-
Data transfer and long-term storage - all devices (GridION, Linux/Mac/Windows workstations)
It is essential that data is streamed from the device in real-time to prevent runs from terminating due to lack of storage space (this is common for high specification laptops). For this, a customer site must ensure that connectivity to the local infrastructure/external SSD is of sufficient bandwidth to prevent data backing up. We recommend using a remote server or network-attached storage (NAS). These typically offer a choice of NFS, SMB, or CIFS shares. If the workstation attached to the P2 Solo uses a Linux-based operating system, we recommend using NFS mounts due to the supported Linux user and group permissions. For Windows workstations, use SMB and for Mac OS use either SMB or NFS. To stream data to storage in real-time, SSD is required due to its high write speed compared to HDD. After initial writing to networked SSD drives, data can be moved to storage with a slower write speed for long-term storage.
The form and volume of data to be stored will depend on your requirements and whether you wish to basecall your sequencing data in the future when more advanced basecalling algorithms are available:
- Storing .POD5 files with raw read data in will permit re-basecalling of data when new algorithms are released by Oxford Nanopore Technologies. In such cases, new releases of basecallers have enabled significant improvements in basecalling accuracy of existing datasets through re-basecalling. Further, selected Oxford Nanopore and third-party tools use the raw signal information contained within the POD5 to extract additional information from the raw signal, e.g. calling modified bases, reference-guided SNP calling, or polishing of data.
- Retaining only FASTQ files will allow use of standard downstream analysis tools using the DNA/RNA sequence, but no further sequence data can be generated when improvements in basecalling become available.
Oxford Nanopore is unable to provide exact recommendations for storage, as these will be site-specific.
-
For more information about offloading data, refer to our Nanopore Learning video:
GridION: Data offload to remote storage
Our device FAQs are located here.
-
GridION data transfer
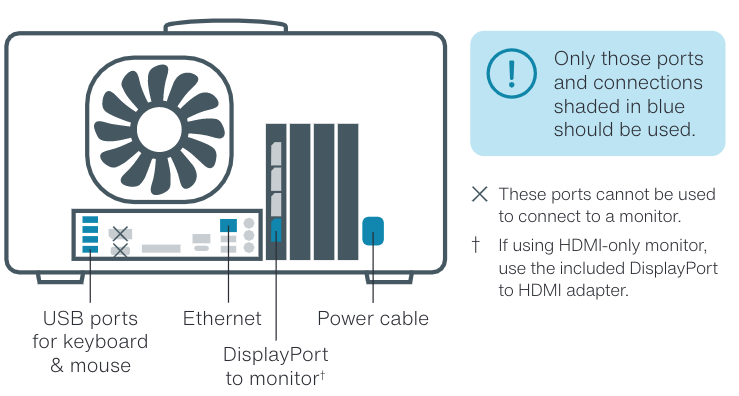
If you are using a P2 Solo in combination with a GridION and require additional SSD storage, ensure you are using the correct USB port/Ethernet on the rear of the device. Do NOT use either the USB port with a white, rectangular centre or those at the front of the device (if your GridION has front-facing USB ports).
Instead, use the blue USB Type-A ports on the rear of the GridION (see the image below for reference). Alternatively if using Ethernet, ensure that you are using a cable capable of at least 1 Gbps (CAT5e) and the minimum length for reduced latency.
-
From MinKNOW version 21.02 onwards, we have added new functionality to enable smoother data transfer off the box during a sequencing run. The instructions below are an example method to mount an external NFS and transfer data. Please consult your local IT department before implementing any code to ensure it is compatible with the local infrastructure and the correct permissions are in place.
-
Run the following command:
sudo sed -i 's/prom/grid/g' /lib/systemd/system/ont-platform-data-offload.service -
Mount your local NFS file system on the device (note that Linux mounts a remote filesystem into a directory locally). Below is an example setup through the terminal:
grid@GXB02000:~$ sudo su -
grid@GXB02000:~$ apt install autofs
grid@GXB02000:~$ echo -en “+auto.master\n/nfs /etc/auto.ont\n” > /etc/auto.master
grid@GXB02000:~$ echo -en "NETWORKSTORAGE -nfsvers=3,rw,bg,async,actimeo=300,soft,intr,noatime,tcp,nolock IPADDRESS:/PATH/TO/SHARE" > /etc/auto.ont
grid@GXB02000:~$ ln -s /nfs/NETWORKSTORAGE /media/NETWORKSTORAGE
grid@GXB02000:~$ ls -al /media/NETWORKSTORAGE/* -
Install the latest version of the MinKNOW software:
grid@GXB02000:~$ sudo apt update
grid@GXB02000:~$ sudo apt install ont-gridion-release -
Make a source directory that your experiments will be saved to in /data/ , for example:
grid@GXB02000:~$ mkdir /data/data-offload -
Make a destination directory on your networked storage. This also tests that the ‘grid’ user can write to the networked storage:
grid@GXB02000:~$ mkdir /media/mounted_drive/destination_directory/ -
As root, edit /etc/systemd/ont-platform-data-offload.conf to set the SOURCE_DIR and DESTINATION_DIR variables. SOURCE_DIR is the directory the PromethION software writes to, and DESTINATION_DIR is on your networked storage.
