- Materials
-
- <1 µg of each DNA sample to be barcoded in 45 µl
- Consumables
-
- NEBNext Ultra II End repair/dA-tailing Module (NEB, E7546)
- Freshly prepared 70% ethanol in nuclease-free water
- Nuclease-free water (e.g. ThermoFisher, AM9937)
- Agencourt AMPure XP beads (Beckman Coulter, A63881)
- 0.2 ml 96-well PCR plate
- Equipment
-
- Thermal cycler
- Ice bucket with ice
- Centrifuge capable of taking 96-well plates
- Magnetic rack, suitable for 1.5 ml Eppendorf tubes
- Optional equipment
-
- Qubit fluorometer (or equivalent for QC check)
-
Prepare the DNA in nuclease-free water.
- Transfer <1 μg DNA of each sample into a fresh 0.2 ml PCR tube or plate
- Adjust the volume to 45 μl with nuclease-free water
- Mix thoroughly by flicking the tube to avoid unwanted shearing
- Spin down briefly in a microfuge
-
In a 0.2 ml 96 well PCR plate, set up the end-repair / dA-tailing reactions as follows:
Between each addition, pipette mix 10-20 times.
Reagent Volume <1 µg DNA 45 µl Ultra II End-prep reaction buffer 7 µl Ultra II End-prep enzyme mix 3 µl Nuclease-free water 5 µl Total 60 µl -
Ensure the components are thoroughly mixed by pipetting, and spin down.
-
Seal the plate with adhesive film or PCR strip caps, spin down in a centrifuge and incubate for 5 minutes at 20 °C and 5 minutes at 65 °C using the thermal cycler.
If condensation is observed in the plate after the thermocycling, briefly spin down the plate contents in a centrifuge.
-
Resuspend the AMPure XP beads by vortexing.
-
Add 60 µl of resuspended AMPure XP beads to the end-prep reaction and mix by pipetting.
-
Allow DNA to bind to beads for 5 minutes at room temperature.
-
Prepare sufficient fresh 70% ethanol in nuclease-free water.
-
Place on a magnetic rack, allow beads to pellet and pipette off supernatant.
-
Keep the tube on the magnet and wash the beads with 180 µl of freshly-prepared 70% ethanol without disturbing the pellet. Remove the ethanol using a pipette and discard.
-
Repeat the previous step.
-
Cover the plate with adhesive film and leave plate on magnet for 2 minutes to allow residual liquid to collect at the bottom. Remove the adhesive film, return the plate to the magnet and aspirate residual wash solution.
-
Briefly incubate the plate on a thermal cycler at 37° C with the lid open and the plate wells unsealed.
-
Remove the plate from the magnet and resuspend pellet in 31 µl nuclease-free water. Incubate for 2 minutes at room temperature.
-
Pellet the beads on a magnet until the eluate is clear and colourless.
-
Remove eluate once it is clear and colourless. Transfer each eluted sample to a new 96-well PCR plate.
-
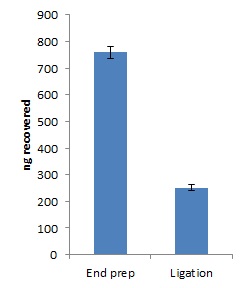
Quantify 1 µl of end-prepped DNA using a Qubit fluorometer - recovery aim >700 ng.