-
How Cas9 works
The CRISPR/Cas family of proteins comprises DNA- or RNA-cleaving enzymes that can be programmed to cut specific sequences using oligonucleotide-length RNA, making them suitable candidates for genome editing but also targeted sequencing, given their high specificity.
CRISPR RNAs (crRNAs) program Cas9 to bind and cleave DNA at sites that are identical (or highly similar) to the crRNA sequence. Candidate crRNAs need to be determined by the user and can be found by bioinformatic searches of reference genomes (detailed below). When Cas9 is loaded with crRNA identical to the intended target sequence, together with trans-activating CRISPR RNA (tracrRNA - a structural RNA required for catalytic activity) it forms a ribonucleoprotein complex (RNP). This complex of RNA and protein searches the genomic DNA for the target region using the crRNA sequence. The complex attempts to seed the crRNA by melting duplex DNA, and, if the crRNA matches and base-pairs with the target sequence, Cas9 cuts both strands of the target sequence.
Cas9 requires a 20mer target site (the protospacer), plus the protospacer-adjacent motif (PAM - sequence NGG). The PAM is immediately next to the 3’ end of the target and defines the boundaries of the protospacer sequence. Cas9 cuts both strands of the target sequence 3 bp upstream of the PAM. The production of a newly-exposed end, what we call ‘deprotection’, at a defined site is the basis for the selectivity of the Cas-mediated enrichment method.
For Cas9 targeted sequencing, we are able to selectively protect and deprotect ends using CRISPR/Cas and enrich for regions of interest in a background prior to long read nanopore sequencing of the native DNA molecule.
-
How to design a targeted sequencing approach using Cas9
There are several ways to design a Cas9 targeting experiment, and the method depends on the target region of interest. Multiple factors determine which method should be used:
- Size of region of interest (ROI)
- Length of input DNA
- Are the sequences either side of the region of interest known?
- How much coverage of region of interest is required?
There are three main options, detailed below.
-
Excision approach
crRNA probes are design to target regions either side of the region of interest to excise that region (ideal for particular gene targets).
This method should be used:
- When the region of interest is <20 kb (largest target region if multiple regions are being targeted)
- When both ends of the region of interest are known, to be able to design probes on either side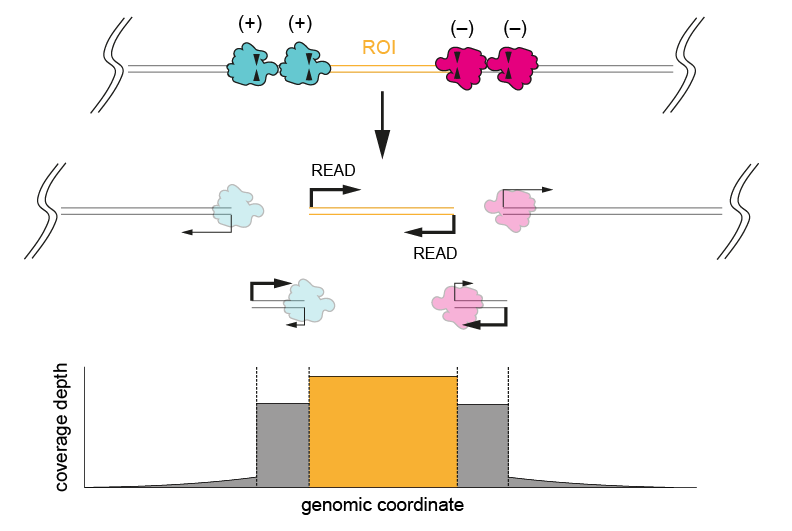
Figure 1. Excising a region of interest using four cuts (two each side of the ROI). Cutting once on each side results in incomplete cleavage, as shown by the high coverage depth of the region behind the ROI after the first cut (bottom panel).We generally recommend excising a ROI by making four cuts, two upstream of the ROI targeting the (+) strand, and two downstream targeting the (-) strand. Four cuts instead of two are for redundancy, in case one or more crRNAs yield incomplete cleavage. This method should provide the highest coverage of a target region, as on-target strands will have sequencing adapters ligated to both ends. Regions larger than 20 kb can be targeted using this approach, but this is limited by the length of the input DNA and users may experience drops in coverage towards the middle of the region of interest.
