-
MinKNOW GUI introduction
The MinKNOW™ software carries out several core tasks: data acquisition, real-time analysis and feedback, basecalling, data streaming, providing device control, and ensuring that the platform chemistry is performing correctly to run the samples. MinKNOW takes the raw data and converts it into reads by recognition of the distinctive change in current that occurs when a DNA strand enters and leaves the pore. The reads are then basecalled, and written into .fast5, .POD5 or FASTQ files. Post-run analysis can now also be carried out through the GUI without needing to use the command line.
-
Login screen
Before a device is connected, the following screen appears:
We recommend users to log into the MinKNOW software using their Community credentials.
If you experience login issues, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community and use Continue as guest for temporary use.
-
Sequencing Overview
This page displays the inserted flow cell state and progress of a current sequencing experiment, including pausing, pore scan and basecalling.
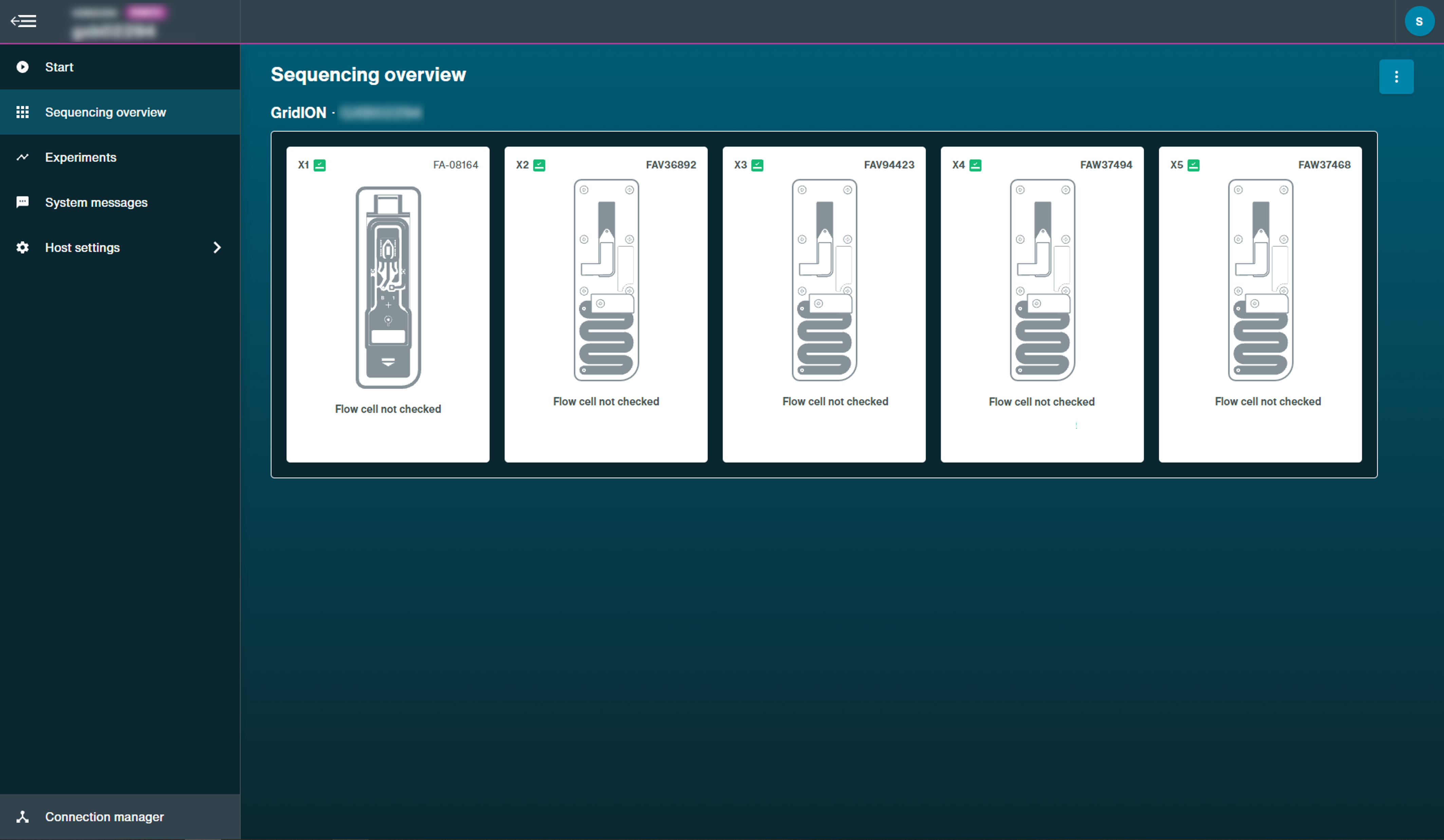
Flow cell with no checks:
This example is on a GridION device.
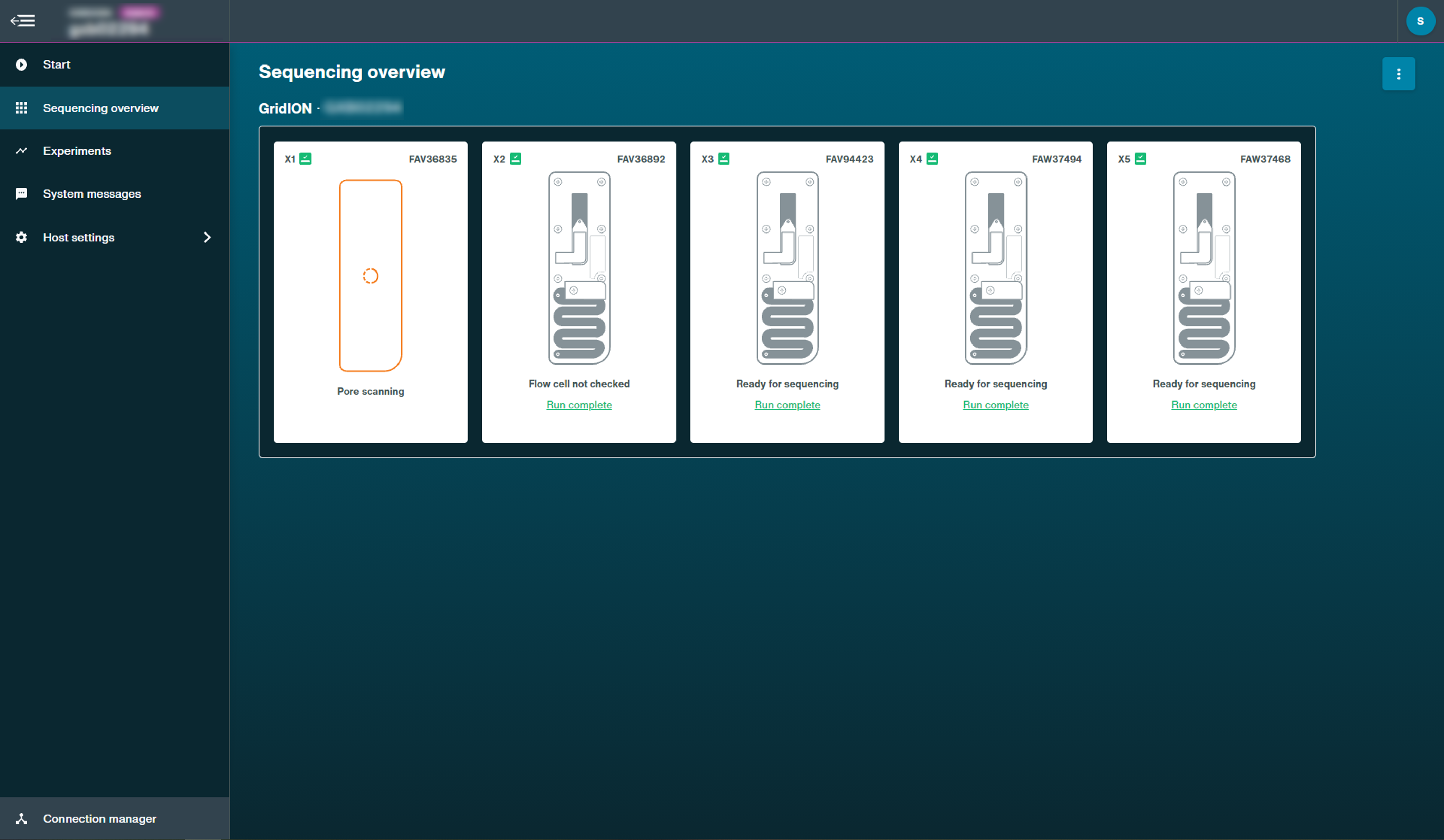
Sequencing states:
These examples are on a GridION device.Pore scan:
Flow cell health after the first pore scan:
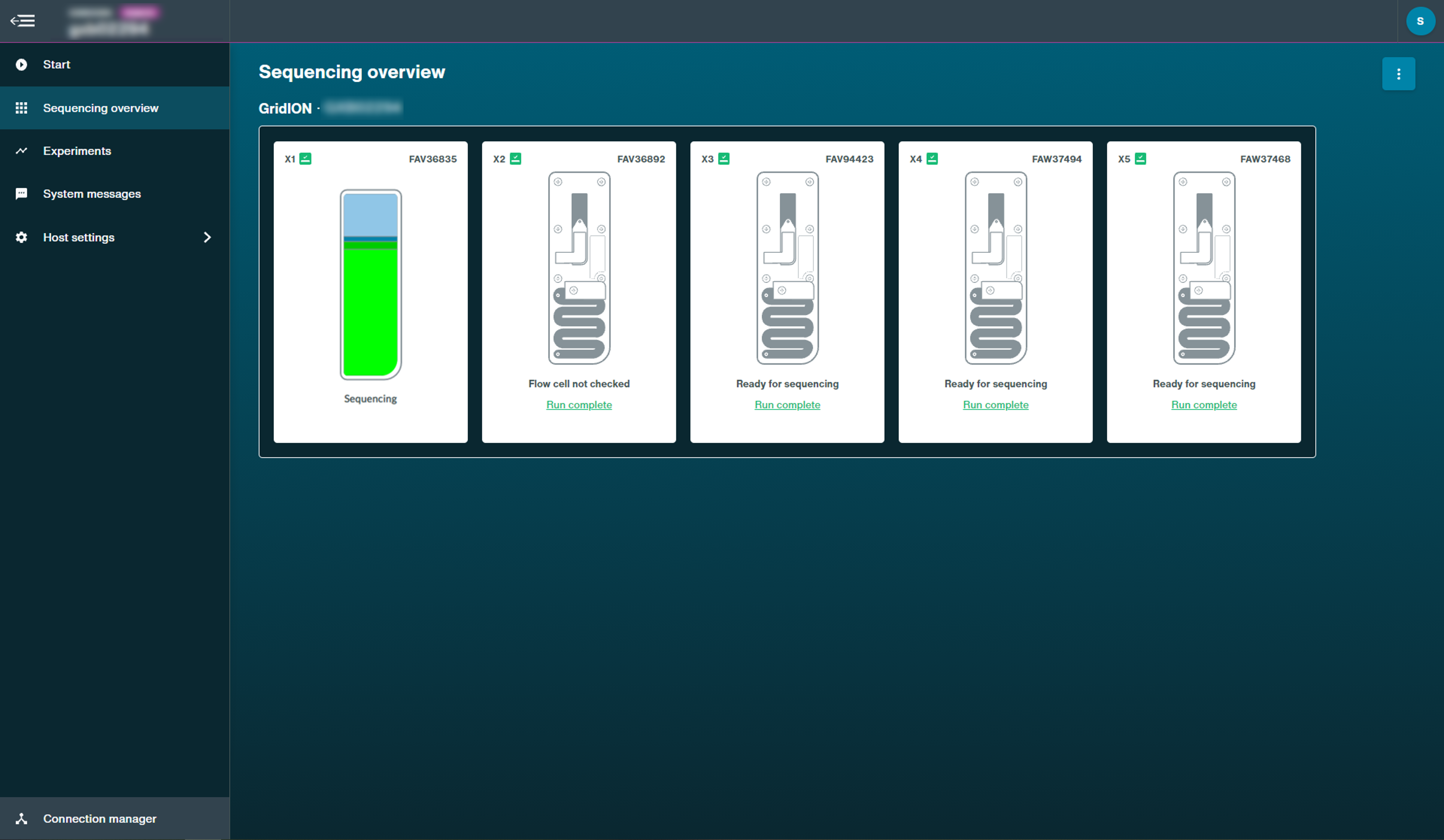
Run paused:
Basecalling catch up post-sequencing:
Click the flow cell to open the quick view of the current experiment.
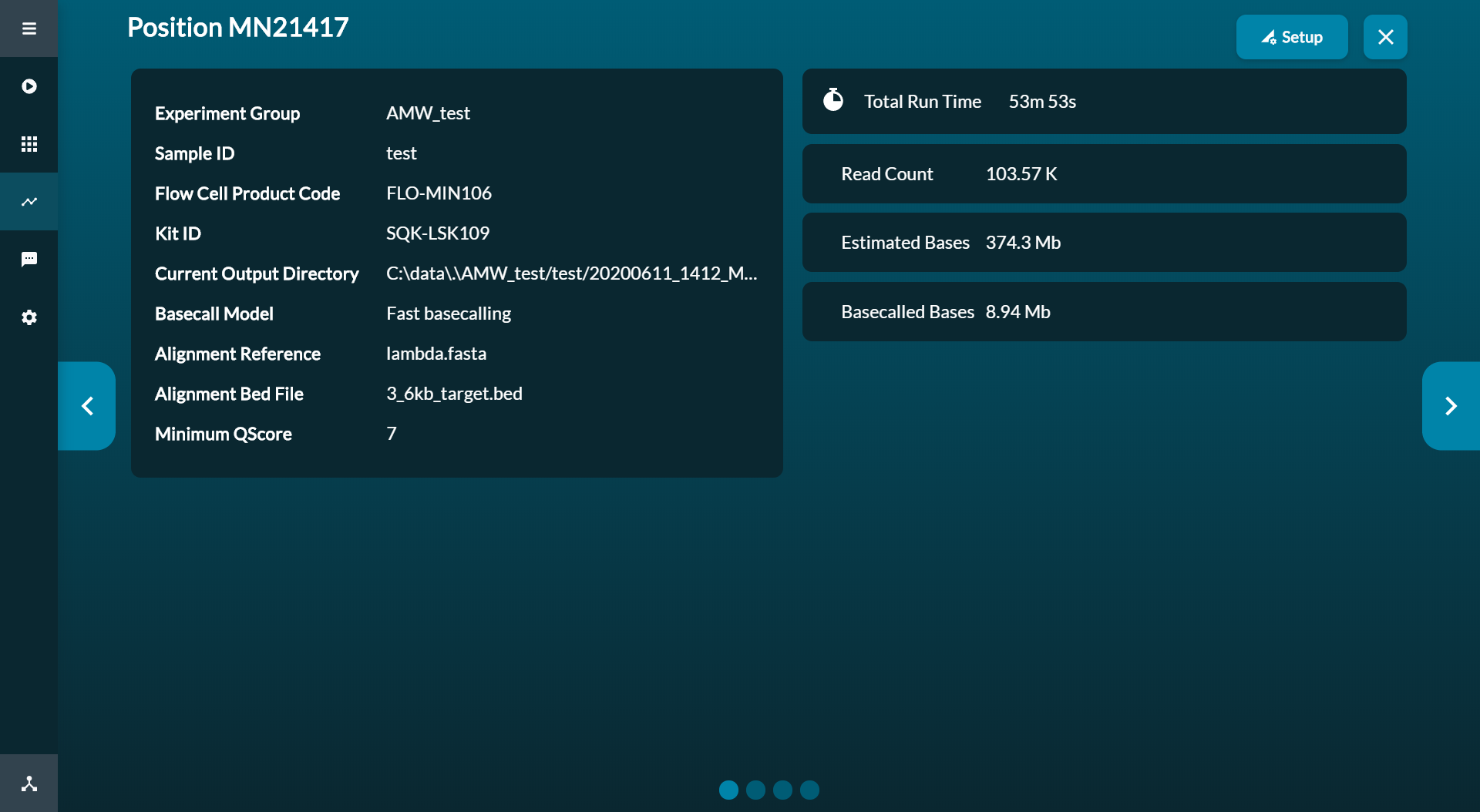
-
Experiment configuration
To start an experiment, the user clicks Start Sequencing to open the experiment configuration options.
