-
Overview of nanopore data analysis
For a full overview of nanopore data analysis, which includes options for basecalling and post-basecalling analysis, please refer to the Data Analysis document.
-
How to start sequencing
The sequencing device control, data acquisition and real-time basecalling are carried out by the MinKNOW software. It is assumed you have already installed MinKNOW on your computer. Further instructions for setting up a sequencing run can be found in the MinKNOW protocol.
In the current MinKNOW software version, the PCR Barcoding Expansions are not available in "Kit selection" when setting up a sequencing run. These will be included in the next software update.
For the meantime, we recommend starting a sequencing run in the MinKNOW software using Ligation Sequencing Kit V14 (SQK-LSK114) to perform real-time basecalling. Once basecalling is complete, the data can be demultiplexed using post-run barcoding on MinKNOW, as outlined below.
-
MinKNOW settings for real-time basecalling and post-run barcoding:
Real-time basecalling
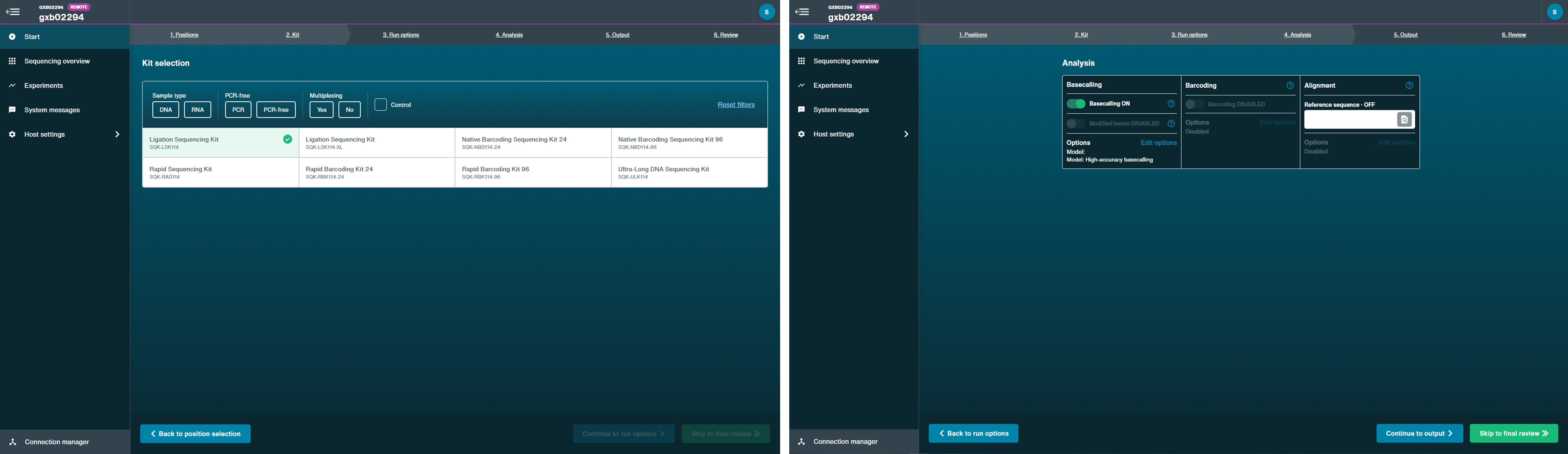
Follow the instructions in the MinKNOW protocol beginning from the "Starting a sequencing run" section for your device until the end of the "Completing a MinKNOW run" section.
Select Ligation Sequencing Kit (SQK-LSK114) in "Kit selection". The barcoding option will be unavailable as default. Other parameters can be kept at their default settings.
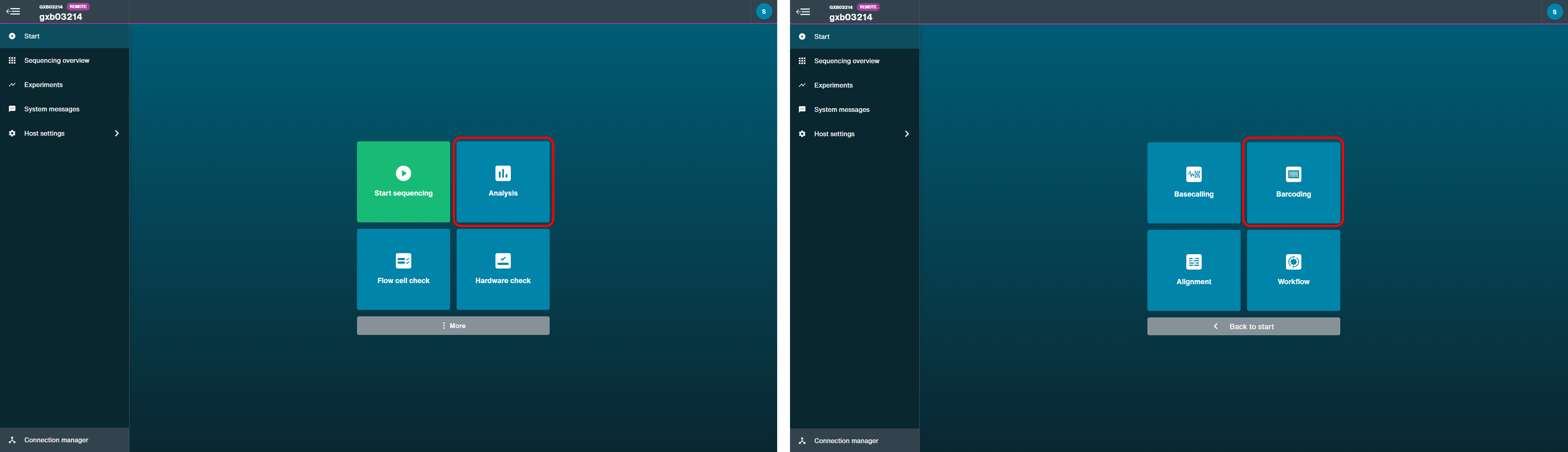
Post-run barcoding
Follow the instructions in "Post-run barcoding" of the MinKNOW protocol.
