-
Below is a list of the most commonly encountered issues, with some suggested causes and solutions.
We also have an FAQ section available on the Nanopore Community Support section.
If you have tried our suggested solutions and the issue still persists, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community.
-
Low sample quality
Observation Possible cause Comments and actions Low DNA purity (Nanodrop reading for DNA OD 260/280 is <1.8 and OD 260/230 is <2.0–2.2) The DNA extraction method does not provide the required purity The effects of contaminants are shown in the Contaminants document. Please try an alternative extraction method that does not result in contaminant carryover.
Consider performing an additional SPRI clean-up step.Low RNA integrity (RNA integrity number <9.5 RIN, or the rRNA band is shown as a smear on the gel) The RNA degraded during extraction Try a different RNA extraction method. For more info on RIN, please see the RNA Integrity Number document. Further information can be found in the DNA/RNA Handling page. RNA has a shorter than expected fragment length The RNA degraded during extraction Try a different RNA extraction method. For more info on RIN, please see the RNA Integrity Number document. Further information can be found in the DNA/RNA Handling page.
We recommend working in an RNase-free environment, and to keep your lab equipment RNase-free when working with RNA. -
Low DNA recovery after AMPure bead clean-up
Observation Possible cause Comments and actions Low recovery DNA loss due to a lower than intended AMPure beads-to-sample ratio 1. AMPure beads settle quickly, so ensure they are well resuspended before adding them to the sample.
2. When the AMPure beads-to-sample ratio is lower than 0.4:1, DNA fragments of any size will be lost during the clean-up.Low recovery DNA fragments are shorter than expected The lower the AMPure beads-to-sample ratio, the more stringent the selection against short fragments. Please always determine the input DNA length on an agarose gel (or other gel electrophoresis methods) and then calculate the appropriate amount of AMPure beads to use. 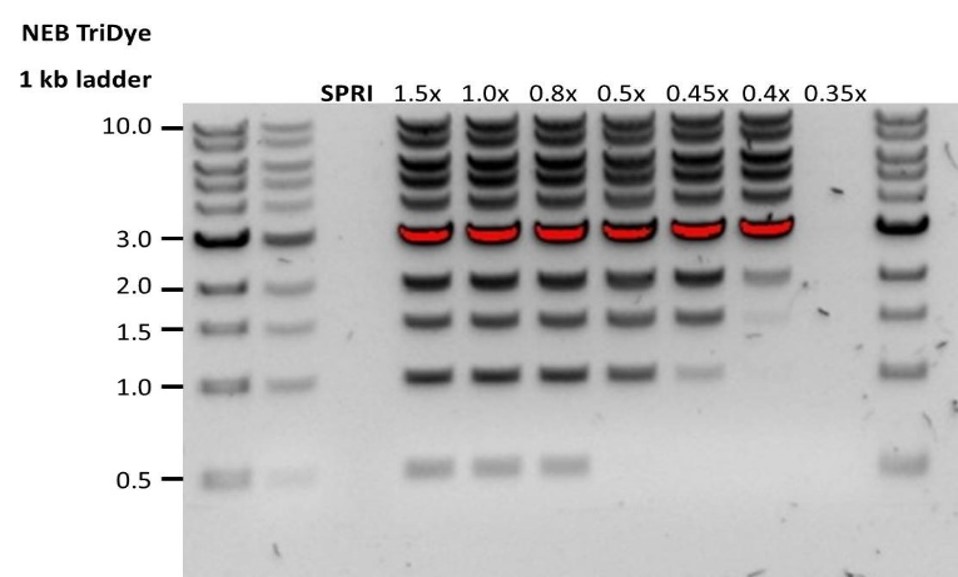
Low recovery after end-prep The wash step used ethanol <70% DNA will be eluted from the beads when using ethanol <70%. Make sure to use the correct percentage. -
The VolTRAX run terminated in the middle of the library prep
Observation Possible cause Comments and actions The green light was switched off
or
An adapter was used to connect the VolTRAX USB-C cable to the computerInsufficient power supply to the VolTRAX The green LED signals that 3 A are being supplied to the device. This is the requirement for the full capabilities of the VolTRAX V2 device. Please use computers that meet the requirements listed on the VolTRAX V2 protocol. -
The VolTRAX software shows an inaccurate amount of reagents loaded
Observation Possible cause Comments and actions The VolTRAX software shows an inaccurate amount of reagents loaded Pipette tips do not fit the VolTRAX cartridge ports Rainin 20 μl or 30 μl and Gilson 10 μl, 20 μl or 30 μl pipette tips are compatible with loading reagents into the VolTRAX cartridge. Rainin 20 μl is the most suitable. The VolTRAX software shows an inaccurate amount of reagents loaded The angle at which reagents are pipetted into the cartridge is incorrect The pipetting angle should be slightly greater than the cartridge inlet angle. Please watch the demo video included in the VolTRAX software before loading.