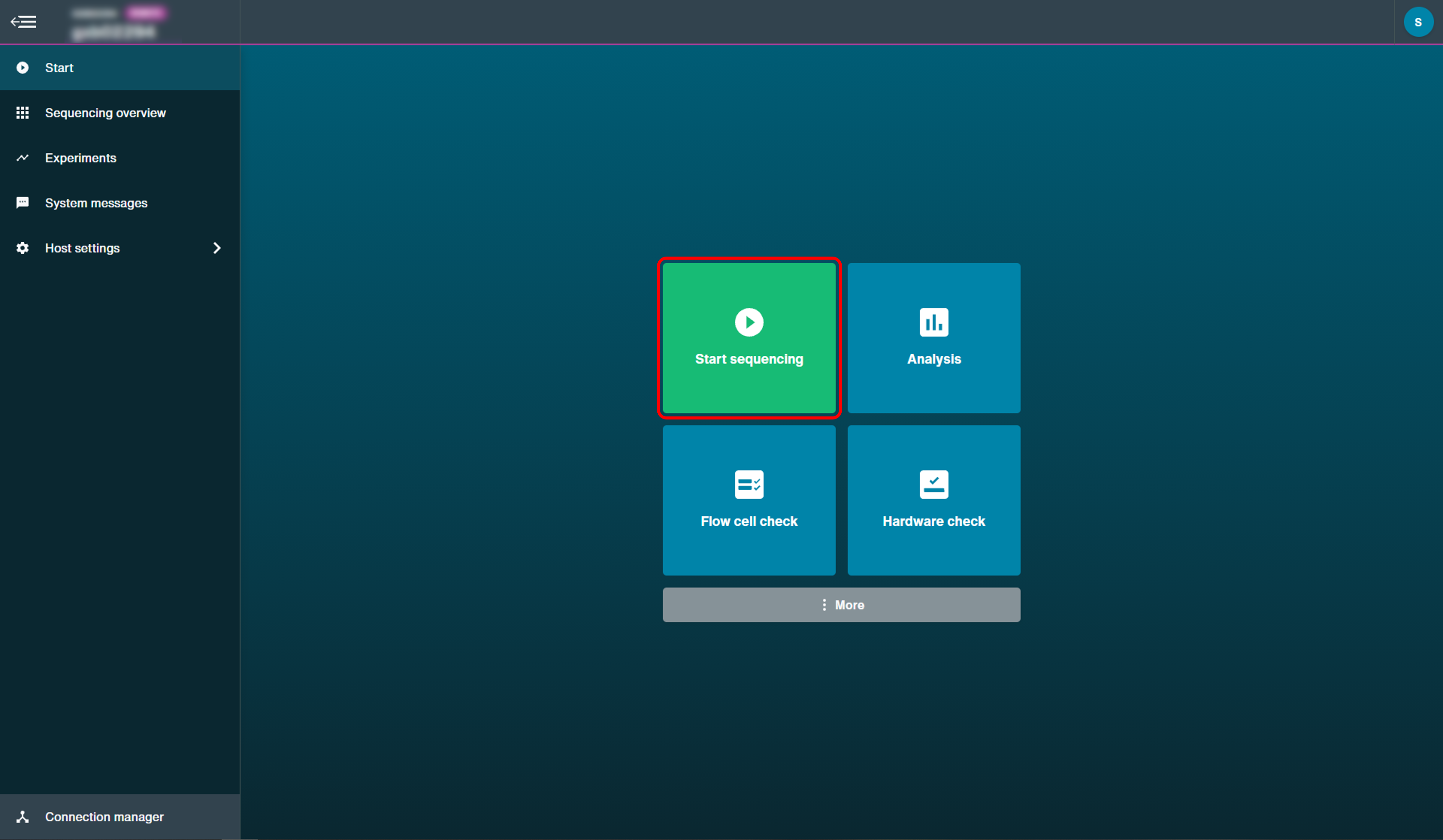
There are a number of parameters to be configured:
Flow cell position and experiment name
The "Positions" tab will show the chosen flow cell. An experiment name can then be assigned.
The other tabs will not become available until an experiment name has been provided.Kit selection
The "Kit" tab will provide a table of available kits. If a kit selected is compatible with a barcoding expansion pack, the barcoding options will become available for selection.Run Options
The "Run options" tab provides variables for run time, minimum read length and the option to use adaptive sampling. Adaptive sampling can be enabled from here on MinION Mk1C, GridION and PromethION.Analysis
Select whether data will be basecalled live on the instrument.
Note: MinKNOW provides the option to re-basecall .fast5 files during post-run analysis.Output
Specify where to save the output data for the run, and the file type.
-
Barcode demultiplexing
During kit selection, if you have used a barcoding kit or a barcoding expansion pack for your library preparation, MinKNOW will split your reads by barcode without having to use commandline tools. Demultiplexing places reads into barcode-specific folders.
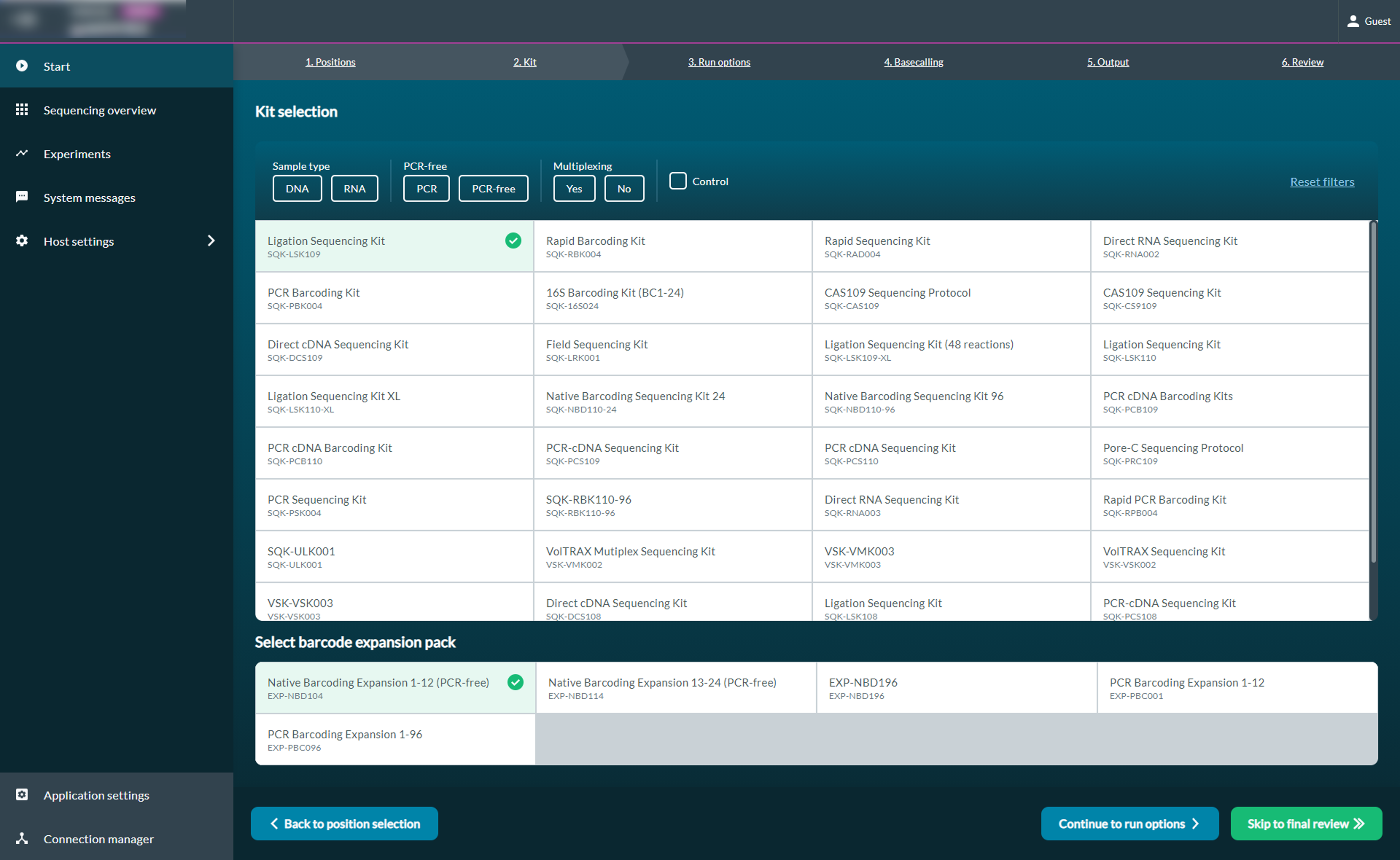
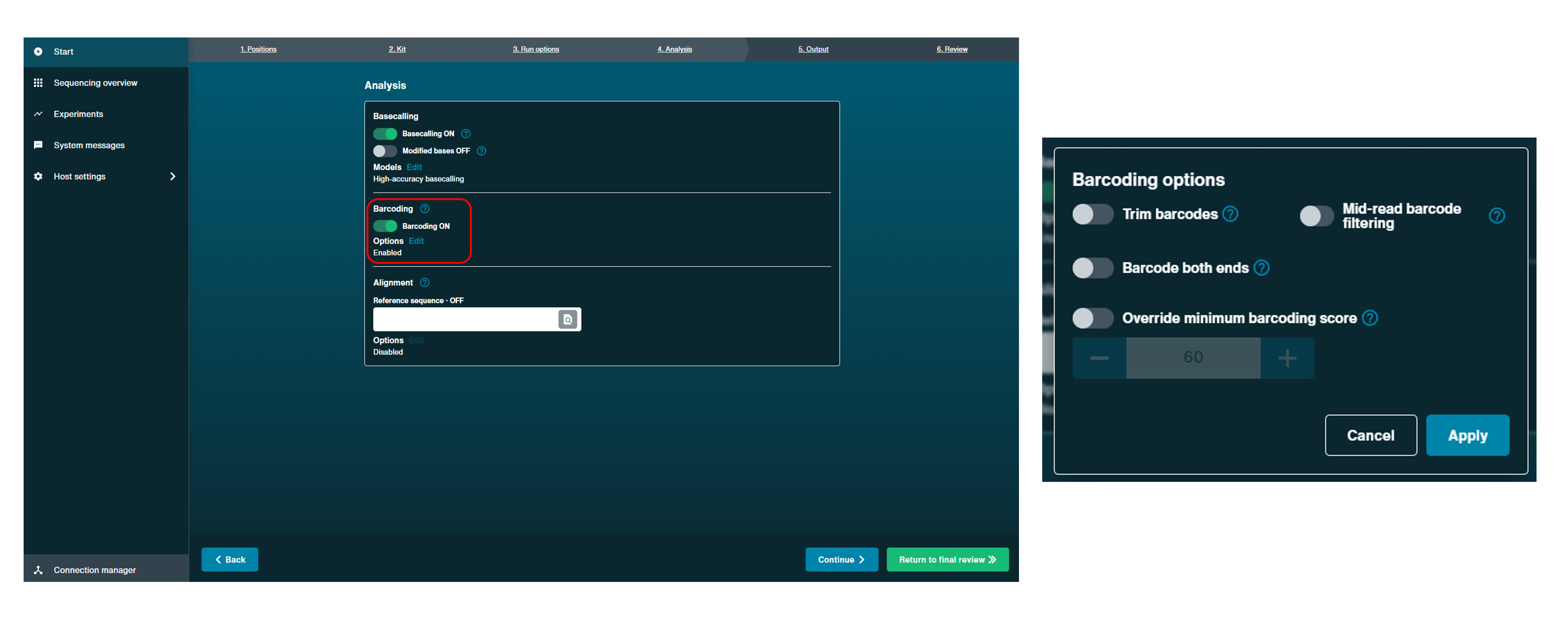
In the "Analysis" tab, the barcoding option can be toggled on and off when setting up an experiment. Further options are also available by selecting Edit options, including trimming barcodes, which can be used to enable barcode trimming in read files. Please note that some primer sequences may also be trimmed together with the barcodes.
