-
Analysis workflow
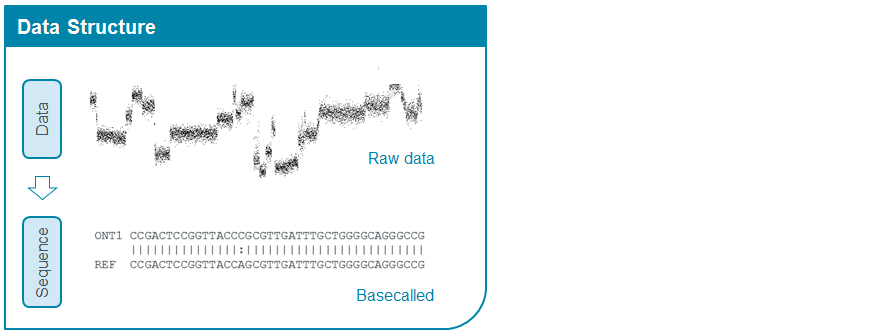
MinKNOW acquires raw signal from the device, and sends it to the analysis pipeline (basecaller) in chunks of a defined size. Before being sent to the pipeline, MinKNOW assesses the data and determines where the individual reads start and end by detecting the abrupt signal change when DNA enters and leaves the nanopore. Only data that is considered to be within a read is sent for analysis.
-
The format and location of your data will depend on the options chosen in the new experiment settings screen:
All files output directly from your experiment are located in the same directory. The directory has the structure:
{output_dir}/{experiment_id}/{sample_id}/{start_time}_{device_ID}_{flow_cell_id}_{short_protocol_run_id}/
-output_diris the configured output directory
-experiment_idis the user-entered identifier for a group of runs
-sample_idis the user-entered value for a specific sample or run. It is intended that multiple sample_ids may exist beneath an individual experiment_id
-start_timeis the time at which the protocol started in YYYYMMDD_HHMM format
-device_idis the serial ID of the MinION Mk1B or device position for GridION/PromethION
-flow_cell_idis the flow cell id (eg: FAH12345), either programmed on the ASIC or entered by the user
-short_protocol_run_idis a unique identifier of 7 characters from the protocol IDExamples of the above naming convention:
/data/MyExperiment/Sample1/20181011_1759_X1_FAH12345_0ffe109Individual read files will be split into .fast5 pass and fail folders, as well as .fastq pass and fail folder within this directory:
{flowcell_id}_{basecall_state}_{short_run_id}_{batch_number}.fastq
{flowcell_id}_{basecall_state}_{short_run_id}_{batch_number}.fast5If the reads are barcoded, the barcode will be included in the file name before the
short_run_id.
Examples of the above file naming:fast5_pass/FAK12345_pass_9aad0448_0.fast5
fasq_pass/FAK12345_pass_9aad0448_0.fastqRun reports can be found in the data folder (e.g. C:\data\reports).
