-
crRNA probe design guidelines
Independent of which Cas9 cut method is used, the probe design process is the same and the same features that make a good probe apply to all. The diagram below shows what the RNP complex looks like bound to the target sequence (in black and PAM in purple, Figure 6 (1) and (2)). Upon RNP binding, the duplex is locally melted, and the crRNA hybridises to the non-target DNA (bottom strand in Figure 6 (2) which is complementary to the target and crRNA sequence). This means that the crRNA sequence is same sequence as the ROI (Figure 6 (2), shown in red). Therefore, when designing a crRNA probe, the sequence should be the same as the region of interest.
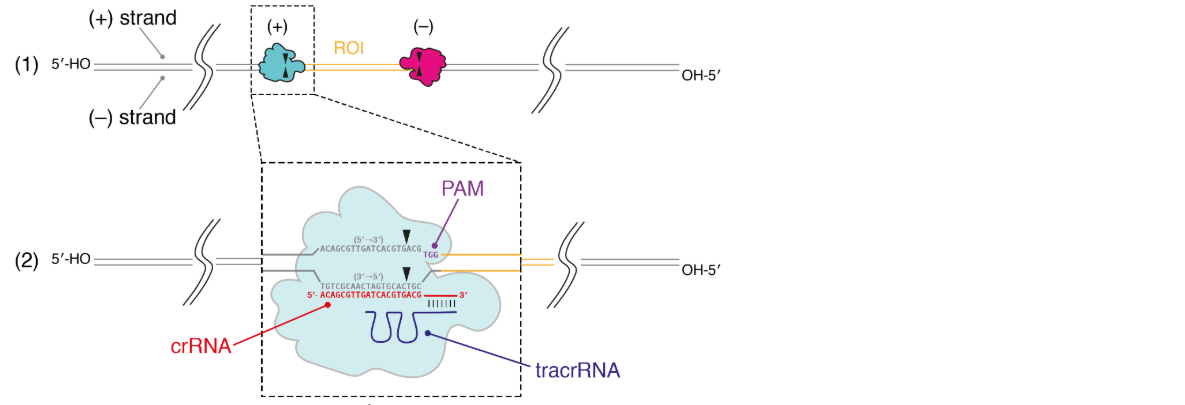
Figure 6. (1) the RNP complex bound to the region of interest. (2) The RNP melts the DNA duplex upon hybridisation with the strand complementary to the target region. The crRNA sequence is identical to the target sequence upstream of the PAM.The orientation of the probes is critical to designing a Cas9 cleavage experiment, the orientation being defined by the PAM and target sequence. Cas9 cuts at the PAM-proximal end of the protospacer 3 bp upstream of the PAM.
Figure 7 (3) shows details of the Cas9 cleavage reaction. The protection of the PAM-distal end by Cas9, which remains bound, provides the directionality for the strand but this process is imperfect. Sometimes Cas9 can release the DNA, but our internal data suggests that the reads towards the PAM site outnumber the reads away by a factor of at least 3:1, up to at least 10:1 (shown by the grey arrows in Figure 7 (3) and the small arrows in Figure 8).
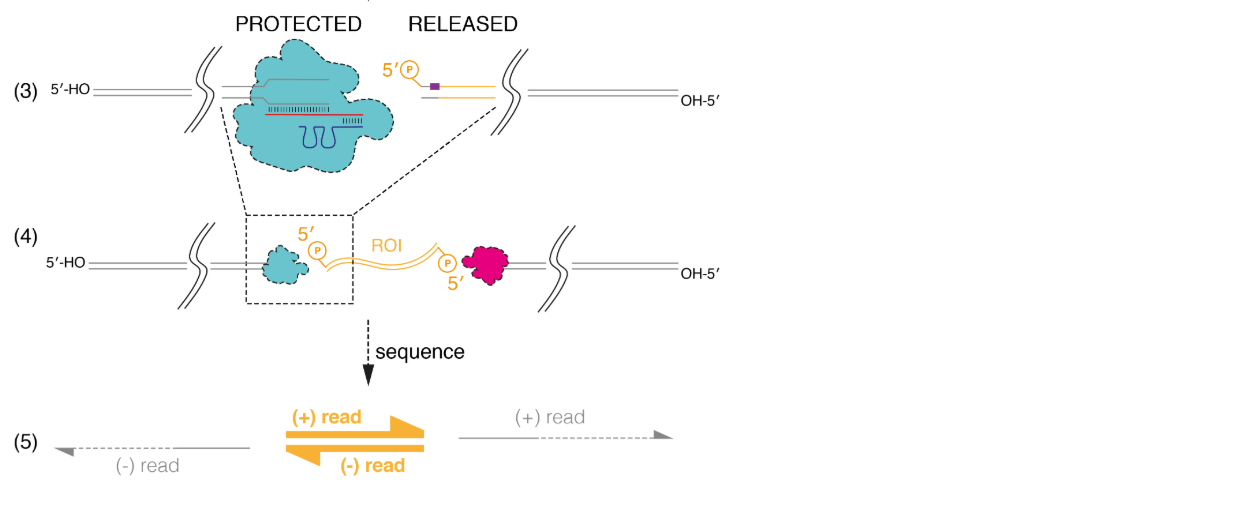
Figure 7. (3) The Region of Interest is cleaved at the 5’ side, and the PAM-distal site is protected by the bound Cas9. (4) The cleavage and protection happens on both sides of the ROI. (5) The directionality of the read is determined by the protection of the PAM-distal site by Cas9. In most cases, the ROI is enriched for, and read by the nanopore. In a minority of cases, the DNA is read in the other direction, away from the ROI.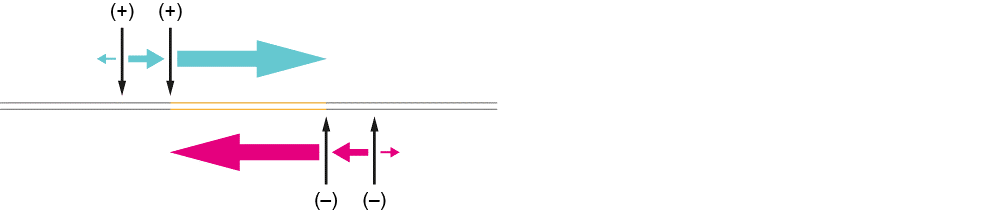