-
Alignment
Alignment can be performed live during sequencing alongside basecalling on MinKNOW. An alignment reference must be uploaded locally on bacterial-sized genomes as a .fasta or minimap index file.
The .fasta or minimap index file can contain multiple entries within the same file (e.g. multiple chromosome). The .fasta and minimap index file alignment hits will populate the alignment graphs.
A .bed file may be uploaded alongside the reference .fasta or minimap index file. The .bed file can be used when the user is interested in a particular region of the reference (e.g. specific gene in the chromosome). The .bed file alignment hits will be highlighted in the sequencing .txt file generated in the data folder
Navigate to the "Analysis" tab and add a reference sequence in the Alignment box to turn on alignment. Basecalling must be enabled for live alignment.
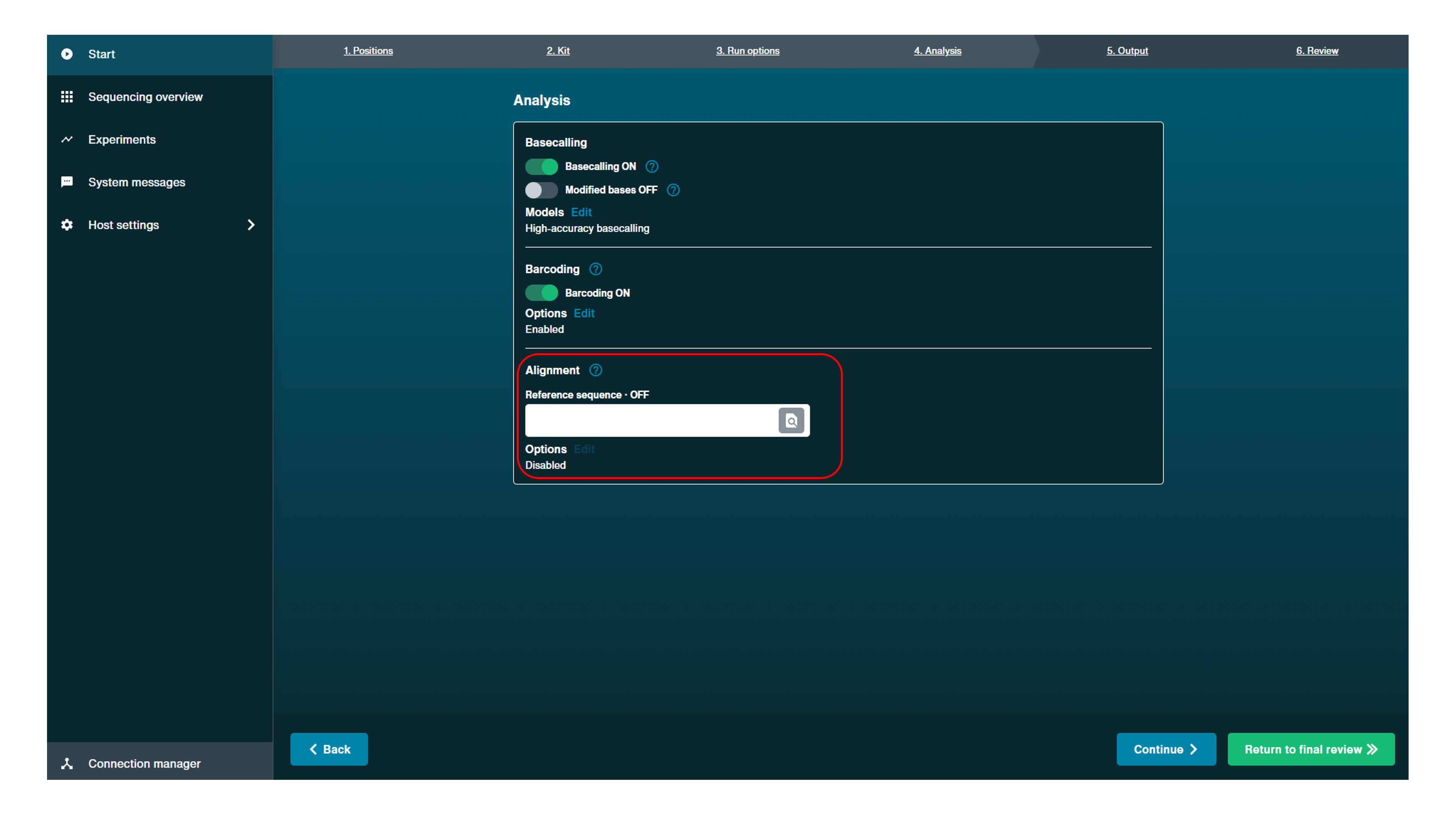
Alignment can be carried out in post-run analysis with basecalling analysis. Alignment can also be carried out independently from basecalling to generate .bam files.
For further information on post-run alignment, refer to the post-run analysis section of this protocol.
-
Flow cell health
During a sequencing experiment, the MinKNOW Sequencing Overview page shows a flow cell icon with coloured bars. The bars represent the combined health of all pores in a flow cell, and indicate how well the flow cell is performing. The colours are:
- Light green: sequencing
- Dark green: open pore
- Dark blue: pore recovering
- Light blue: pore inactive
This information is identical to the last bar of the duty time plot.
GridION flow cell health diagram

-
Experiments page
The experiments page displays summary information for all sequencing flow cells and device checks carried out on the device.
Previous runs can be viewed on the UI at the top of the screen and the number of days to view the last active experiments can be altered by typing in a different integer.

From this page, you can control specific runs and identify real-time information, including flow cell health and reads.
- Run statistics: The total number of reads, estimated and basecalled bases across an experiment, and number of active and total runs
- Run time: The duration of the experiment
- Run state: The current state of the sequencing run; 'Active', 'Basecalling', 'Complete', 'Stopped with error'
- Health: The current flow cell health
The white panel displays a summary of sequencing experiments and the blue panel displays status information of a specific run.
