-
Factors contributing to the nanopore signal
The nanopore and molecules interacting with the nanopore
E.g. the motor protein.
Applied voltage
- Set voltage, defined in the protocol scripts in MinKNOW
- The resistance of the electrochemical path between the common electrode and the well electrode
- The electrochemistry at the electrodes
Temperature
Higher temperatures increase the mobility of charged species and thereby increase the current. A range of set temperatures is used for various steps of the sequencing run:

Buffer
The storage and running buffers contain different concentrations of charged species, and will affect the pores and DNA translocation differently.
-
Platform QC
The choice of which well is used is made via the multiplexer (MUX), a switch which allows the selection of the best channels for sequencing. For a summary of the MUX layout, please see the channel layout description above. The Platform QC is a scan of each MUX in turn to see if a pore is present, with the aim of determining the total number of functional pores present in a chip. Platform QC must be run on a new flow cell within ten days of receipt.
The Platform QC script runs through each MUX in turn, first applying a triangle wave to assess the membrane. Then 180 mV are applied for 30 s, and different metrics contained in the signal, such as current levels, noise measurements and events that can help to classify the signal, are recorded. The pores are classified as 'single pore', 'multiple pores', 'saturated', and 'zero current'. To class a single pore, the software searches for an open pore current level within a given range, combined with the distinct signal of the QC DNA strand that is present in the flow cell storage buffer. The MinKNOW GUI then displays the total number of single pores.
-
The MUX scan
At the beginning of the sequencing run, when a library has been loaded, an additional MUX scan looks at the performance of the nanopores and groups them into collections of 512 to determine which one will provide the best signal and therefore the best sequence during the time of the run. In a similar fashion to the Platform QC, a 180 mV voltage is applied to each MUX in turn. If there are library DNA strands in any of the pores, these are now used to classify channels. Channels are prioritised for sequencing based on 'strand' first, then 'pore', then 'unavailable' pores.
- strand - a DNA strand translocating through a single pore
- pore - an open pore without DNA
- unavailable - signals that the software is not confident about
The MUX scan report shows the number of active pores, defined as channels in strand and open pore. MinKNOW then groups the best 512 channels for sequencing plus 2nd and 3rd choices that can be utilised during a sequencing run. This is the groups 1, 2, and 3 reported on the GUI. Sequencing is begun from group 1.
-
Voltage drift during a sequencing run
Once the MinKNOW script proceeds to sequencing, the system goes through a cycle of voltage changes. The current scripts start with an applied voltage of -180 mV, which is optimal for basecalling with R9.4.1 chemistry. However in the process of a run, the voltage drifts due to the depletion of the redox chemistry in the bulk solution.
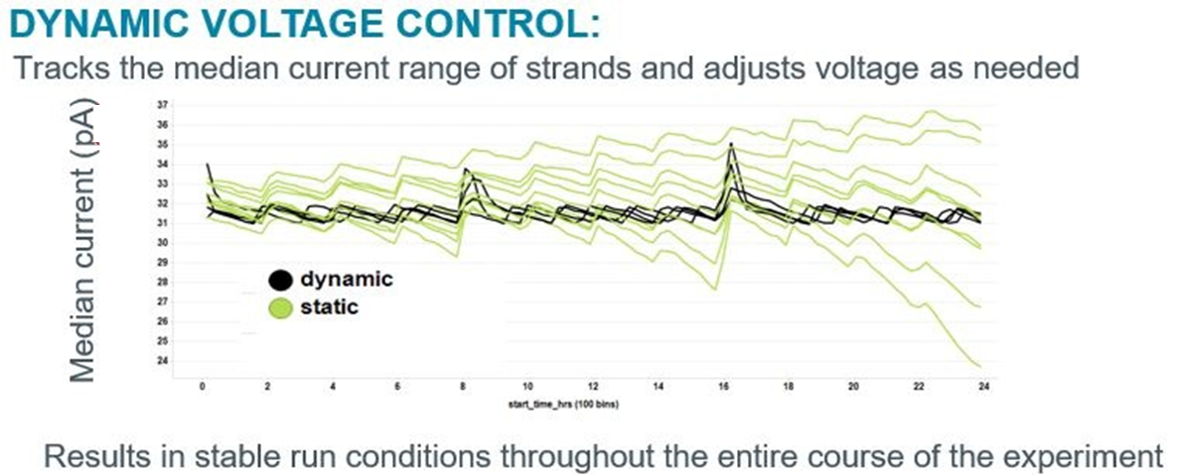
In order to keep the voltage constant, MinKNOW enables "Dynamic Voltage Control", which constantly monitors the raw signal from the device, and adjusts the applied voltage when required. Dynamic Voltage Control (in black) maintains a considerably tighter grouping of current levels, which is more stable over the course of a run than that previously achieved by the Static Voltage Control used in earlier versions of MinKNOW: