-
FASTQ Human Exome workflow in EPI2ME
Human Exome is an EPI2ME™ workflow for aligning basecalled reads to a set of reference sequences from Ensembl using the minimap2 aligner, covering the entire human exome. Alignments are performed against full gene sequences, including exons and introns, but currently not against flanking regions. The reference sequences are taken from the feature strand of the most recent human genome assembly, GRCh38.
The experiment report gives details on sequence length, accuracy, quality score and the amount of data generated during the experiment. It is possible to align to a number of human reference genes, and view more details about alignment coverage and accuracy.
For more information, please refer to the Workflows section in the EPI2ME protocol.
-
Suggested workflow
- Align reads to the human reference genome (GRCh38 or GRCh37) using minimap2. We recommend using -k parameter value 13
- (Optional) Remove reads that do not overlap with pull-down regions
Example data
Expected mean target coverage from one MinION Mk 1B flow cell:
Aligned read length:
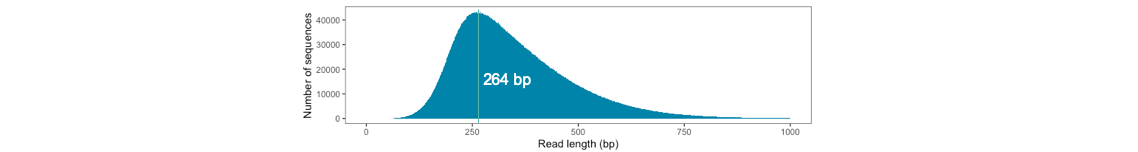
-
Other data analysis options
1. Bioinformatics tutorials
For more in-depth data analysis, Oxford Nanopore Technologies offers a range of bioinformatics tutorials, which are available in the Bioinformatics resource section of the Community. The tutorials take the user through installing and running pre-built analysis pipelines, which generate a report with the results. The tutorials are aimed at biologists who would like to analyse data without the help of a dedicated bioinformatician, and who are comfortable using the command line.
2. Research analysis tools
Oxford Nanopore Technologies' Research division has created a number of analysis tools, which are available in the Oxford Nanopore GitHub repository. The tools are aimed at advanced users, and contain instructions for how to install and run the software. They are provided as-is, with minimal support.
3. Community-developed analysis tools
If a data analysis method for your research question is not provided in any of the resources above, we recommend the Community-developed data analysis tool library. Numerous members of the Nanopore Community have developed their own tools and pipelines for analysing nanopore sequencing data, most of which are available on GitHub. Please be aware that these tools are not supported by Oxford Nanopore Technologies, and are not guaranteed to be compatible with the latest chemistry/software configuration.