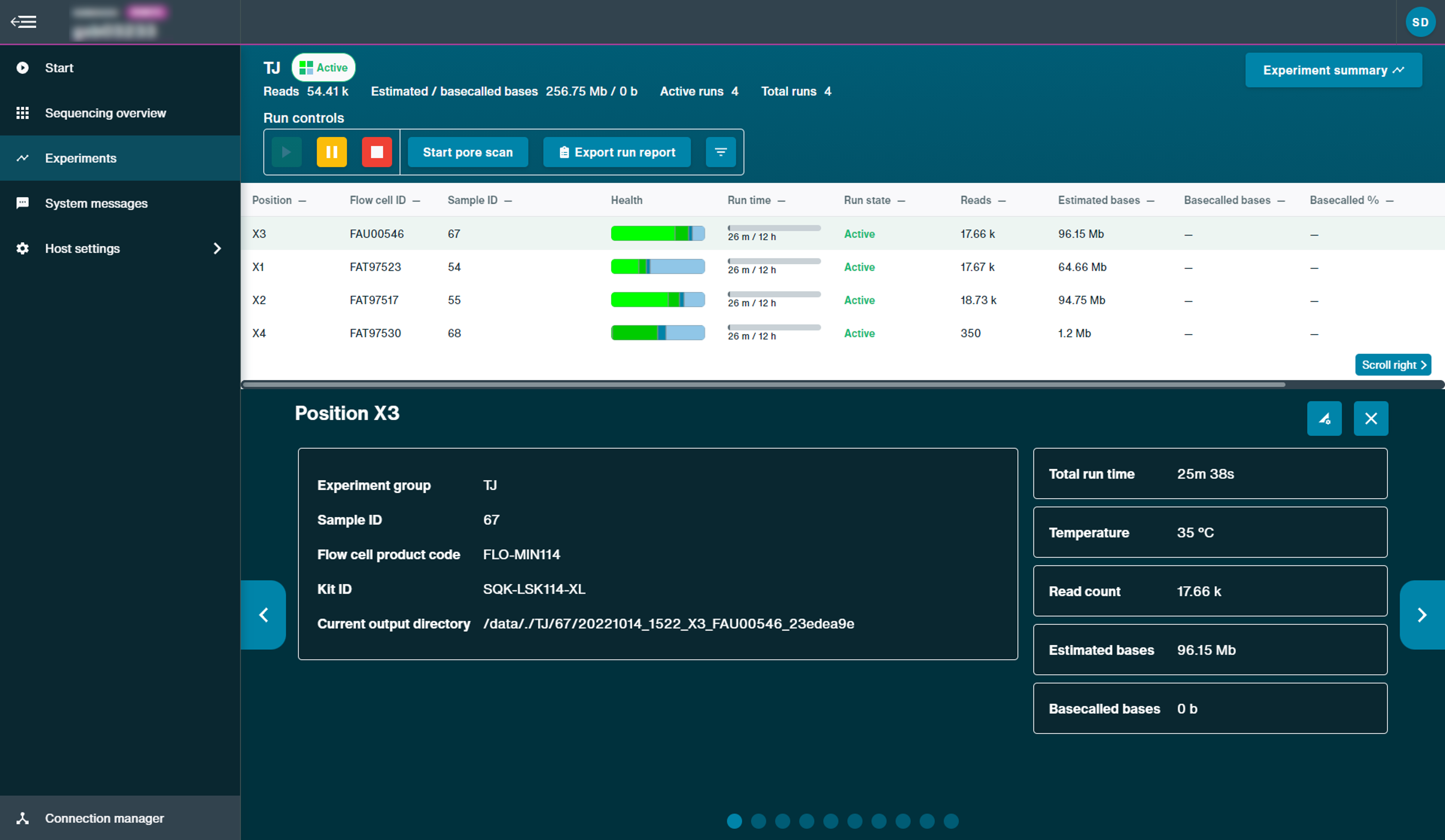
Example of experiments page on GridION:
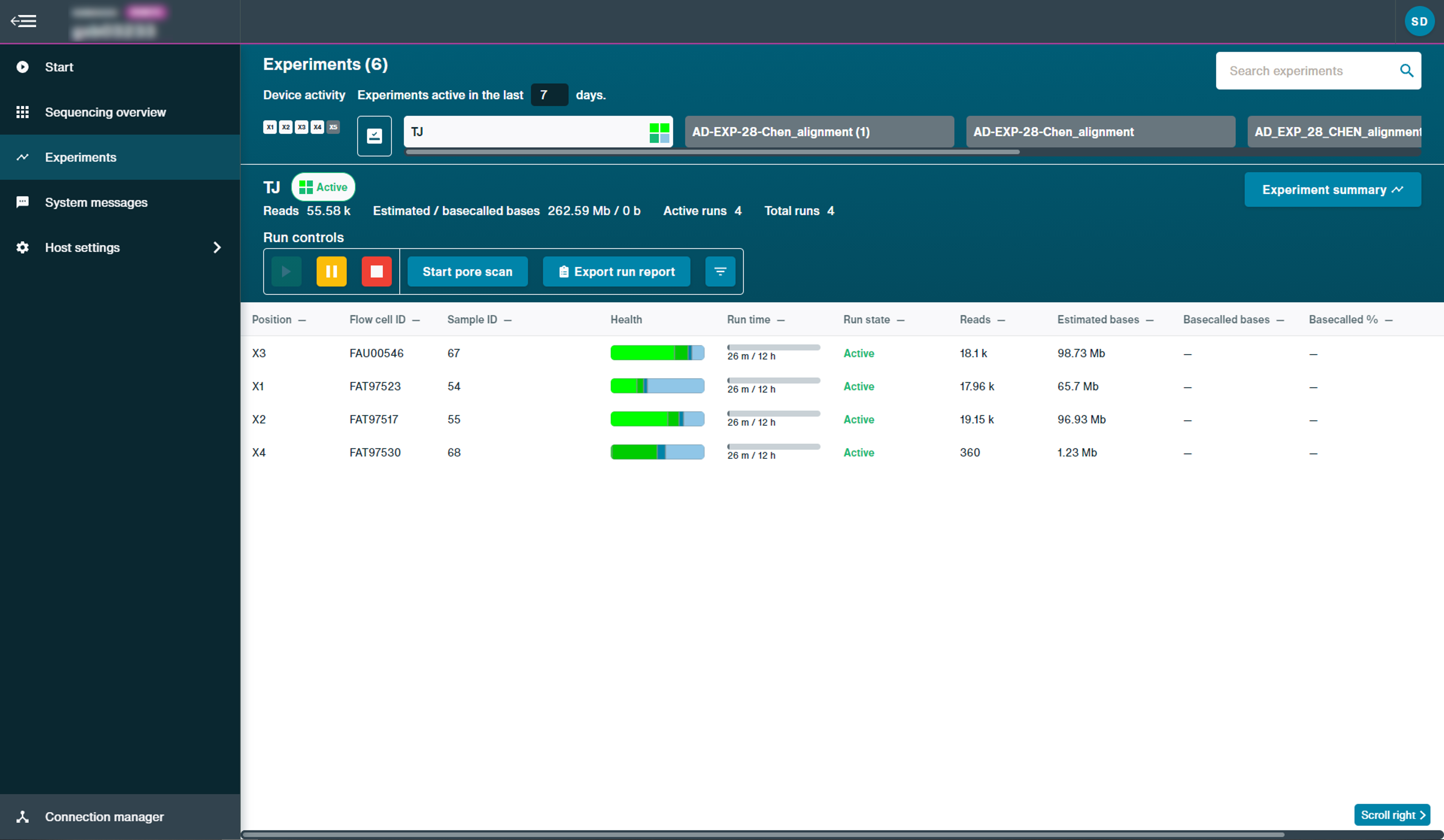
For more status information of a specific run, click the run to open the quick view, including current temperature and voltage. In the example below, the run in position X3 was clicked to open the quick view.
-
Experiment page: stats and physical layout
The experiment page shows how the run is progressing in real-time. Select a run to view the real-time data in the quick view and use the arrows to move through the graphs.
- The quick view below displays information such as the number of generated and basecalled reads, run time, temperature, and voltage.
The run time bar shows how far the experiment has progressed.

