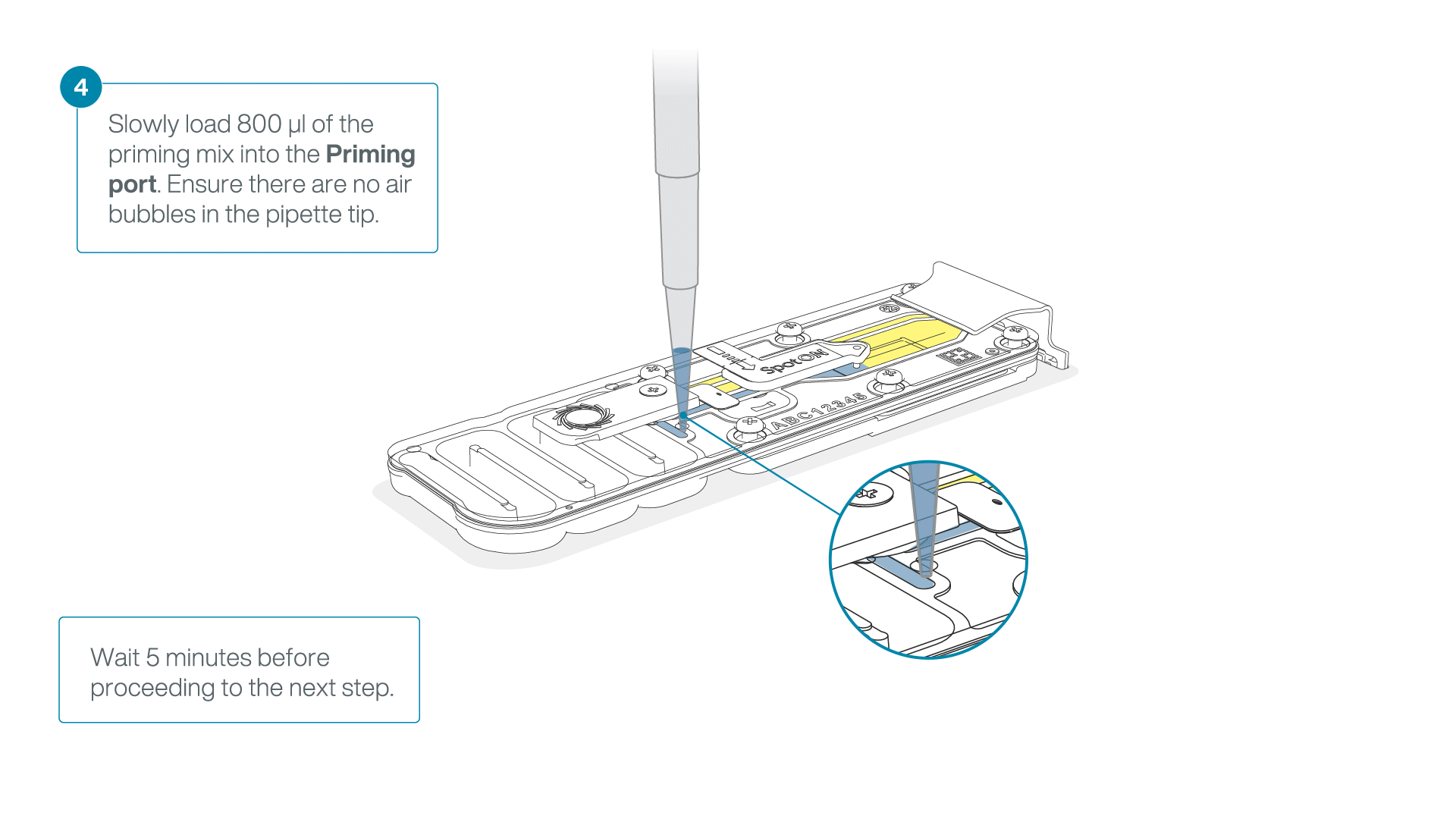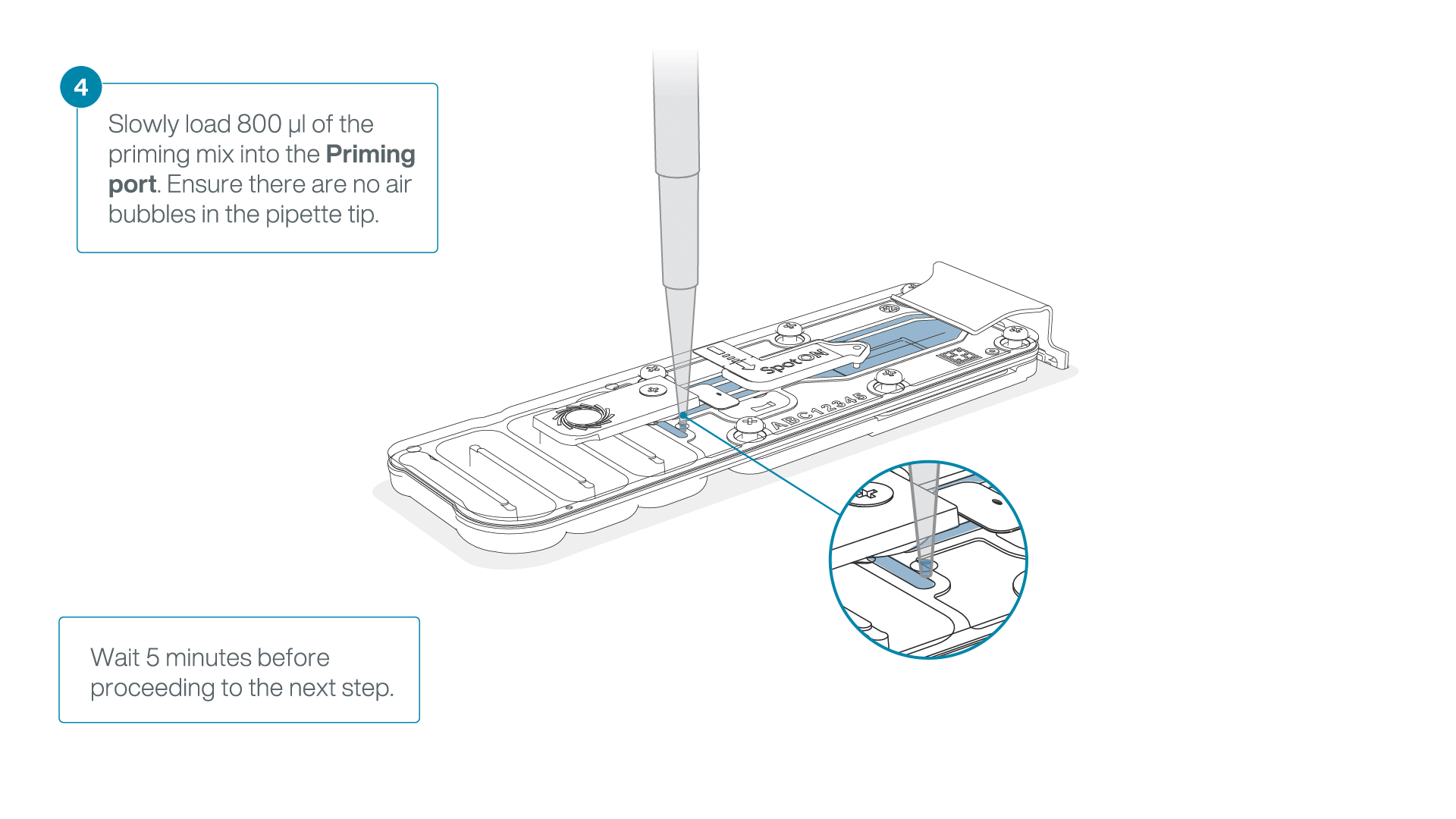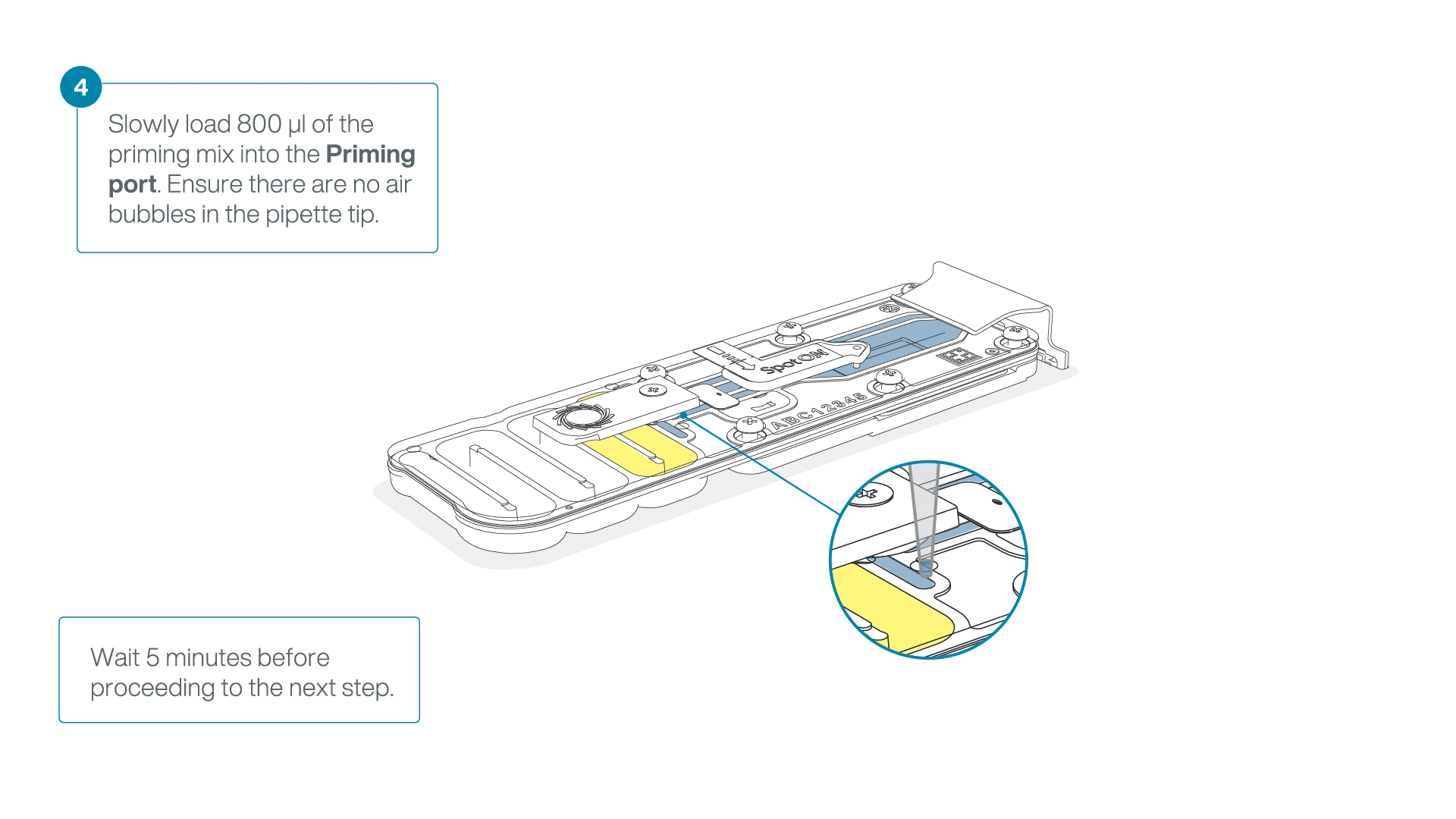
Load 800 µl of the priming mix into the second flow cell via the priming port, avoiding the introduction of air bubbles. Wait for five minutes.
-
Thoroughly mix the contents of the Library Beads/Loading Beads by pipetting.