Figure 8. The directionality of the sequenced read when the Region of Interest is cleaved by the excision approach (twice on each side).To design a crRNA probe, you initially must search the region upstream, and downstream if doing an excision approach, of the region of interest for a PAM sequence (NGG). We recommend a minimum of 1 kb flanking region between the region of interest and probe target sequence. This is very important when looking at repeat expansions and regions of low complexity to help with scaling of signal during basecalling, and this flanking region can be expanded if the region of interest is very small and could be lost in subsequent clean-ups (<3 kb). All crRNA probe sequences should be checked against the whole target genome to see if there are off-target alignments, which would result in cuts elsewhere in the genome. Assess if there are any known SNPs in the crRNA probe target sequence, as these SNP mismatches will reduce the efficiency of a probe, potentially causing the absence of a cut at the region of interest in non-reference samples.
-
Panels of regions of interest
Cut kinetics are acceptable for panels of up to ~100 crRNAs. Single or multiple gene fragments may be targeted in an experiment. To excise a ROI, at least four crRNAs are required, but up to a hundred or more may be used. For simplicity, we advise users to pre-mix crRNAs at equimolar concentration and form all ribonucleoprotein complexes (RNPs) simultaneously in a single tube (unless performing a tiling experiment where each pool should be kept separate). The greater the number of crRNAs in a single cut reaction, the lower the concentration of each probe, and therefore the slower the cut kinetics. The reaction is performed at excess RNPs with respect to target. In-house data at Oxford Nanopore Technologies shows that the kinetic efficiency of the cut reaction is complete within ~15 min and acceptable for panels of up to ~40 crRNAs. Larger panels may require longer cut times.
-
Designing and ordering probes
We recommend that even users familiar with existing CRISPR design criteria read the section below carefully, because there are rules specific to nanopore target enrichment.