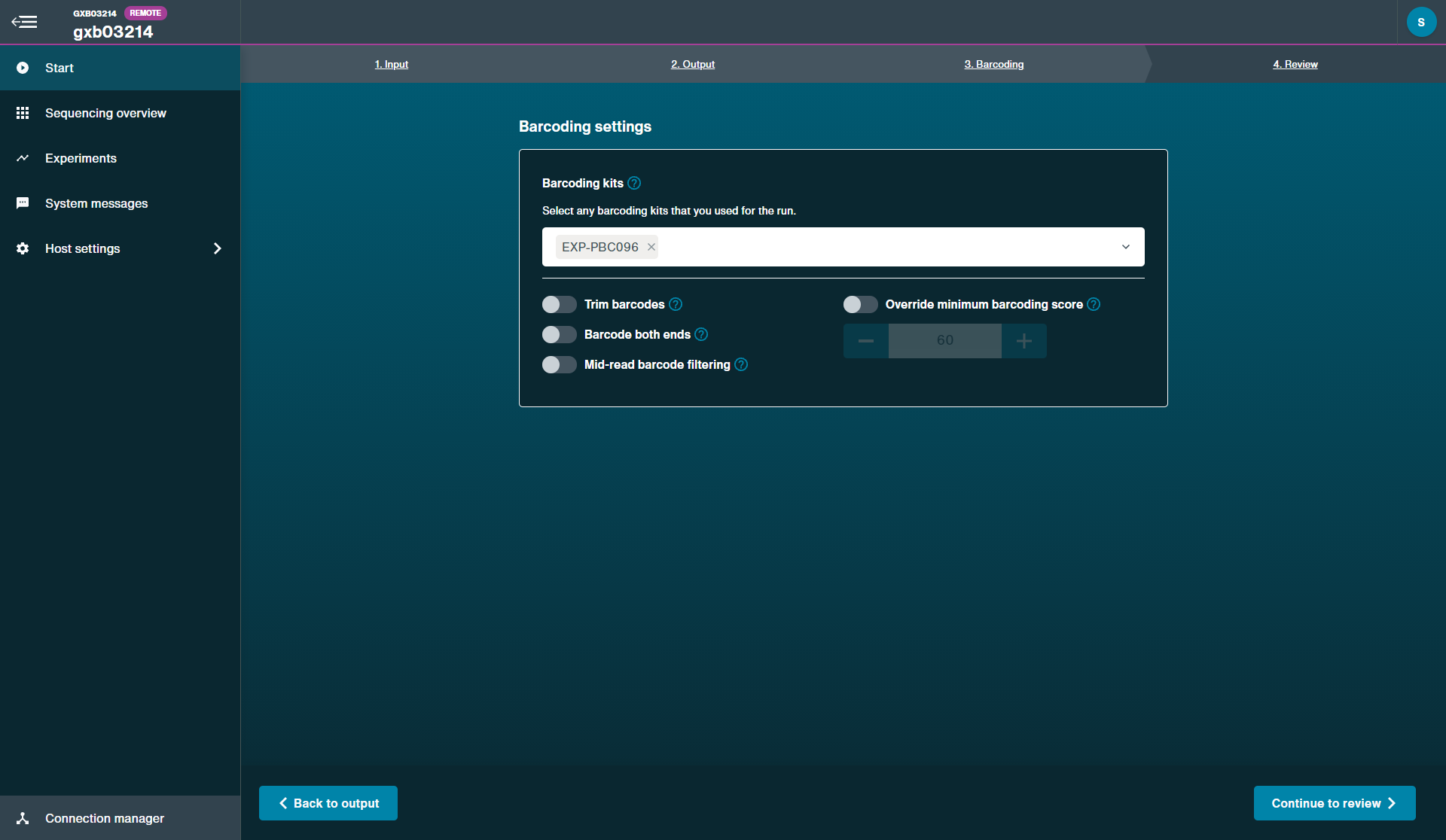
Click Analysis to open post-run options. Choose Barcoding and input your fastq data files. Select the PCR Barcoding Expansion used during library preparation in "Barcode settings" and use the default settings for other parameters.