-
In a new tube, prepare the recovered library for loading according to the "Priming and loading the SpotON flow cell" section of the suitable protocol to ensure you are using the correct reagents and volumes.
For Kit 14 chemistry:
Reagent Volume per flow cell Sequencing Buffer (SB) 37.5 µl Library Beads (LIB) mixed immediately before use, or Library Solution (LIS), if using 25.5 µl Recovered DNA library 12 µl Total 75 µl -
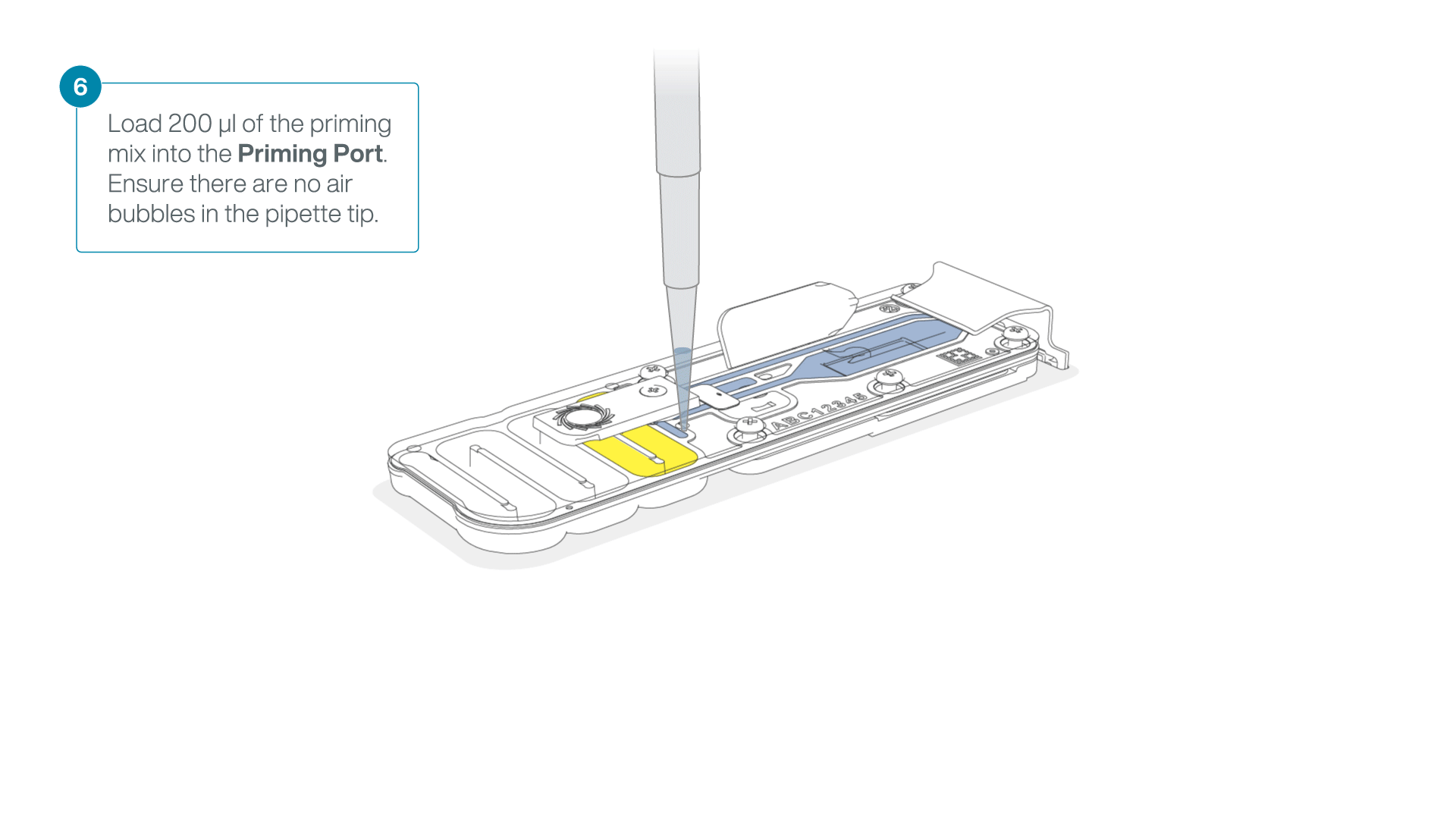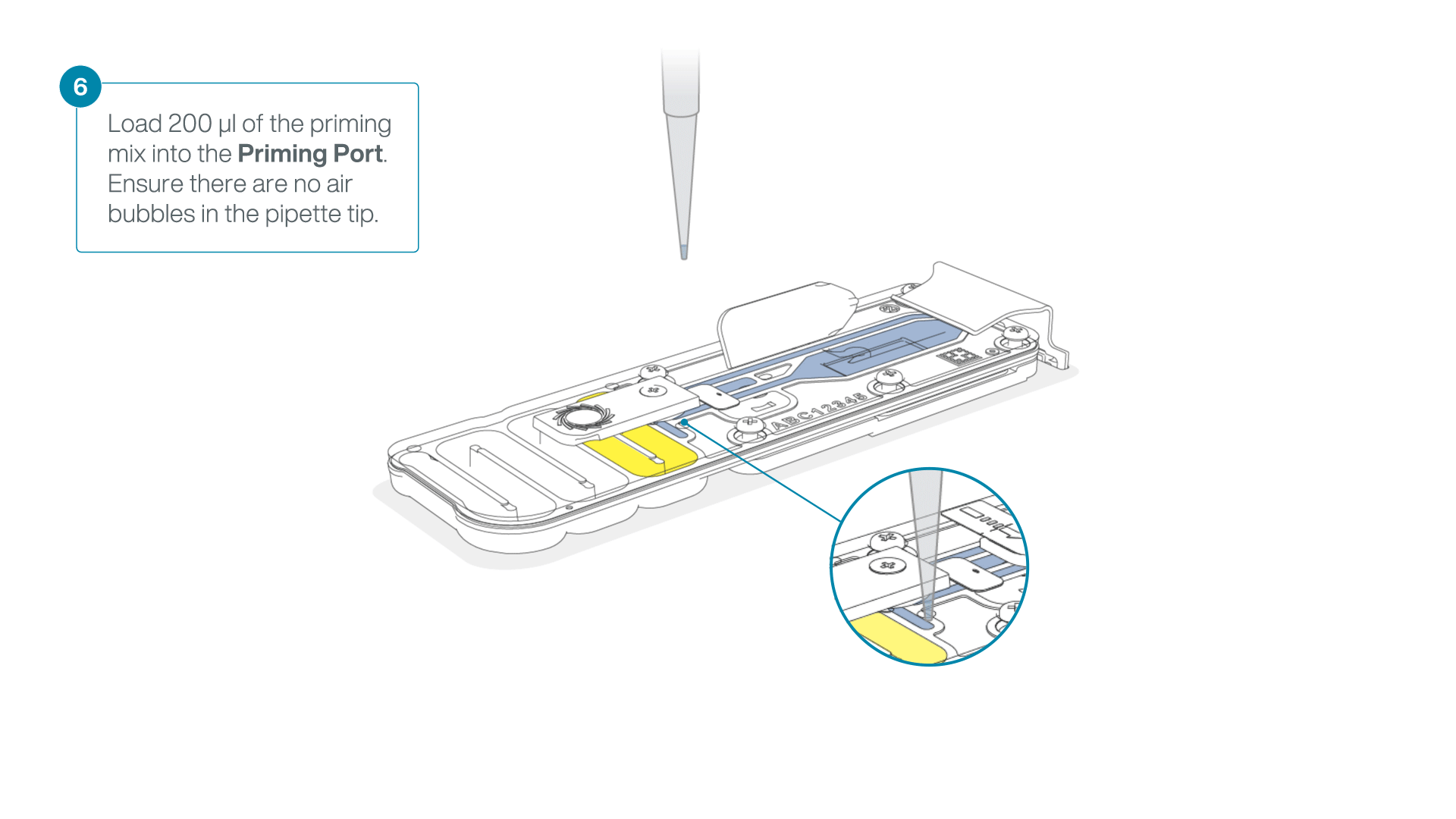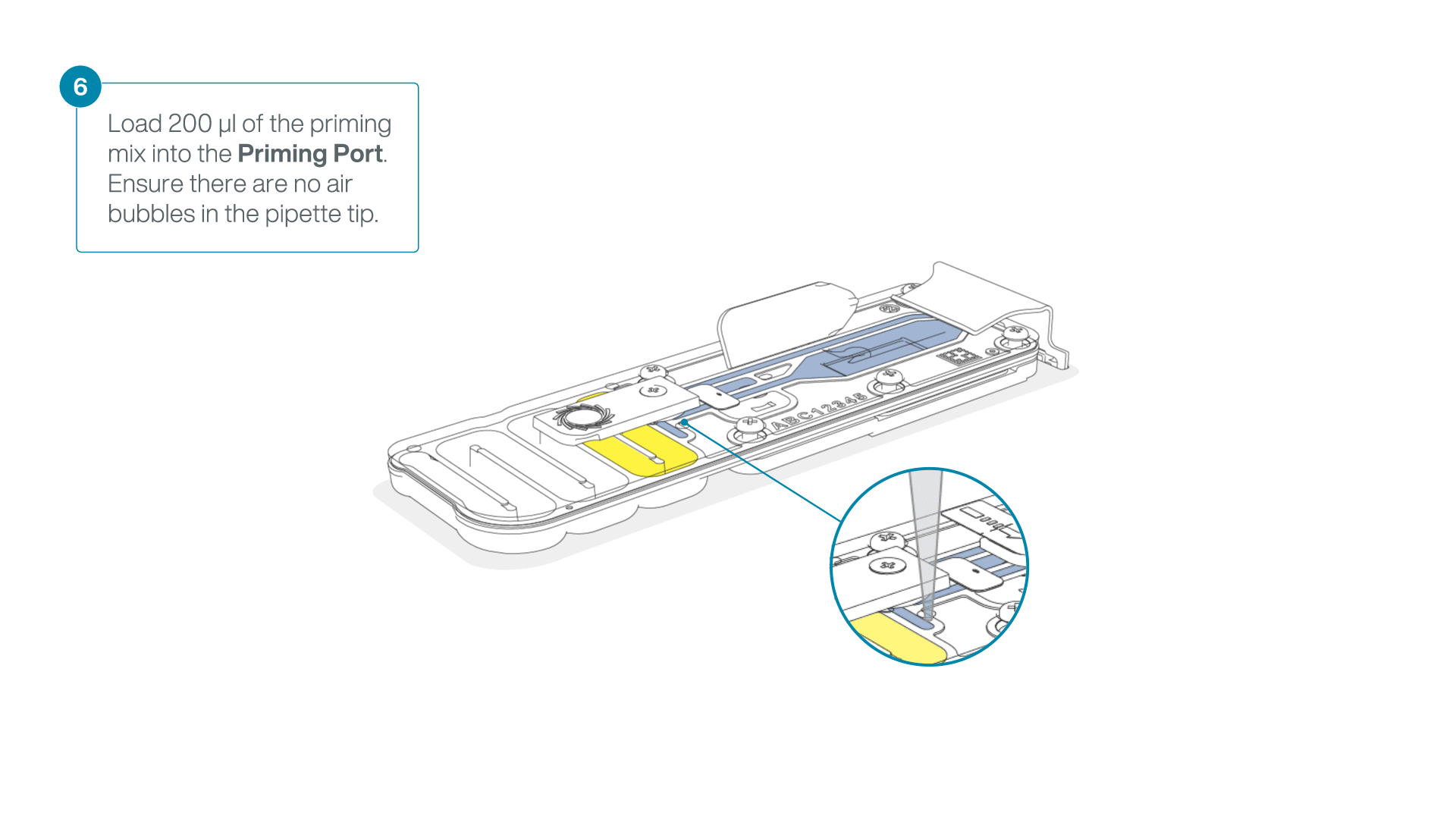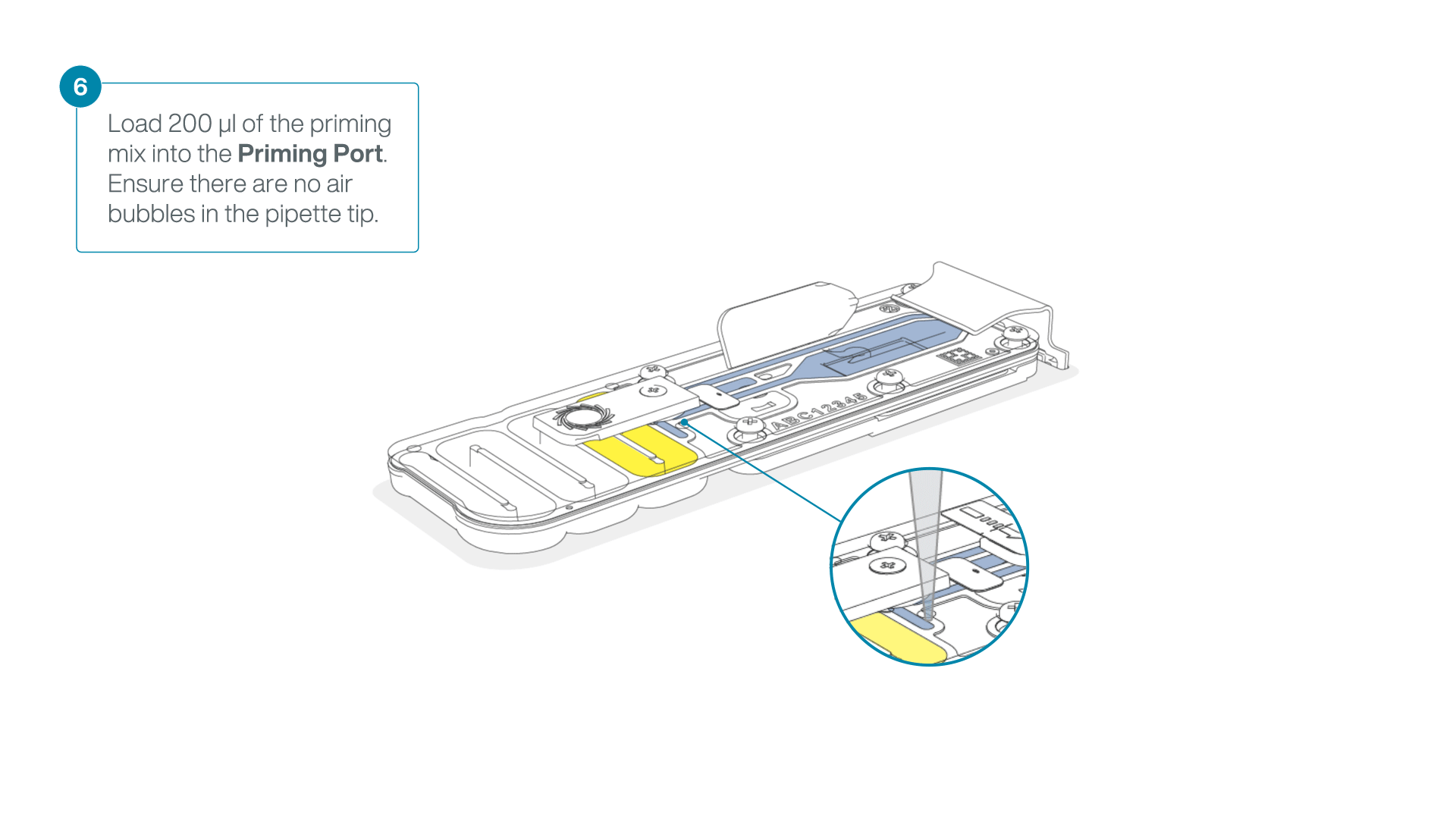
Complete the flow cell priming for the second flow cell:
- Gently lift the SpotON sample port cover to make the SpotON sample port accessible.
- Load 200 µl of the priming mix into the flow cell via the priming port (not the SpotON sample port), avoiding the introduction of air bubbles.
-
Mix the prepared library gently by pipetting up and down just prior to loading.
-
Add the recovered DNA library to the second flow cell via the SpotON sample port in a dropwise fashion. Ensure each drop flows into the port before adding the next.
