-
What is ligation-based sequencing?
Ligation-based chemistry is the method that the sequencing adapter is attached (by ligation) to the DNA ends during library preparation using ligation enzymes.
Our ligation sequencing kits are optimised for output and accuracy. These kits are the most popular kits and require double-stranded DNA as input, including gDNA, amplicons, or cDNA.
Our current ligation sequencing kits available are:
- Ligation Sequencing Kit V14 (SQK-LSK114)
- Ligation Sequencing Kit XL V14 (SQK-LSK114-XL)
- Native Barcoding Kit 24 V14 (SQK-NBD114.24)
- Native Barcoding Kit 96 V14 (SQK-NBD114.96)
- PCR Barcoding Expansion 1-12 (EXP-PBC001)
- PCR Barcoding Expansion 1-96 (EXP-PBC096)
- Multiplex Ligation Sequencing Kit V14 XL (SQK-MLK114.96-XL)
Below, we outline the sample input requirements and library preparation workflows for our simplex kits. For information on the barcoding kits, please see the Barcoding kits section.
-
Sample input recommendations
Before starting the Ligation Sequencing Kit library preparation, it is important to ensure that you are using the correct amount of starting material to ensure a successful sequencing experiment. After DNA extraction, we recommend quantifying your DNA samples:
Quantification Method Mass Qubit Fluorometer and Qubit dsDNA BR Assay Kit Size • Agilent 2100 Bioanalyser for samples of <10 kb
• Agilent FemtoPulse for samples of >10 kb
• Oxford Nanopore FlonglePurity Nanodrop 2000 Spectrophotometer For information regarding how to quantify the mass of DNA samples for library preparation input, please refer to the Sample input and recommendations section.
Depending on fragment lengths, we recommend different starting inputs for the Ligation Sequencing Kit V14 (SQK-LSK114) with R10.4.1 flow cells.
Note: These input recommendations are for our standard protocols and may vary depending if a protocol is written for a specific application.Samples Input Short fragment samples (<10 kb) • 100-200 fmol for MinION and PromethION Flow Cells
• 50-100 fmol for Flongle Flow CellsLong fragment samples (>10 kb) • 1 µg for MinION and PromethION Flow Cells
• 500 ng for Flongle Flow CellsMolar quantification is difficult and unreliable for samples consisting of long fragments and we have found these starting inputs are almost always enough for optimal pore occupancy. The graph below illustrates pore occupancy of long fragment libraries using different mass input.
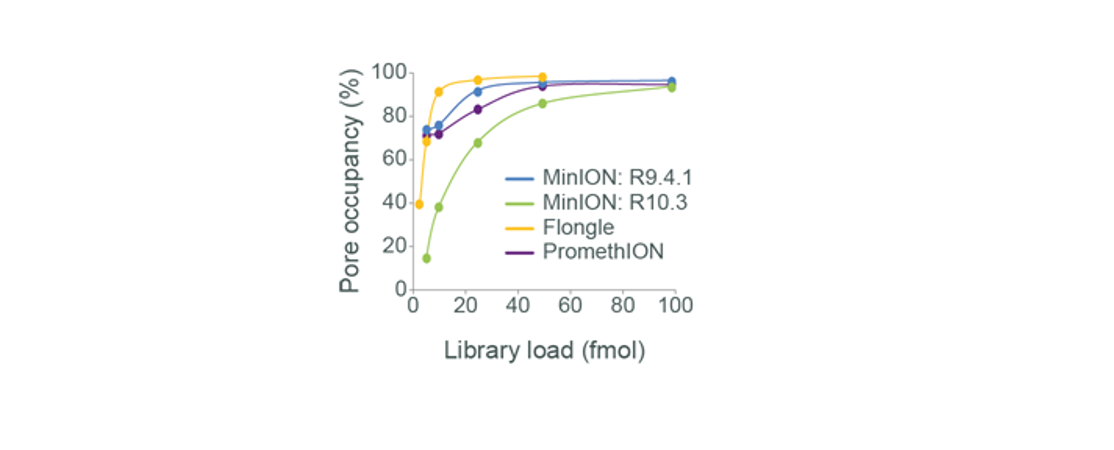
Figure 1. Sequencing libraries were prepared using various starting inputs of a gDNA sample consisting of long molecules, and then run on the various flow cell types. The resulting pore occupancy is shown, and it was found that an input of ~1 µg is sufficient for optimal pore occupancy on R9.4.1 flow cells and an input of ~2 µg was sufficient for optimal pore occupancy on R10.3 flow cells.If you use less than 1 µg in your library preparation, you may have a reduced yield, meaning you will have fewer DNA strands with sequencing adapters (adapted DNA) on each end which may negatively affect sequencing output. This is because optimal pore occupancy may not be reached and it may deteriorate faster as the pores will not always be sequencing, compromising output. For more information on pore occupancy, please see the Sample input and recommendations section.
However, even when starting with as little as 100 ng of high molecular weight (HMW) DNA, we have observed outputs of approximately 10 Gb from R9.4.1 MinION/GridION Flow Cells and approximately 30 Gb from PromethION Flow Cells (Figure 2.). As you decrease input below 100 ng, pore occupancy significantly deteriorates, and we recommend considering amplification (by PCR or multiple displacement amplification) to generate more template. HMW DNA can also be sheared using Covaris g-TUBE or MegaRuptor® to increase strands of DNA for low input samples. For further information, see the optional fragmentation of gDNA in the Sample input and recommendations section.