-
Single cut and read out
A crRNA probe is designed at one end of a target region to read into the unknown (ideal for characterisation of integration sites within a genome).
This method should be used:
- When the region of interest is <20 kb (largest target region if multiple regions are being targeted)
- When only one side of the region of interest is known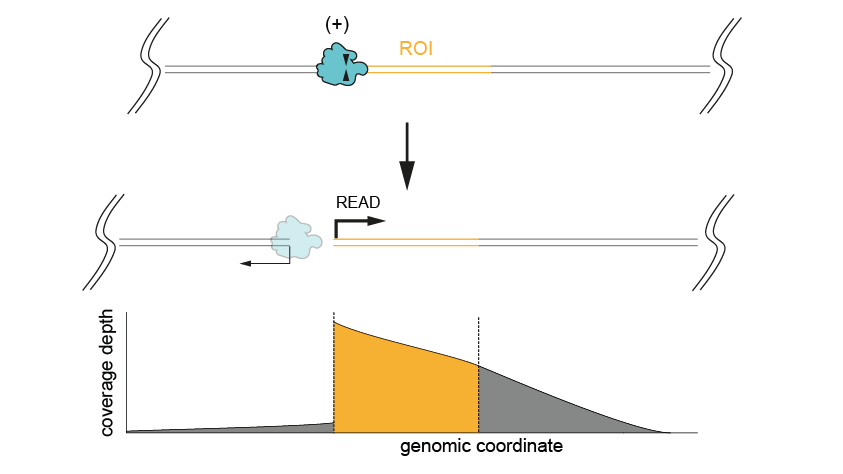
Figure 2. Excising a region of interest using a single cut, when only one side of the ROI is known. The coverage depth is high both for the ROI and the region downstream (bottom panel).This method will give lower throughput due to only ligating sequencing adapters to one end of the strand. Regions larger than 20 kb can be targeted using this approach, but this is limited by the length of the input DNA and users may experience drops in coverage towards the end of the region of interest.
-
Comparison of the single cut and read-out vs excision approach
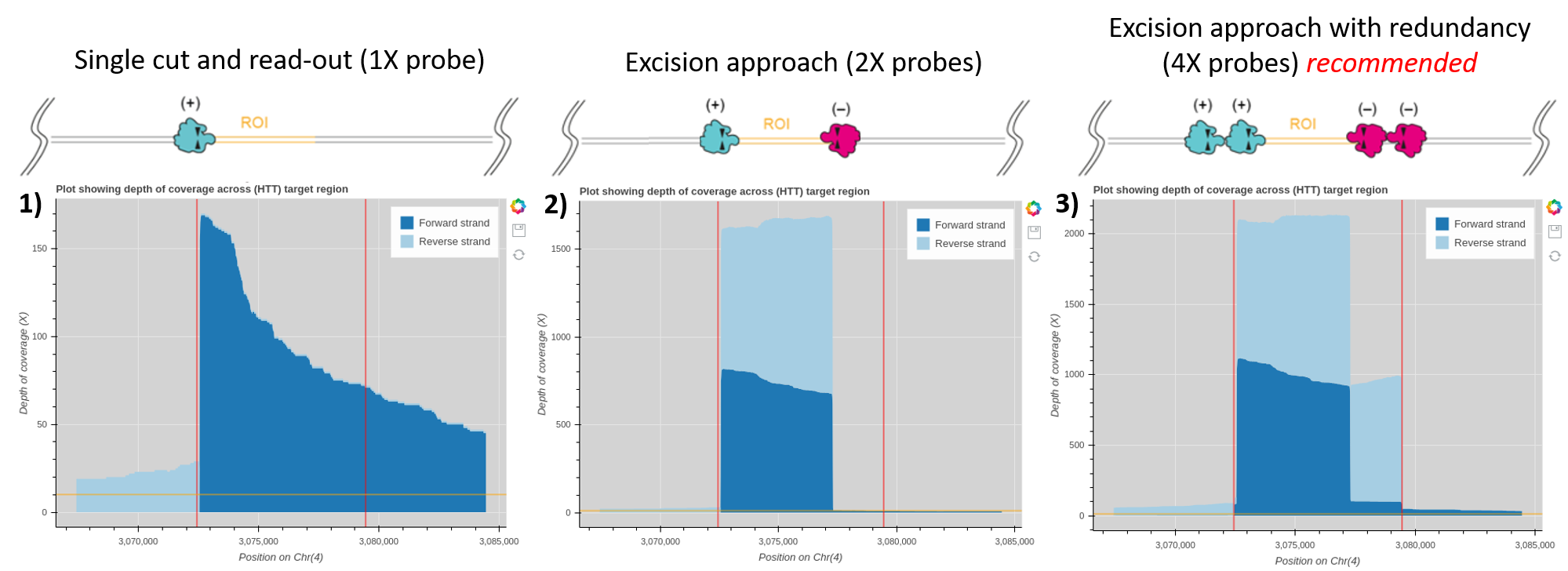
Figure 3. Example coverage plots generated using the Oxford Nanopore bioinformatics tutorial “Evaluation of read-mapping characteristics from a Cas-mediated PCR-free enrichment” for the HTT gene with alternative probe design methods.