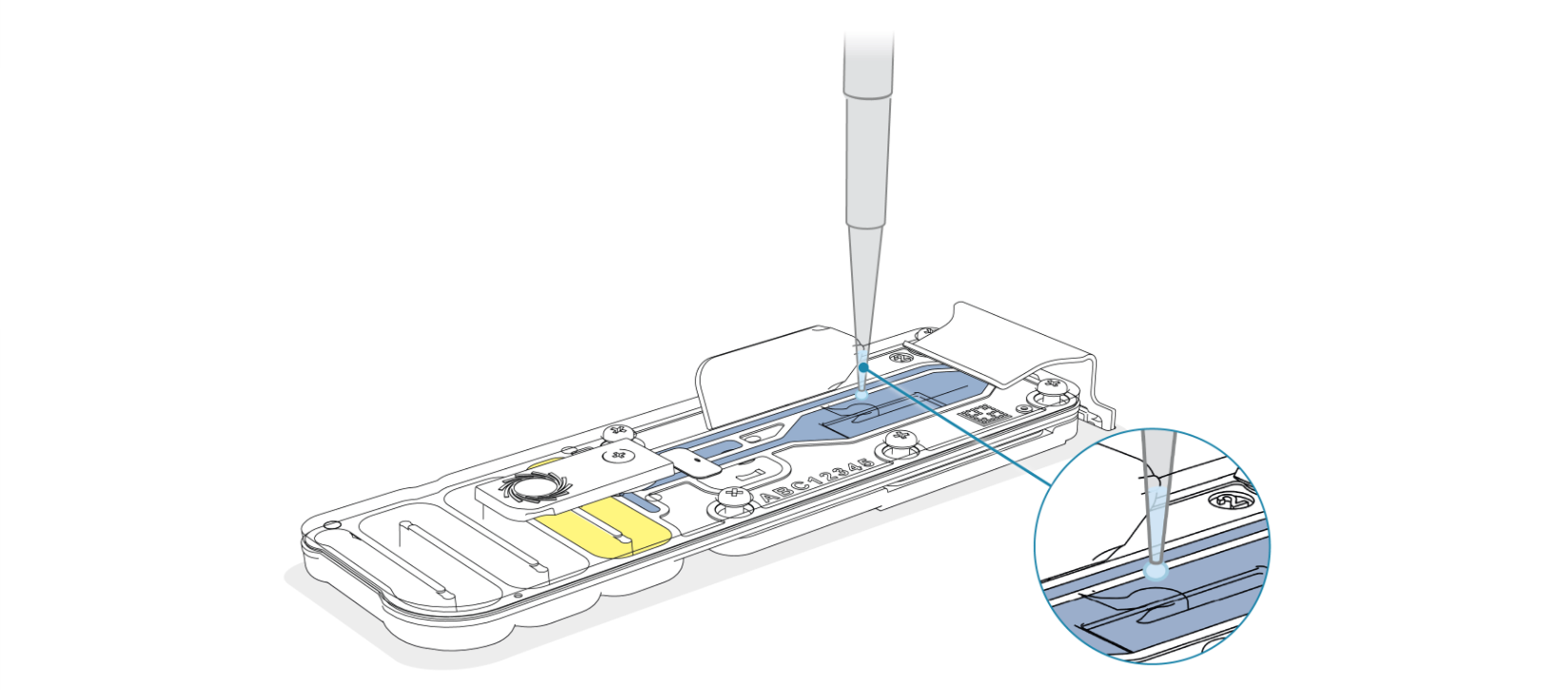
-
Gently replace the SpotON sample port cover of the second flow cell, making sure the bung enters the SpotON port and close the priming port.
-
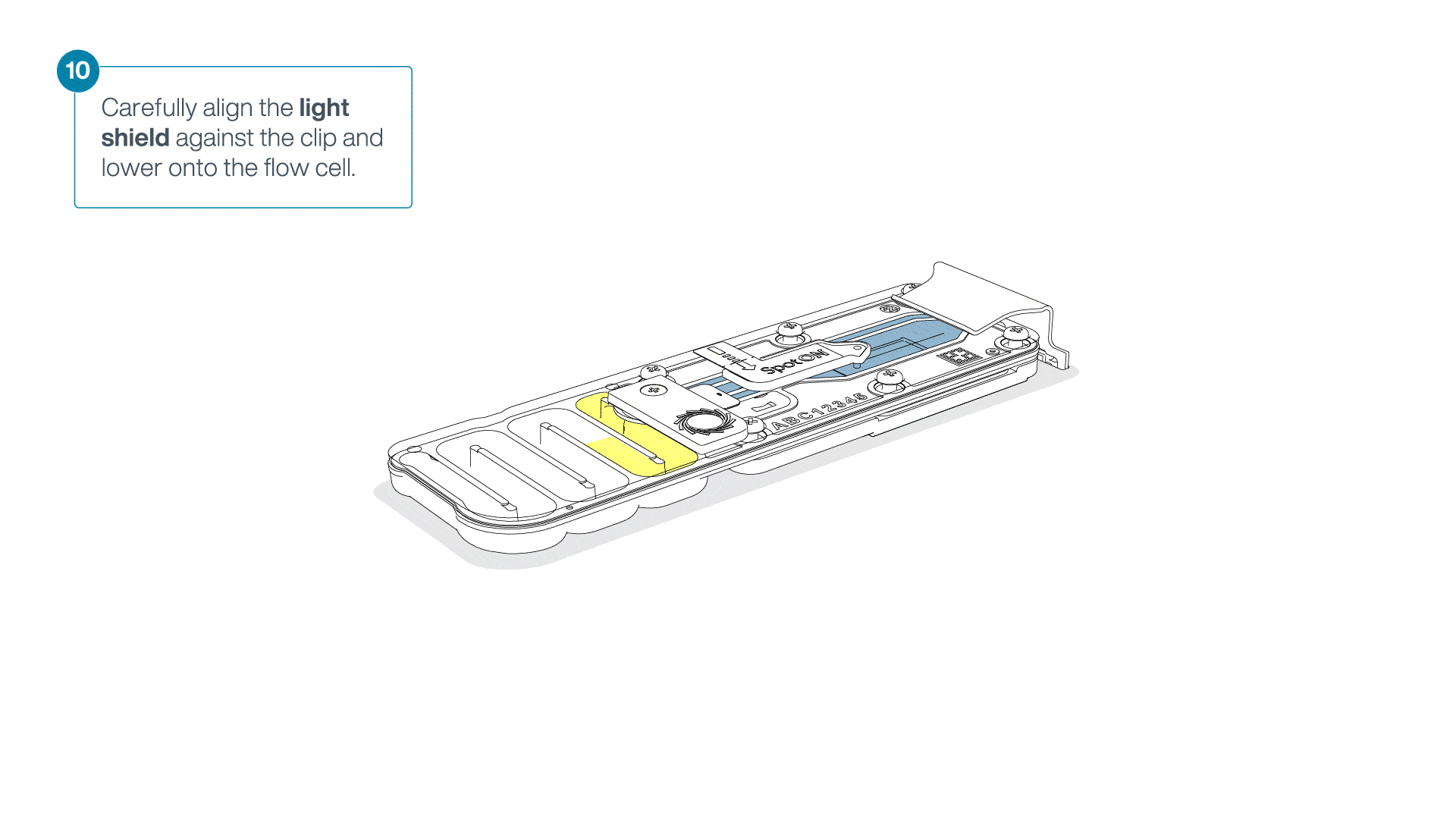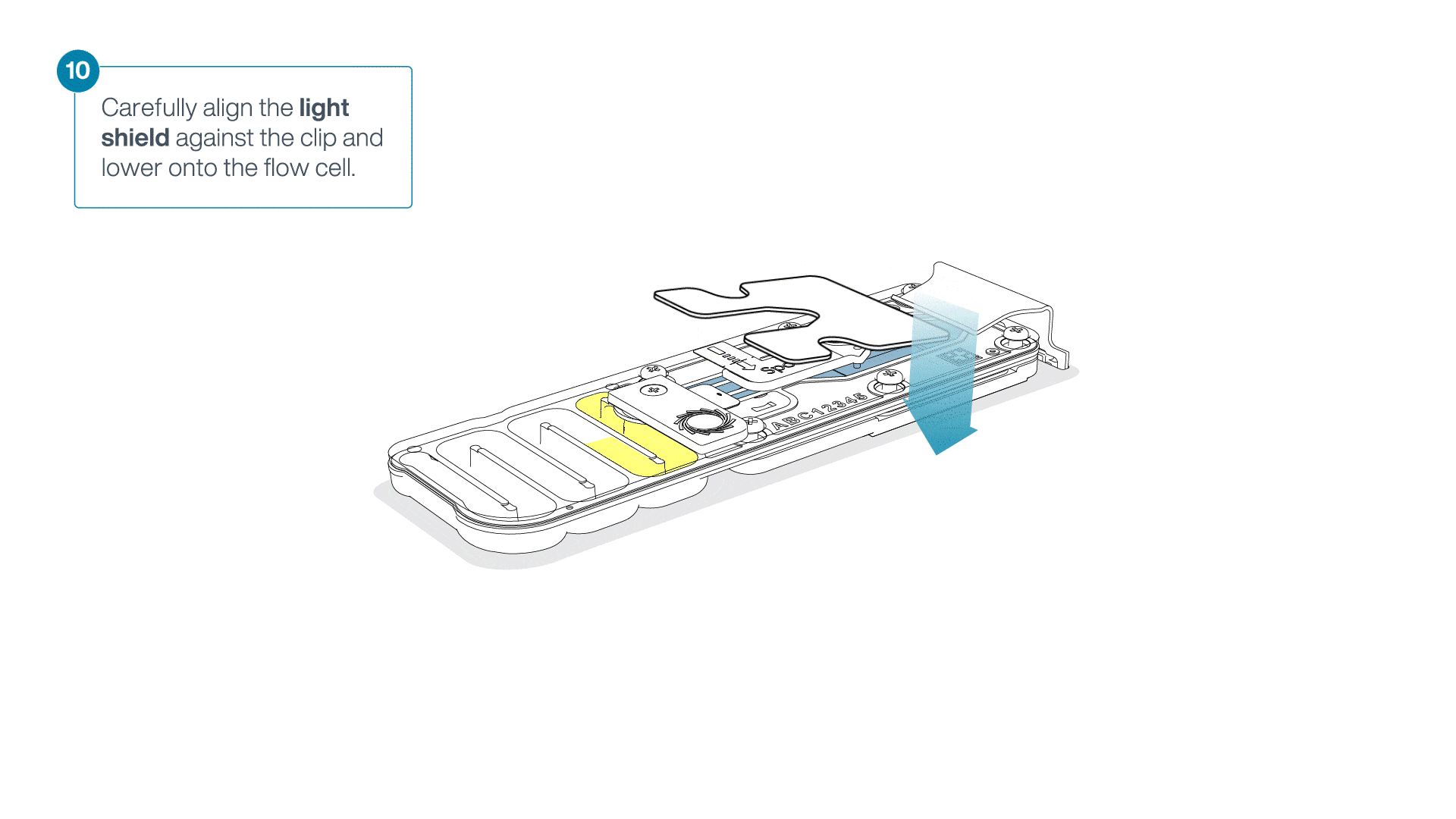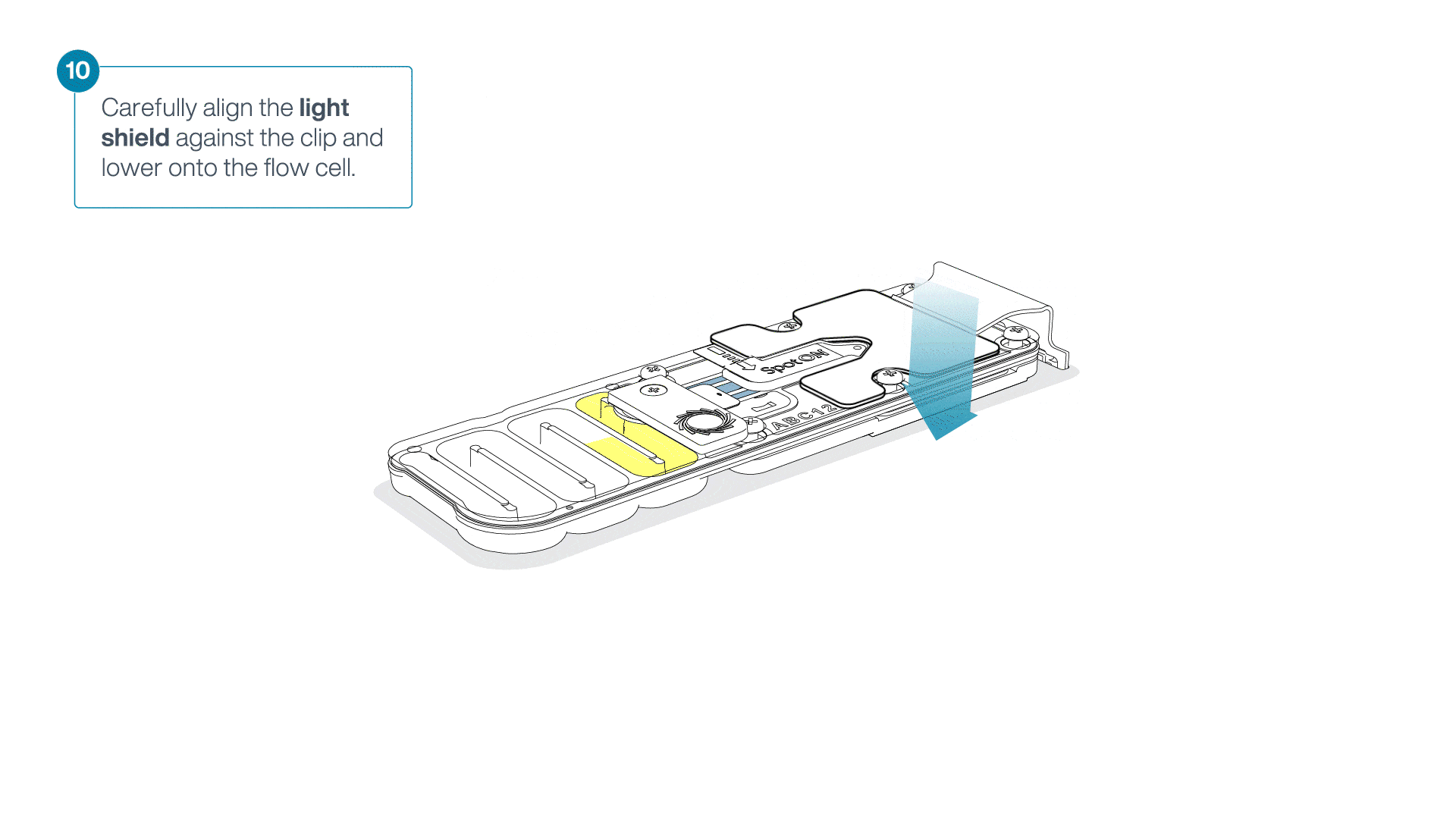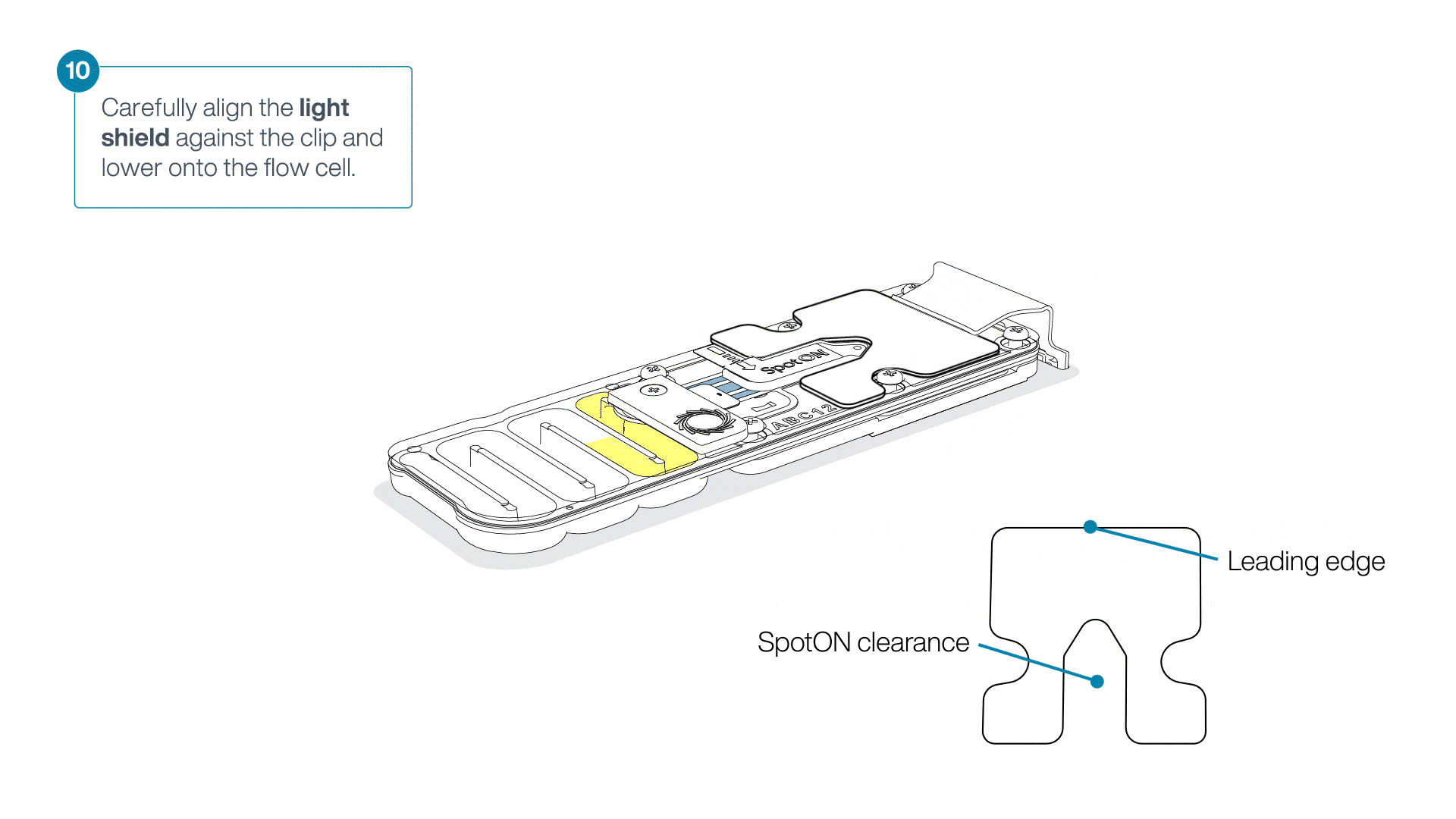
Place the light shield onto the flow cell, as follows:
Carefully place the leading edge of the light shield against the clip.
Note: Do not force the light shield underneath the clip.Gently lower the light shield onto the flow cell. The light shield should sit around the SpotON cover, covering the entire top section of the flow cell.
-
Start a new sequencing run on MinKNOW for the second flow cell.






