-
Alignment overview
A user can align their data directly in MinKNOW after a sequencing experiment has finished, using a FASTQ file from a previous run.
The reference file of bacterial-sized genomes must be uploaded locally as a FASTA file. Alignment hits from these files are used to populate the alignment graphs.
A BED file may also be uploaded alongside the reference FASTA file when the user is interested in a particular region of the reference (e.g. specific gene in the chromosome). The BED file alignment hits will be highlighted in the sequencing .txt file generated in the data folder.
When alignment is run independently from basecalling, BAM files are generated.
-
Accessing post-run alignment
Post-run alignment can be accessed through alignment menu and selecting Alignment.
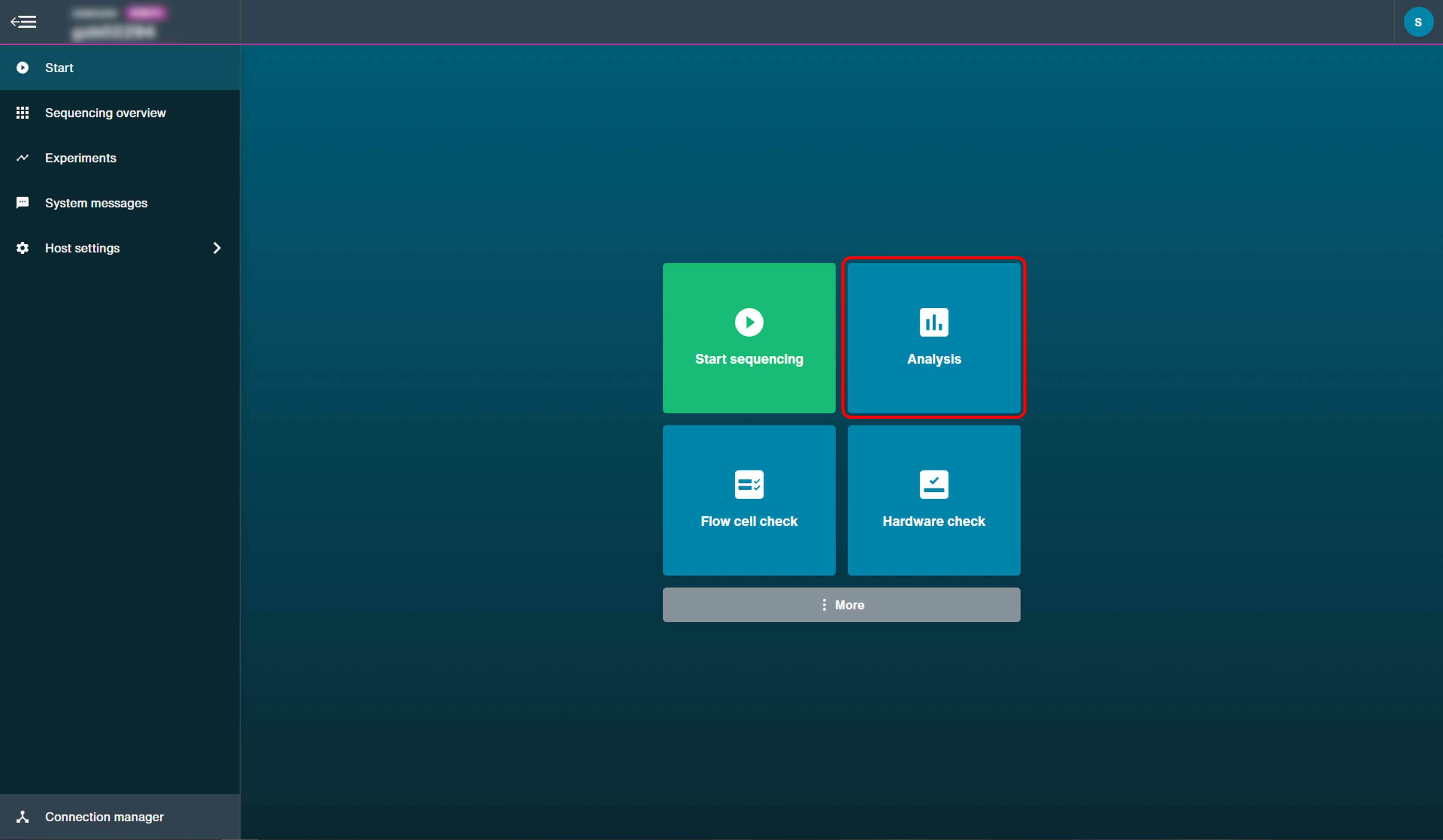
The options provided for alignment data once a run has finished are described in the MinKNOW protocol under 'Post-run analysis'.