An enrichment experiment run on a single MinION R9.4.1 flow cell targeting the HTT gene with a single probe (panel 1), an excision approach using one probe on either side of the region of interest (panel 2) and the recommended excision approach using 4X probes for redundancy (panel 3). Red lines on the coverage plot highlight the range in the input BED file used in the bioinformatics tutorial. This example ranges from outer probe to outer probe for the 4X excision method for all plots. The HTT gene falls between these red lines, so the average coverage of the HTT gene is the highest coverage observed on these graphs. Therefore the average coverage of the HTT gene with a single probe is around 100X, 1600X for an excision approach with 2X probes and 2100X coverage with a 4X probe excision. -
Tiling
crRNA probes are designed along a region of interest in 5-10 kb overlapping chunks that allows for even coverage across the region of interest.
This method should be used:
- When the region of interest is >20 kb (especially when limited by the length of the input DNA)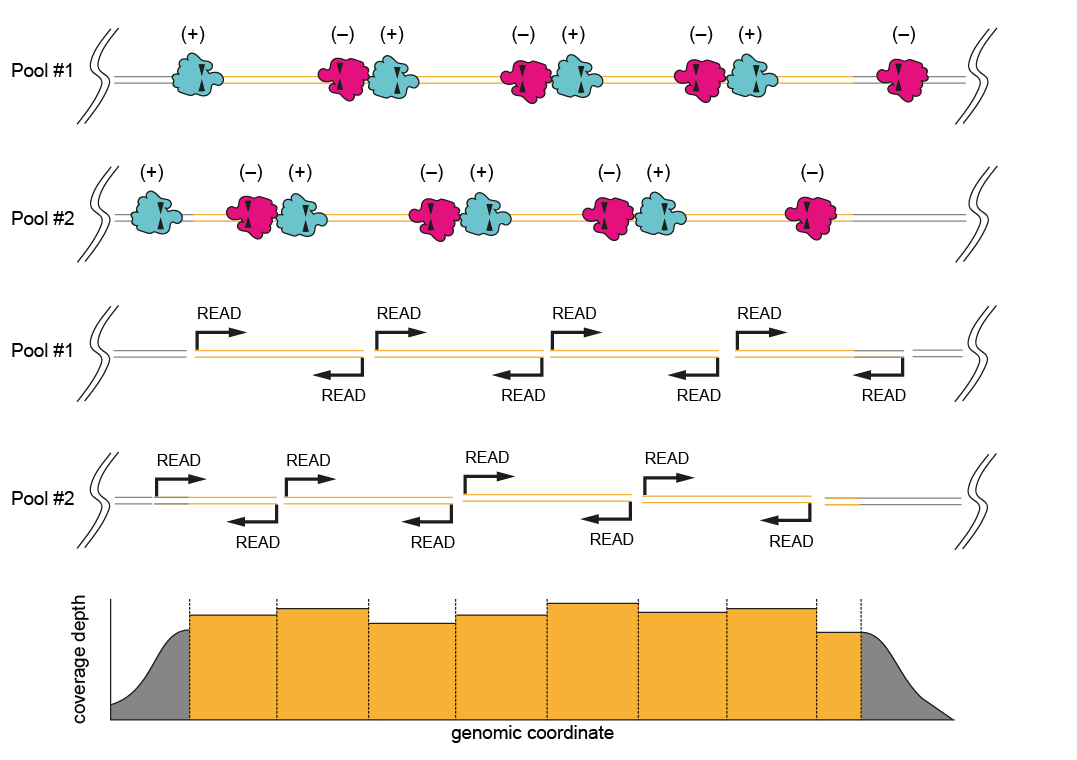
Figure 4. Probe design for a “bricklayer” tiling approach. Two sets of probes are designed covering a large region of interest (>20 kb). This breaks up the ROI into overlapping sections, ensuring good coverage of the whole length of the ROI.This method involves designing two pools of probes which should be prepared as two separate experiments and pooled in the final step (before the final clean up) and loaded together onto the same flow cell. There are two options for designing