Oxford Nanopore Technologies recommends free crRNA probe design tool CHOPCHOP (chopchop.cbu.uib.no) and ordering IDT Alt-R™ Cas9 probes: idtdna.com/crispr-cas9. -
CHOPCHOP
CHOPCHOP (chopchop.cbu.uib.no) identifies all possible target sites, based on the availability of PAM sequences, and evaluate each site for the efficiency of cutting and possible off-target effects (based on experimental data). A CHOPCHOP search would normally involve:
Input parameters (user-defined):
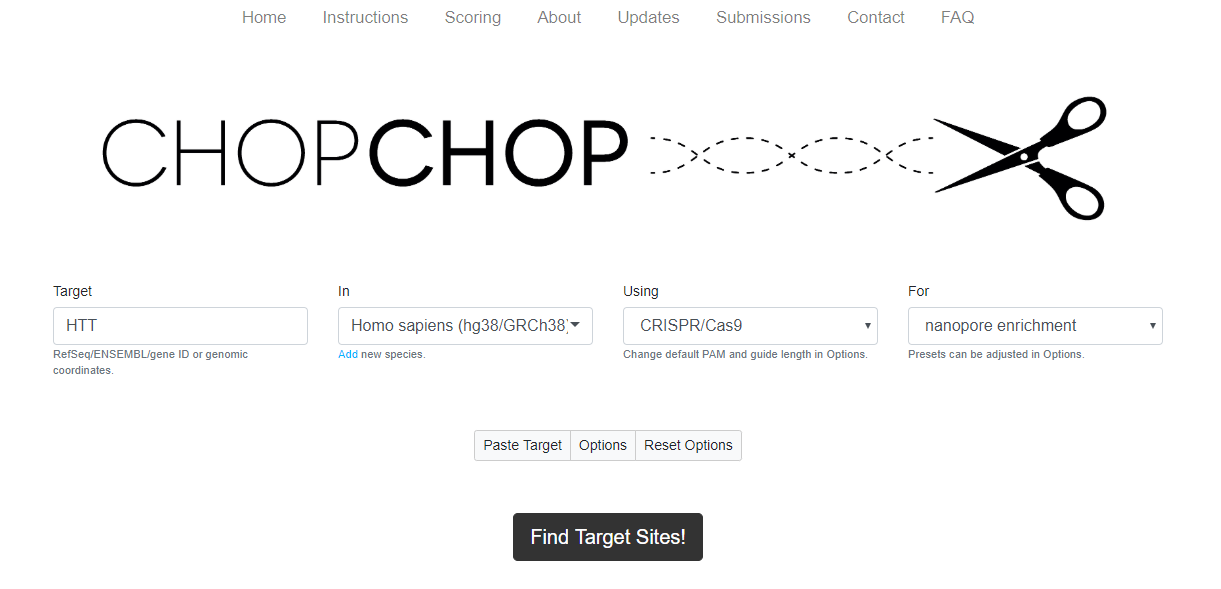
Figure 9. The CHOPCHOP homepage, with the fields selected for CRISPR/Cas-based nanopore enrichment.- A FASTA sequence or specific genomic coordinates. Remember to include 1 kb flanking regions, and a separate search is required for probes upstream and probes downstream of the region of interest (‘Target’ box on CHOPCHOP homepage)
- The target genome to search for off-target sites (‘In’ box on CHOPCHOP homepage)
- Which CRISPR/Cas system to use (‘Using’ and ‘For’ box on CHOPCHOP homepage. Select ‘CRISPR/Cas9’ and ‘nanopore enrichment’)
- Manual parameters, e.g. the PAM, protospacer length, and the algorithms used to score the efficiencies of probes (select ‘Options’ and change Efficiency score to ‘Doench et al. 2014 – only for NGG PAM’)
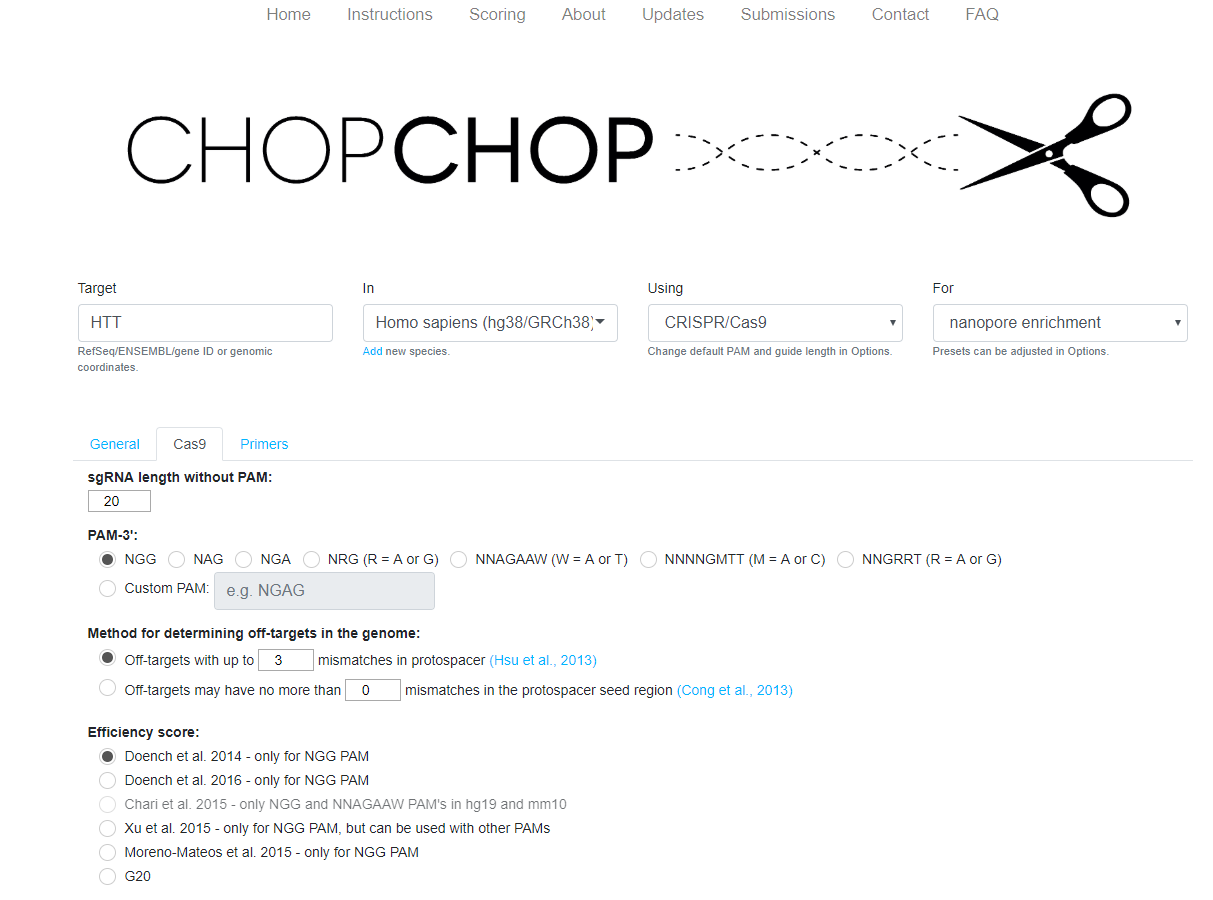
Figure 10. Recommended parameters in CHOPCHOP for probe design.The search tool, performed within CHOPCHOP:
- Identifies all possible protospacer candidates based on the locations of PAM sequences
- Scores and ranks them based on:
- Predicted efficiency
- Self-complementarity (how likely the crRNA is to form unwanted secondary structure)
- Number of off-target sites in the genome bearing 0, 1, 2 or 3 mismatches relative to the candidate protospacer sequence, as judged by an alignment of each candidate protospacer sequence against the entire genomeFiltering the search results, user-performed: to remove candidates with significant secondary structure, off-target effects, or low efficiency.