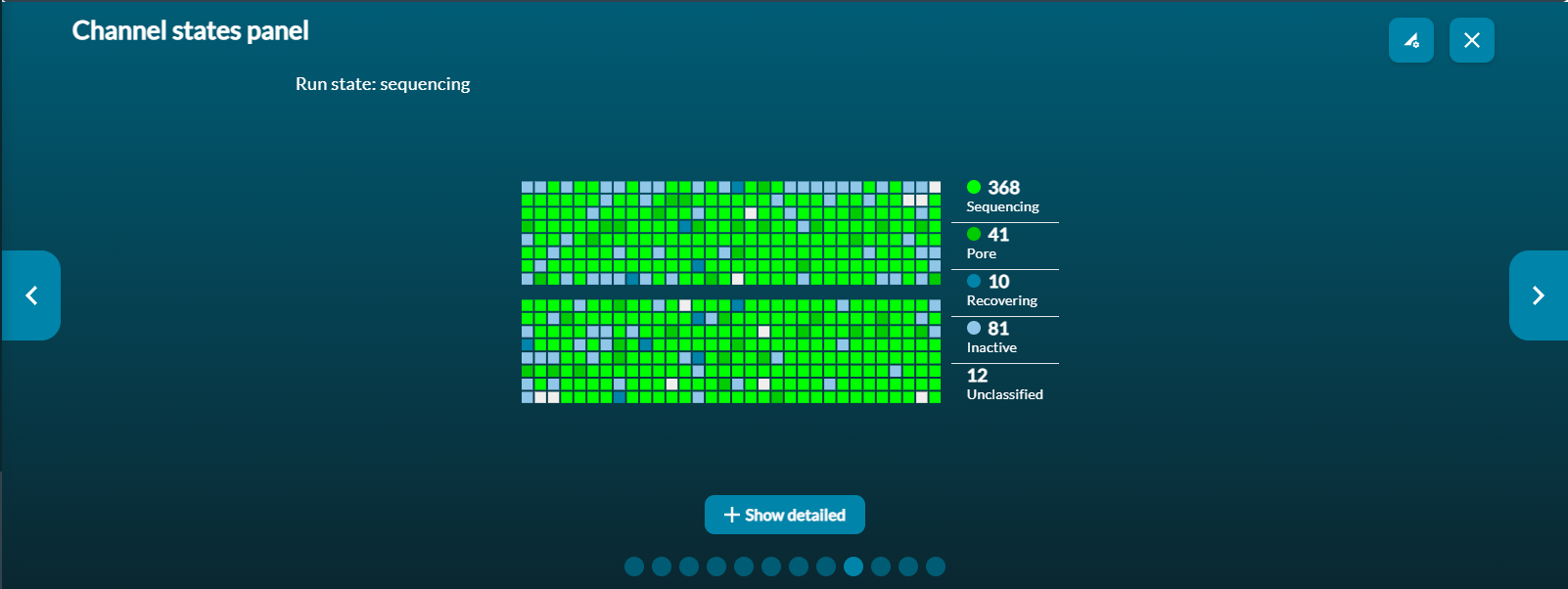
Channel panel: a real-time representation of each of the channels (e.g. 512 for MinION Mk1B)
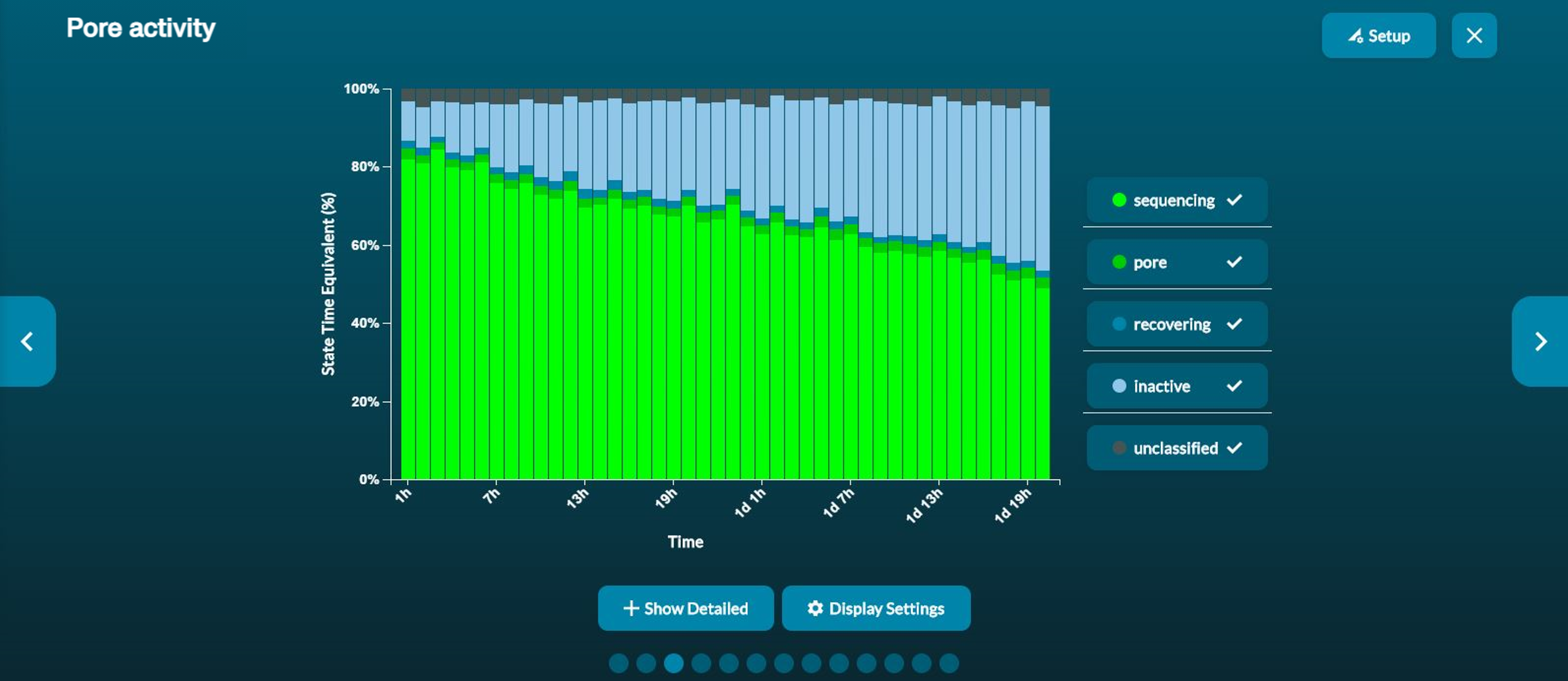
Pore activity plot: a summary of the channel states over time. This can be used to assess the quality of the sequencing experiment
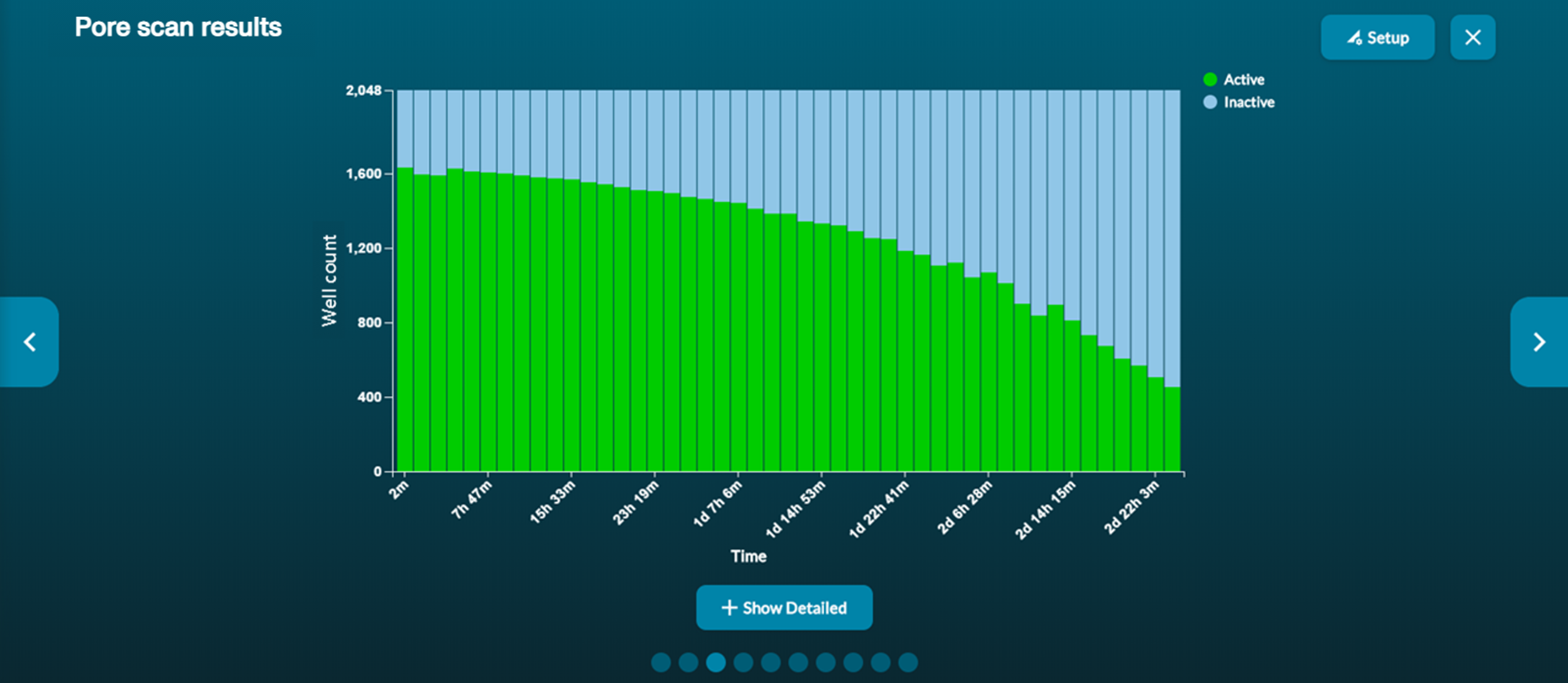
Pore scan results: a summary of the channel states during the most recent pore scan
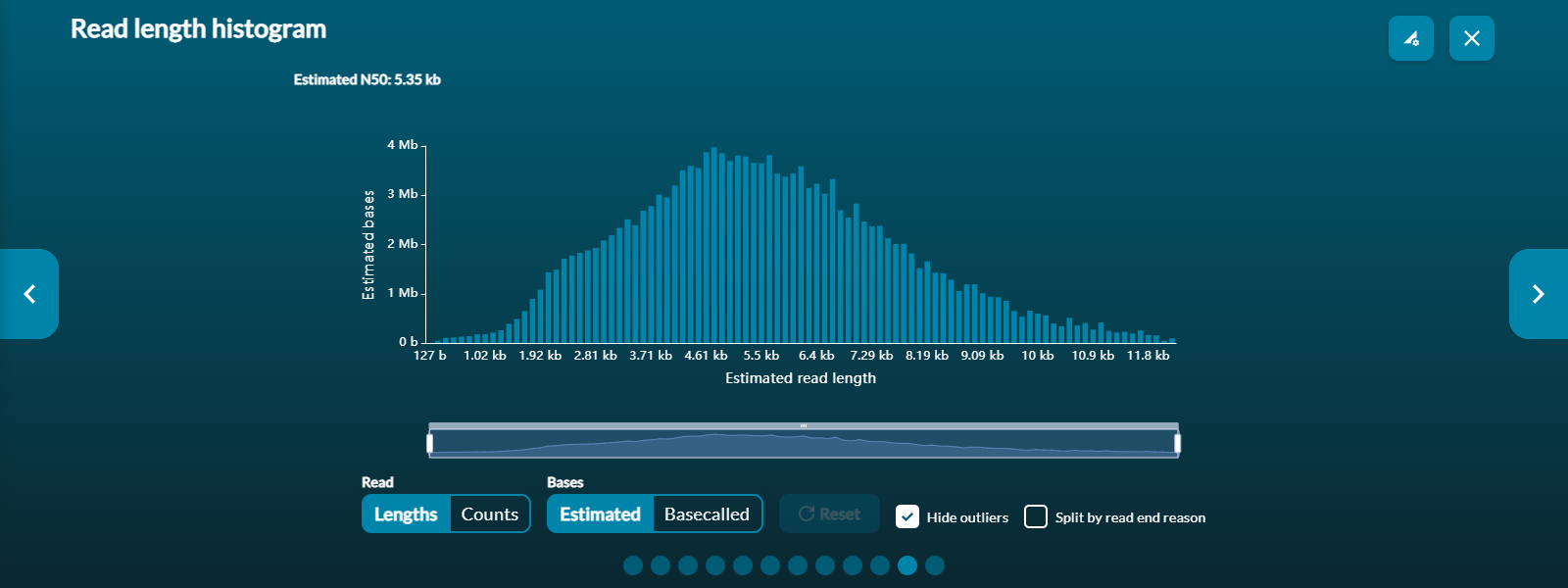
Read length histogram: a real-time representation of the reads generated so far
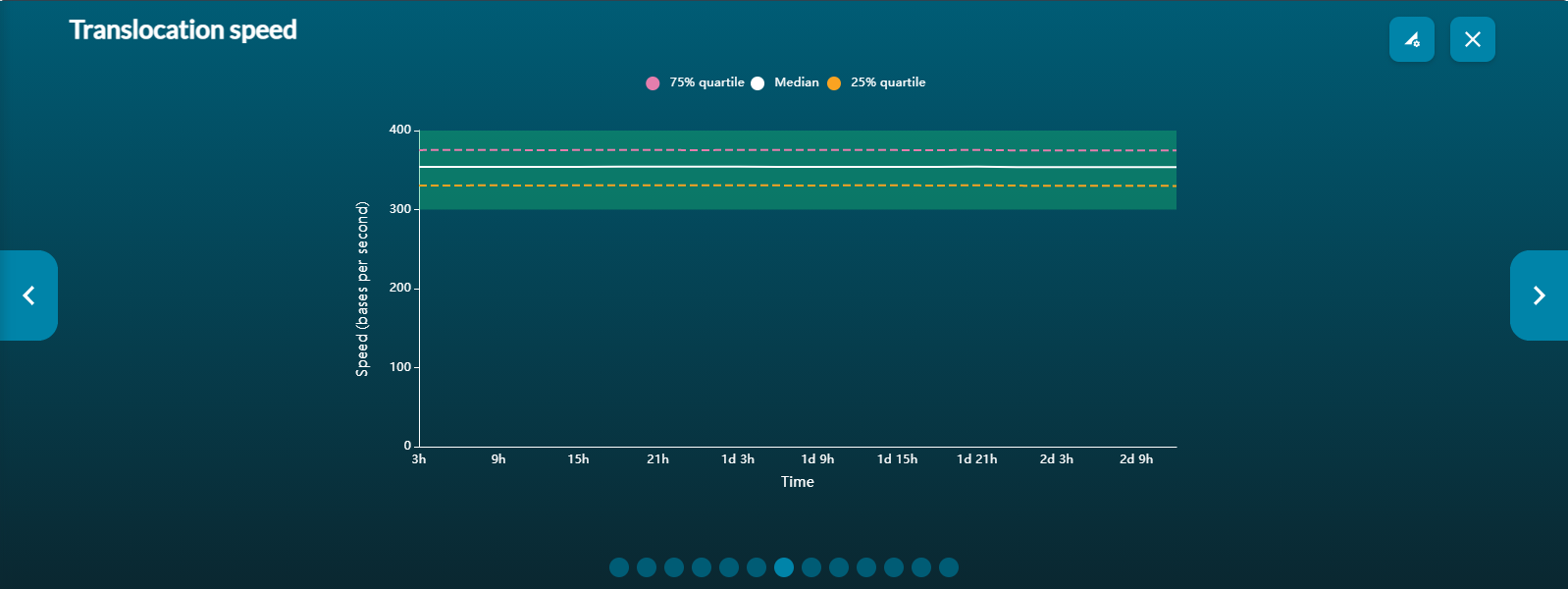
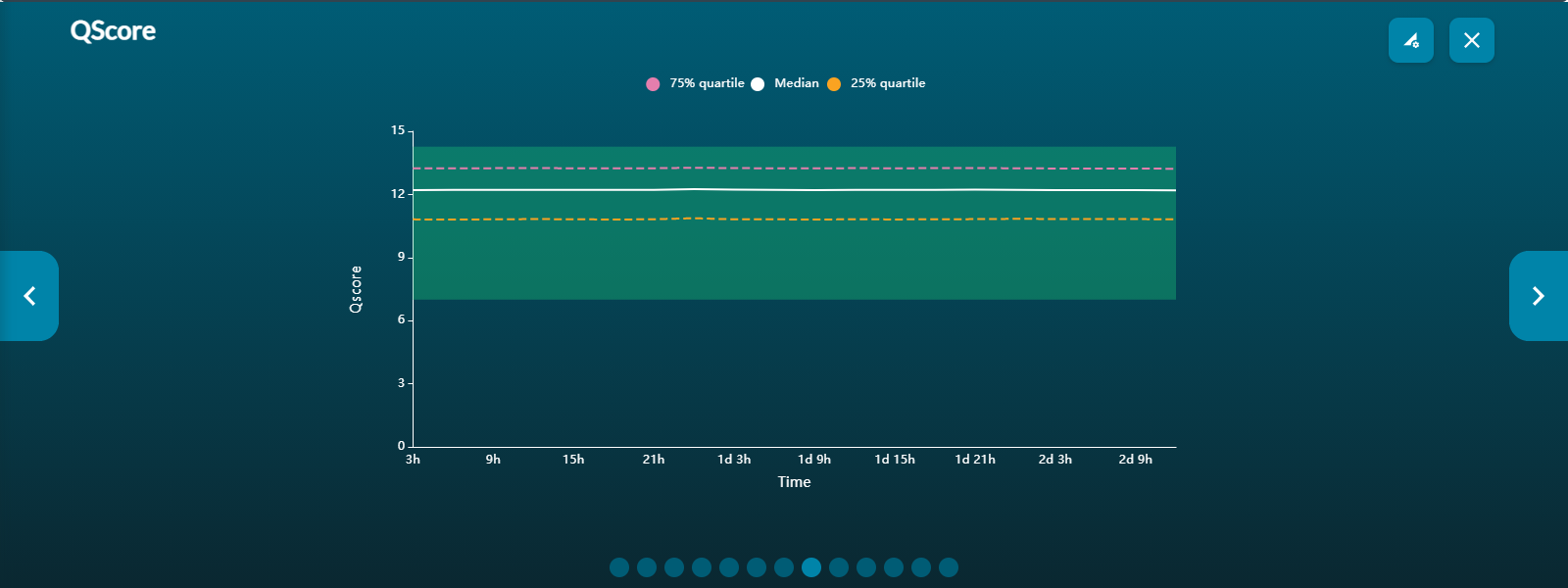
Basecalling: shows the translocation speed and Qscore against time.
