-
Introduction to basecalling
Basecalling is the process of converting the electrical signals generated by a DNA or RNA strand passing through the nanopore into the corresponding base sequence of the strand. The general data flow in a nanopore sequencing experiment is shown below.
Raw data – a direct measurement of the changes in ionic current as a DNA/RNA strand passes through the pore, which are recorded by the MinKNOW software. MinKNOW also processes the signal into "reads", each read corresponding to a single strand of DNA/RNA. These reads are written out as POD5 files: a custom Oxford Nanopore file type.
Basecalling – the basecalling algorithm uses signal processing techniques based on machine learning to transform the raw signal of the reads into basecalls. The software writes out the results of these analyses into BAM files (unaligned, or containing modified base information and/or alignment information), with a default of 4000 reads per file. Additionally, FASTQ files are also produced. Similarly, the default is 4000 reads per file.
Oxford Nanopore Technologies provides several platforms to allow users to carry out basecalling in real-time, as well as executables for users' local infrastructure. You can carry out basecalling live during the experiment, as post-processing after an experiment has finished, or a combination of these.
-
Basecalling with neural networks
The production version of Oxford Nanopore basecallers convert raw signal data to basecalls using algorithms that incorporate bi-directional Recurrent Neural Networks (RNNs).
A neural network models processes that occur inside the human brain. The network contains nodes arranged in layers, which carry out computations. Neural networks receive and process data, but crucially, they have been trained to have exceptional performance for particular signal processing tasks. They have been successfully used for diverse applications like pattern recognition (such as handwritten characters, speech recognition), or predicting trends over time.
A recurrent neural network is a class of neural networks in which the output is dependent on past computations. An RNN keeps an internal memory of previously-seen data, so each new computation can use information from several preceding computations. A bi-directional RNN can set data in the context of what comes both before and after in the signal.
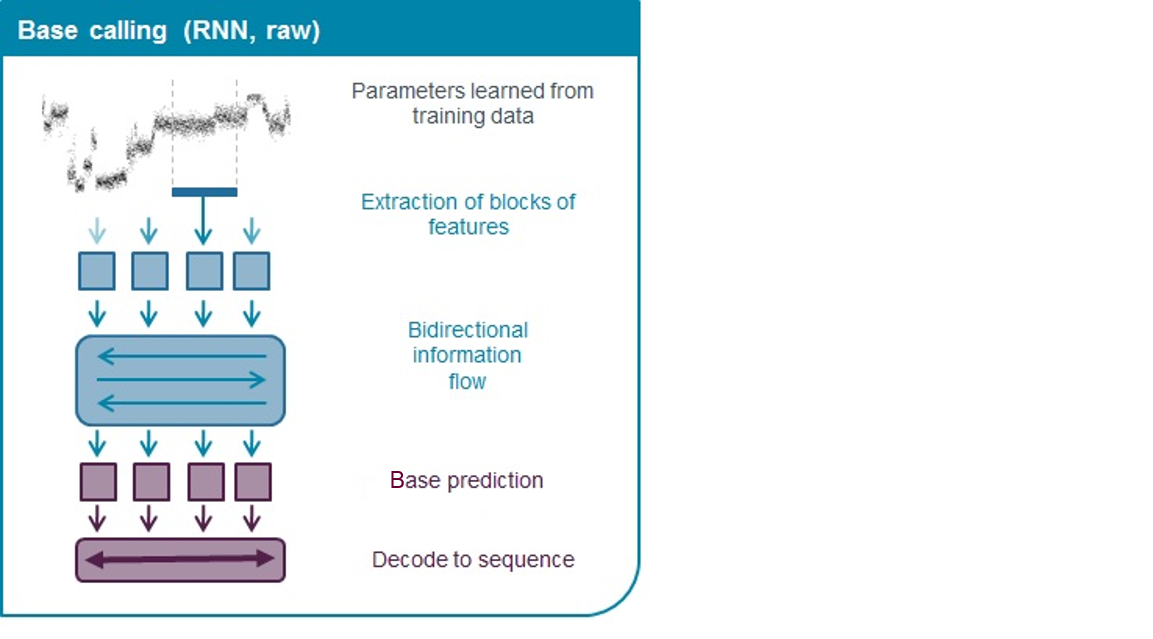
Oxford Nanopore's basecallers use neural networks that have been trained on a range of example DNA sequences (described in more detail in the Basecaller training section of Basecalling algorithms). The network learns how to translate the series of measurements into the sequence.
-
Oxford Nanopore basecallers
Basecaller Algorithm Availability MinKNOW basecaller Production basecaller on the device software. This is identical to the algorithm used by our stand-alone basecaller, but may be a version behind. Available as a free download (further details in the MinKNOW protocol). Select the basecalling option when starting the sequencing experiment, and MinKNOW will display the experiment progress via the user interface.
A Dorado-powered basecall server installed with MinKNOW is also available as a package for advanced users. ont-dorado-server is available as a free download, and is also included in MinKNOW installations. You can find the changelog.md file containing the documentation file (DOCUMENTATION.md) in the root of the archives.Dorado basecaller Dorado is the production basecaller that is also available in MinKNOW. Available as a free download. You can run the executable version of the software on the host computer via the command line. Dorado is heavily optimised for NVIDIA A100 and H100 GPUs and will deliver maximal performance on systems with these GPUs. GPU compatibility is also expected to work on other NVIDIA GPUs with at least 8 GB of GPU memory and architecture from Volta onwards. Research algorithms Varied Research algorithms are available through GitHub. The releases are varied, and often include features that will be included in future versions of the production basecaller.
