-
Below is a list of the most commonly encountered issues, with some suggested causes and solutions.
We also have an FAQ section available on the Nanopore Community Support section.
If you have tried our suggested solutions and the issue still persists, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community.
-
Low sample quality
Observation Possible cause Comments and actions Inefficient lysis Enzyme activity has degraded in the solution or the isolate species is hard to lyse. - Make a fresh enzyme solution.
- Follow the hard to lyse gDNA extraction method.
- Increase the enzyme incubation for longer than 10 minutes.Low DNA concentration Low input into the extraction method - Check the cell input used
- Add more input and perform the extraction again
- Elute in less Elution Buffer
- Concentrate the DNA with a 0.4X AMPure XP Bead wash.Low DNA integrity number (DIN) Low quality or concentration of sample input - Repeat the extraction with freshly made enzyme solution
- Concentrate the DNA input with a 0.4X washLow sequencing yield Low sample concentration - Concentrate the DNA with a 0.4X AMPure XP wash step to remove potential inhibitors.
- Check the DNA concentration and quality. RNA presence may affect quantification of total DNA.Low DNA purity (Nanodrop reading for DNA OD 260/280 is <1.8 and OD 260/230 is <2.0–2.2) The DNA extraction method does not provide the required purity The effects of contaminants are shown in the Contaminants Know-how piece. Please try an alternative extraction method that does not result in contaminant carryover.
Consider performing an additional SPRI clean-up step. -
Low DNA recovery after AMPure bead clean-up
Observation Possible cause Comments and actions Low recovery DNA loss due to a lower than intended AMPure beads-to-sample ratio 1. AMPure beads settle quickly, so ensure they are well resuspended before adding them to the sample.
2. When the AMPure beads-to-sample ratio is lower than 0.4:1, DNA fragments of any size will be lost during the clean-up.Low recovery DNA fragments are shorter than expected The lower the AMPure beads-to-sample ratio, the more stringent the selection against short fragments. Please always determine the input DNA length on an agarose gel (or other gel electrophoresis methods) and then calculate the appropriate amount of AMPure beads to use. 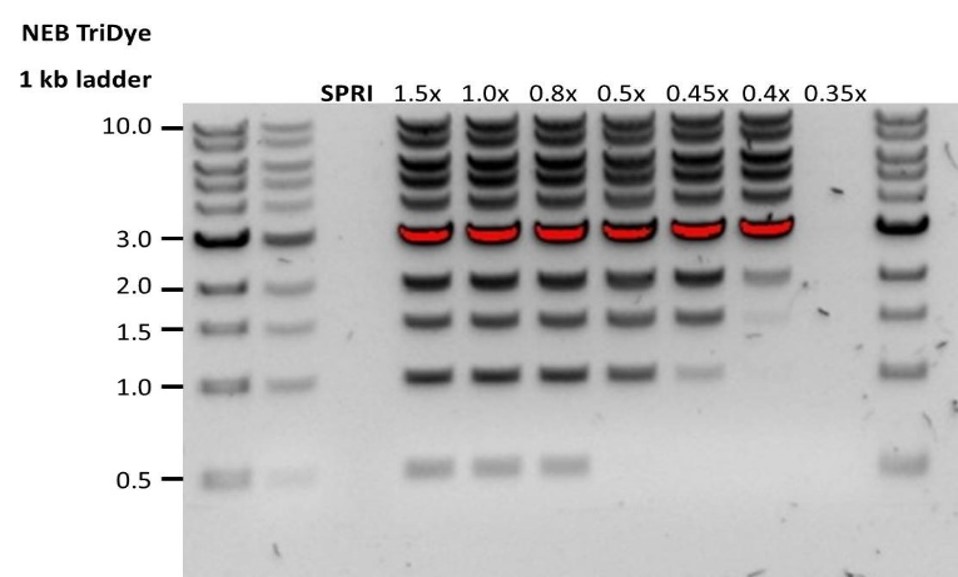
Low recovery after end-prep The wash step used ethanol <70% DNA will be eluted from the beads when using ethanol <70%. Make sure to use the correct percentage.