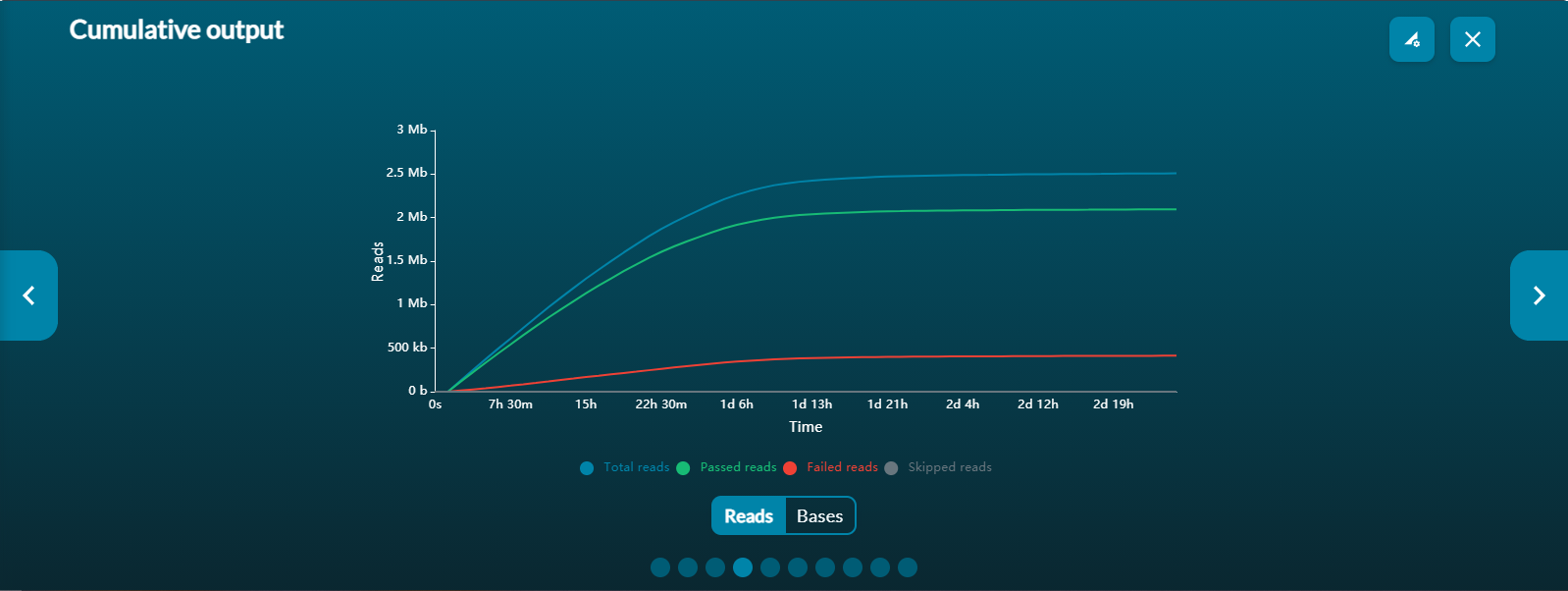
Cumulative output: shows the number of bases that have been sequenced and basecalled
Trace viewer: shows the current level fluctuations in the selected channels
-
Channel states
The channel states pannel gives an overview of the states the flow cell pores are in to give the user an idea of how well the sequencing run is performing in real time. A good library will be indicated by a higher proportion of light green channels in "Sequencing" than are in "Pore available". The combination of "Sequencing" and "Pore available" indicates the number of active pores at any point in time. A low proportion of "Sequencing" channels will reduce the output of the run.