the probes. For most use cases, we recommend a “bricklayer” approach, where each pool contains (+) and (–) probes that are designed to overlap each other. The second “highway” option is to have one pool of probes cutting in one direction (+) and one pool of probes cutting in another (–) (probe directionality is explained below). The “bricklayer” approach has shown to give a higher coverage of the target region.
-
Decision tree for selecting the Cas9 targeting approach
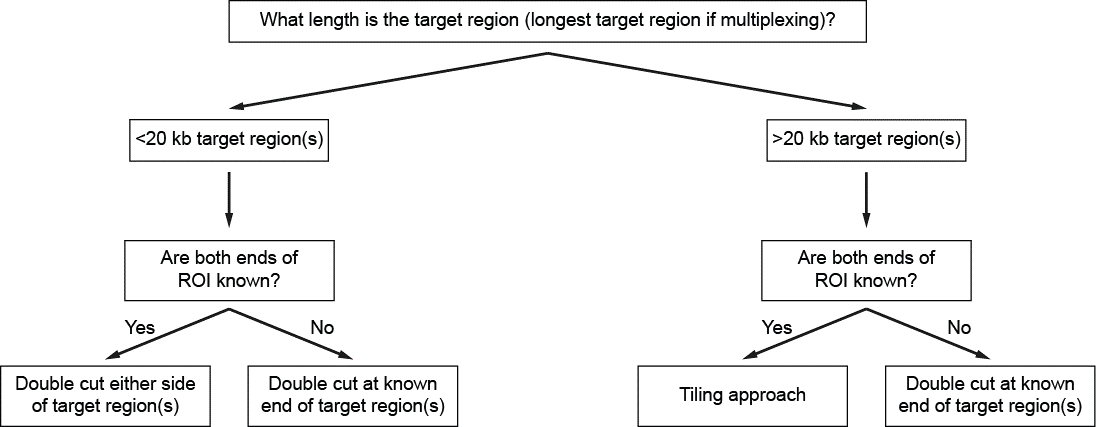
Figure 5. Decision tree for designing the Cas9 targeting approach based on the size of the target region. -
Enrichment experiment steps and associated instructions
Step Instructions 1. Extract and prepare DNA Extraction methods 2. Design probes Targeted, amplification-free DNA sequencing using CRISPR/Cas (Probe design) 3. QC input DNA Input DNA/RNA QC 4. Perform enrichment, and prepare sequencing library Cas9 Sequencing Kit (SQK-CS9109) - this is the protocol using the Oxford Nanopore Technologies Cas9 Sequencing Kit for Cas9 enrichment and library preparation
Cas9 targeted sequencing - this is the protocol that uses the Ligation Sequencing Kit (SQK-LSK109) for library preparation after Cas9 enrichment. It requires more 3rd party reagent than if doing library preparation using the Cas9 Sequencing Kit5. Sequence on device Cas9 Sequencing Kit (SQK-CS9109)
Cas9 targeted sequencing6. Take basecalled FASTQ files into analysis pipeline Cas9 Targeted Sequencing Tutorial (EPI2ME Labs) 7. Assess success of experiment and feed back into probe design and quality of input