-
Using CHOPCHOP to design probes for a tiling approach
CHOPCHOP will only allow for a search within 20 kb at a time. For loci >20 kb, multiple searches should be performed, but the results from multiple searches can be pooled. This is required if using a tiling approach.
CHOPCHOP returns the target sequence, i.e. the protospacer + PAM sequence. For S. pyogenes Cas9, this will be a 20mer sequence + NGG. Cas9 will not cleave the target if the PAM is included in the crRNA sequence.
Example for Cas9: if target = GTTAGTGTCCCCATACAACG*GGG*, the 20mer crRNA = GTTAGTGTCCCCATACAACG.
-
Probe ordering: IDT
Oxford Nanopore Technologies recommends ordering IDT Alt-R™ Cas9 probes: idtdna.com/crispr-cas9. Alt-R™ probes contain proprietary modifications that increase the stability of the crRNA and its resistance to nuclease-mediated degradation.
The crRNA requires a short sequence at the 3’ end of the protospacer sequence that hybridises to tracrRNA; however, this sequence is automatically applied to the crRNA sequence at the time of ordering; only the protospacer sequence (no PAM, as DNA provided by CHOPCHOP) is required. As an RNP complex is required to perform a Cas9 experiment, users must ensure that they also order tracrRNA (IDT Alt-R™, Cat # 1072532, 1072533 or 1072534) and S. pyogenes Cas9 (Alt-R® S. pyogenes HiFi Cas9 nuclease V3 IDT Cat #1081060 or 1081061).
-
Replacing IDT synthetic crRNA and tracRNA is not recommended by Oxford Nanopore Technologies for the following reasons:
- A 5' G is needed to make the in vitro transcript, which limits the candidates by 4-fold. Adding one where it does not already exist causes issues.
- Transcribing RNA in vitro can yield highly variable results.
If you would still like to try, skip the annealing step and add 500 nM final sgRNA during the RNP formation step.