grid@GXB02000:~$ sudo vi /etc/systemd/ont-platform-data-offload.conf
-
A log of the actions taken by the script will be written to /data/data-offload.log
-
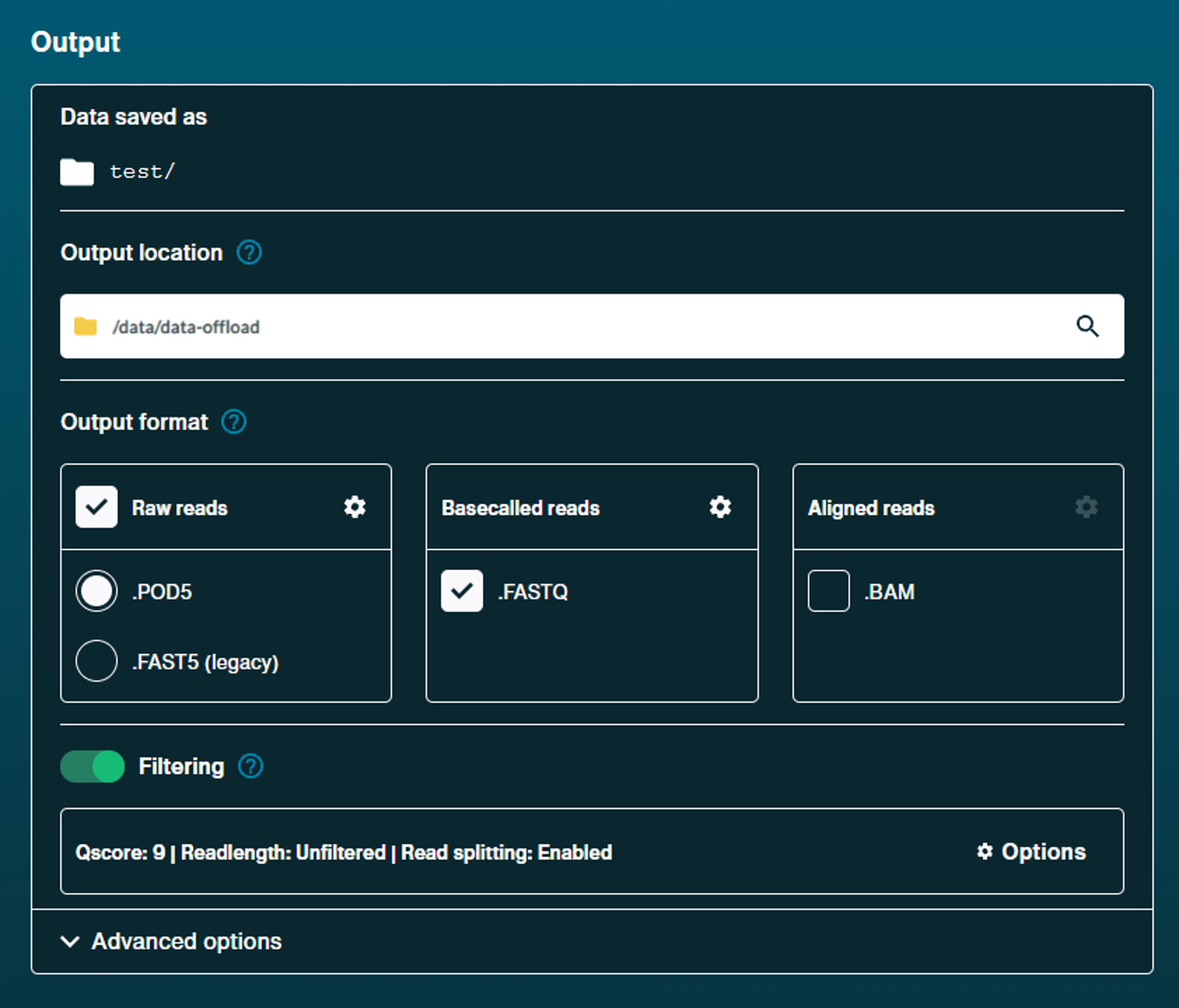
Start your sequencing in the MinKNOW UI, setting the output location to SOURCE_DIR set above.
-
We currently recommend manually controlling the service, as the data offload activity could affect active runs.
To start the offload service as root:
sudo systemctl start ont-platform-data-offloadTo stop the offload service as root:
sudo systemctl stop ont-platform-data-offload -
To check that the service is running, run the following command:
sudo systemctl status ont-platform-data-offload