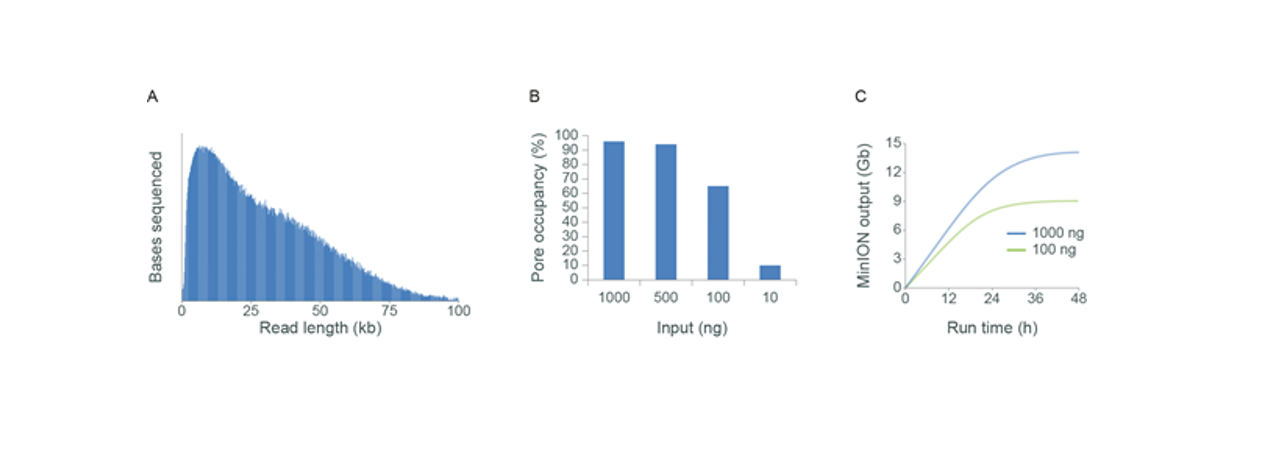
Figure 2. Flow cell output may be decreased when starting with less than 1 µg of HMW gDNA. In the experiment above, gDNA extracted from human cells (GM24385) was titrated as input into the Ligation Sequencing Kit. The resulting libraries were sequenced on MinION (R9.4.1 flow cells). Panel A: The typical read length distribution of a library. Panel B: Starting with 1 µg of HMW DNA results in efficient pore occupancy. As input mass decreases, the number of adapted DNA strands decreases and this leads to reduced pore occupancy. Panel C: The reduction in pore occupancy at a lower input mass means that the pores spend less time sequencing, which results in a reduced sequencing output. -
Ligation Sequencing Kit V14
The Ligation Sequencing Kit V14 (SQK-LSK114) is our newest ligation-based sequencing kit optimised to achieve sequencing accuracies of over 99% (Q20+), with high output on our latest nanopore: R10.4.1. Further improvements include duplex sequencing which allows users to sequence both the template DNA strand and the complement strand. For more information, see the Kit 14 sequencing and duplex basecalling document.
This Kit 14 upgrade includes previous updates such as higher capture rate of DNA to enable lower flow cell loading amounts, and fuel fix technology. Note, due to the higher capture of the adapter, it is important to follow the flow cell loading recommendations in the protocols.
The library preparation is simple and highly versatile, accommodating any double-stranded DNA sample input of 100‒200 fmol of short fragment libraries (<10 kb) or 1 µg of long fragment libraries (>10 kb). Fragment length can be controlled by optional fragmentation or size selection methods which can be found in our DNA/RNA handling page.
This kit is also compatible with upstream processes such as targeted enrichment by sequence capture and whole-genome amplification.
Details of our previous Ligation Sequencing kits are available in the Legacy kit page of this document.
Workflow:
The library preparation involves two enzymatic steps to prepare the DNA ends for sequencing. The first enzymatic step repairs any damage in the DNA molecules, such as nicks and generates uniform ends with 5’ phosphates and 3’ adenine overhangs. The second enzymatic step ligates the sequencing adapters that have complementary thymine tails with the dA-tailed template.
This kit has a standard protocol and multiple associated protocols due to the multiple applications this kit can be used for:
• Ligation Sequencing gDNA V14 (SQK-LSK114)
• Ligation Sequencing Amplicons V14 (SQK-LSK114)
• Ligation Sequencing gDNA - Lambda Control V14 (SQK-LSK114)
• Ligation sequencing gDNA V14 - low input by PCR (SQK-LSK114 with EXP-PCA001)
• Ligation sequencing V14 - PCR Barcoding (SQK-LSK114 with EXP-PBC001 or EXP-PBC096)
• Ligation sequencing V14 - Direct cDNA sequencing (SQK-LSK114)
• Ligation sequencing gDNA V14 - reduced representation methylation sequencing (RRMS) of human samples (SQK-LSK114)
• Ligation Sequencing gDNA V14 - human sample (N50 30 kb) on PromethION (SQK-LSK114)
• Ligation sequencing gDNA V14 - human sample (N50 10 kb) on PromethION (SQK-LSK114)
• Ligation sequencing gDNA V14 - whole genome amplification (SQK-LSK114)
• High duplex ligation sequencing DNA V14 (SQK-LSK114)
• Ligation sequencing V14 - Single-cell transcriptomics with cDNA prepared using 10X Genomics on PromethION (SQK-LSK114) -
Ligation Sequencing Kit XL
The Ligation Sequencing Kit V14 XL (SQK-LSK114-XL) is a scaled-up version of the Ligation Sequencing Kit V14 and contains larger quantities of the same components.
In this kit, there are sufficient reagents to generate 48 sequencing libraries and is recommended for users who would like to process multiple samples simultaneously, either with a multichannel pipette or a liquid handling robot.

