-
Basecalling overview
A user can basecall and demultiplex their data directly in MinKNOW after a sequencing experiment has finished to avoid having to use command-line tools, or to re-analyse old data using the latest basecalling models. Reads can also be aligned against a reference post-sequencing.
Note: Both barcoding and alignment can be run on FASTQ, POD5 or FAST5 reads when coupled with basecalling.
-
Accessing post-run basecalling
Post-run analysis can be accessed by selecing Analysis on the start page, then selecting Basecalling.
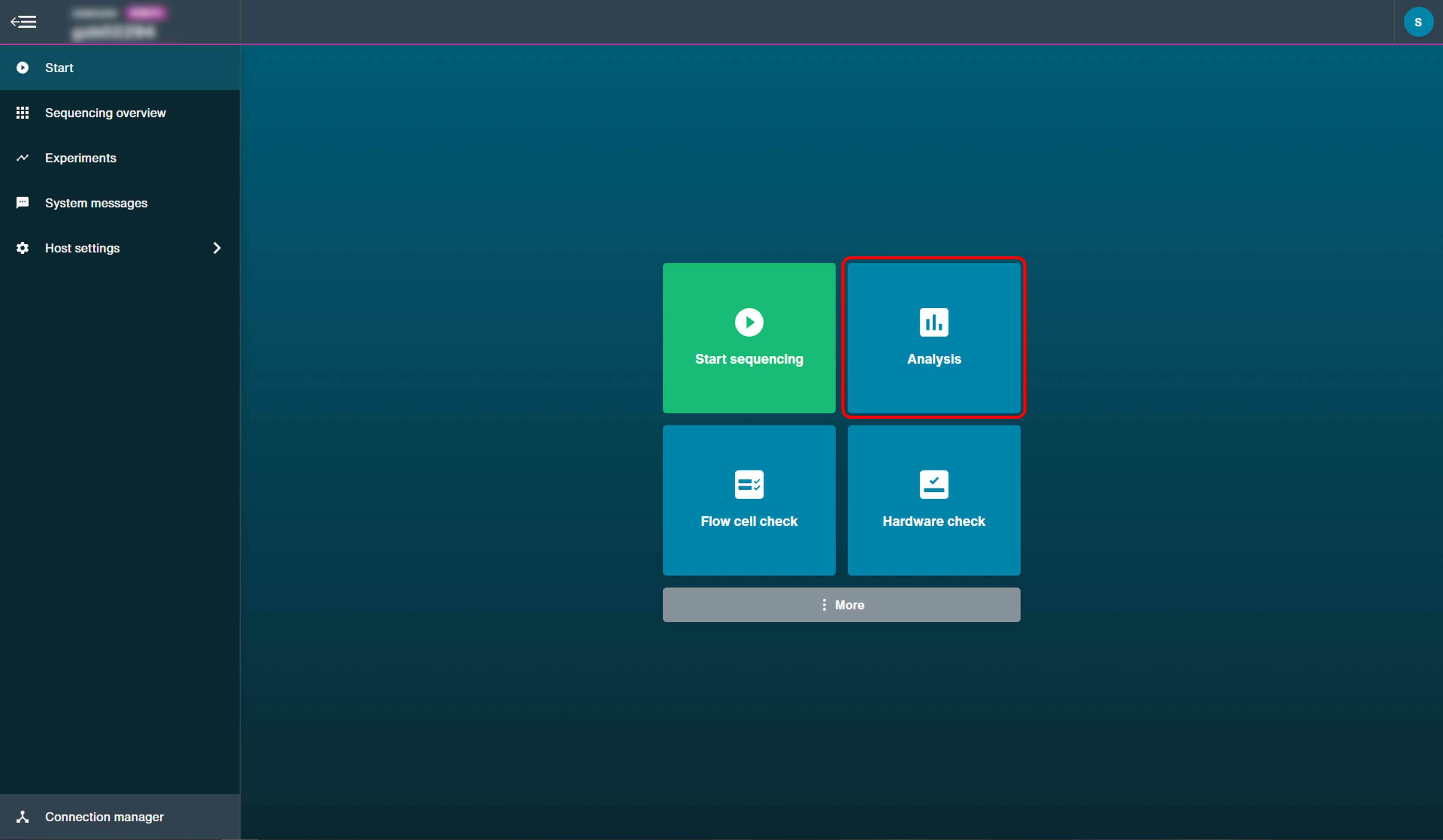
The options provided for basecalling data once a run has finished are described in the MinKNOW protocol under 'Post-run analysis'.
The user also has the option to use barcoding and alignment during basecalling analysis. However, both options can be carried out in independent post-run analyses.
-
Basecalling model selection
Dorado (and previously Guppy), the basecallers used by MinKNOW, provide multiple models for basecalling nanopore data in post-run basecalling. The models that can be selected are described in Local basecalling ealier in this document.