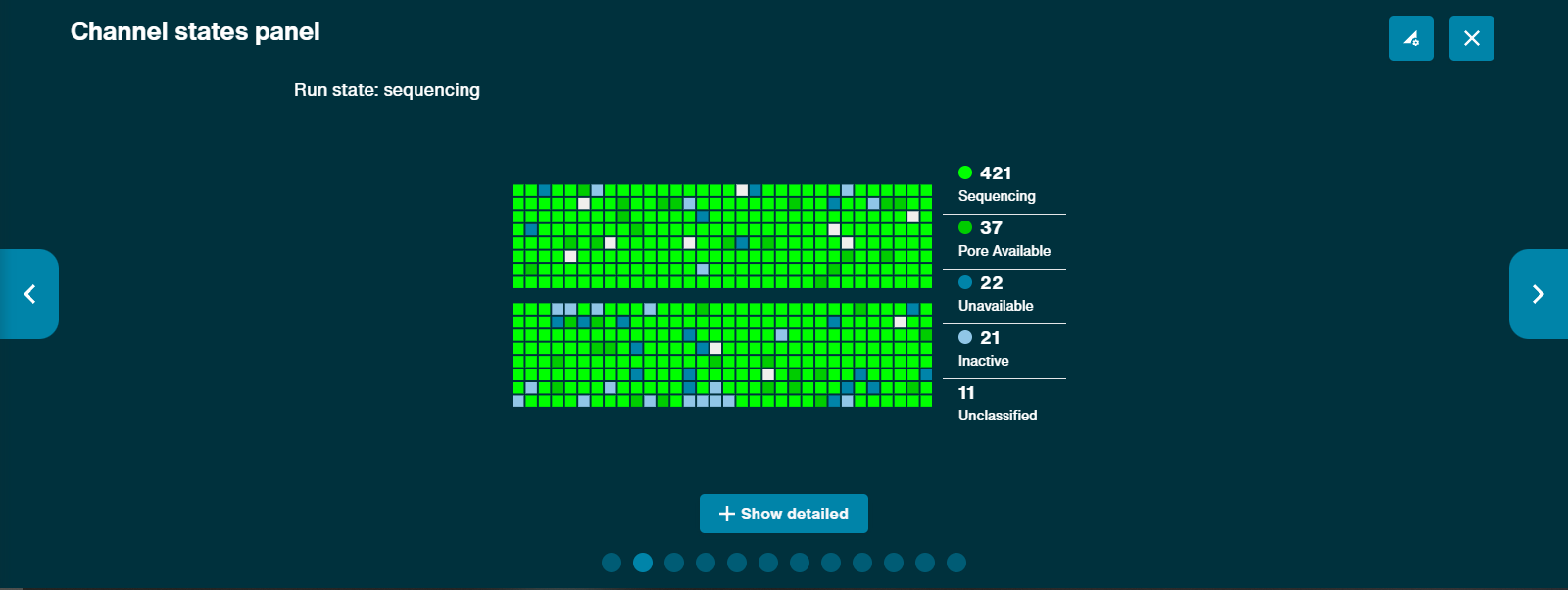
Clicking on the Show Detailed button reveals a more detailed array of channel states:
- Sequencing: Pore currently sequencing.
- Pore Available: Pore available for sequencing.
- Adapter: Pore currently sequencing adapter.
- Active feedback: Channel ejecting analyte.
- No pore: No pore detected in channel.
- Multiple: Multiple pores detected. Unavailable for sequencing.
- Unavailable: Pore unavailable for sequencing.
- Unclassified: Pore status unknown.
- Saturated: The channel has switched off due to current levels exceeding hardware limitations.
- Out of range-high: Current is positive but unavailable for sequencing.
- Out of range-low: Current is negative but unavailable for sequencing.
- Zero: Pore currently unavailable for sequencings.
-
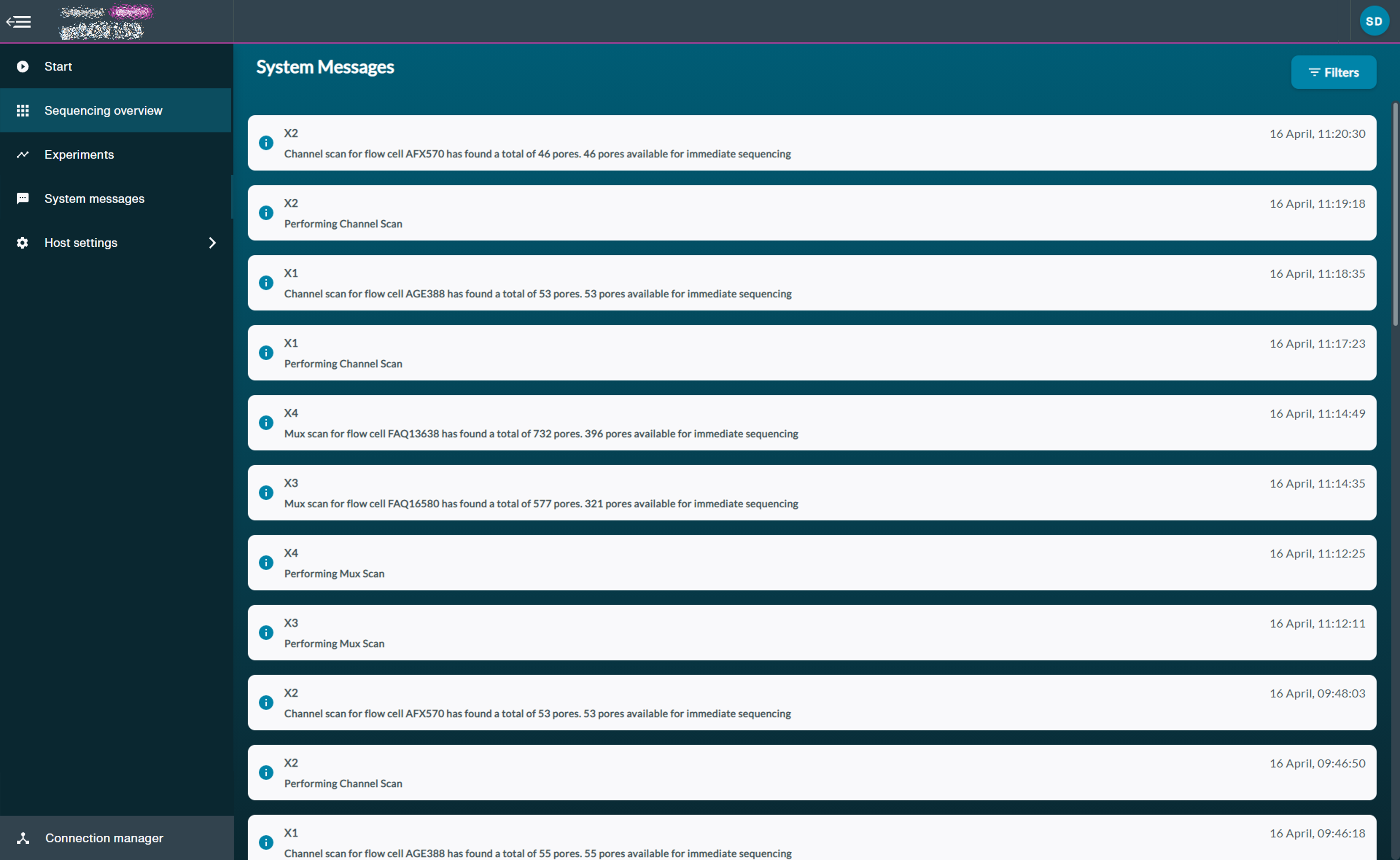
System Messages
All device reports and messages are displayed here.