-
Overview of flow cells
The flow cell is the platform an DNA/RNA library is loaded onto and sequenced in a device. Sequencing devices can only use compatible flow cells but they all contain the proprietary sensor array, Application-Specific Integrated Circuit (ASIC) and nanopores for sequencing DNA/RNA libraries. The flow cell contains sufficient buffer to run for up to or more than 72 hours under optimal conditions depending on the flow cell. A user can choose to run continuously or run, stop, wash and load a new library until the buffer and nanopores are exhausted. This enables a single flow cell to be used multiple times before the nanopores are exhausted. An individual flow cell may also be used for different experiments.
Flow cell Flongle MinION/GridION PromethION Device MinION Mk1B, Mk1C and GridION MinION Mk1B, Mk1C and GridION PromethION Storage and stability (unopened) 2-8°C: 4 weeks Room temp: 1 month
2-8°C: 12 weeksRoom temp: 1 month
2-8°C: long termPore count warranty ≥ 50 ≥ 800 ≥ 5000 Nanopore technology allows for real-time assessment of sequencing experiments with the visualisation of run statistics on the MinKNOW UI. This makes it easier for the user to adjust sequencing parameters during run set up and to make the most of a flow cell and experiment.
For libraries of high purity and quality, MinKNOW will be able to generate high data output. We recommend monitoring the pore activity plot to ensure pore occupancy remains high, as well as the pore scan plot to check whether there is a build up of unavailable channels. The translocation speed plot should also be checked to ensure sequencing speed remains between the median target. This is discussed in more depth further in the document.
We recommend live basecalling to allow the user to assess in real-time how the experiment is being conducted on the flow cell. However, fast basecalling also allows real-time assessment on computers that struggle to keep up with basecalling and the raw data can be saved and basecalled at a later date.
For further information on how to monitor your sequencing experiment, we go into further detail in the MinKNOW protocol.
-
Flow cell wash
The Flow Cell Wash Kit (EXP-WSH004) contains a nuclease, DNase I, which is used to digest any remaining library on a flow cell for immediate reuse. The wash step is recommended when there is an accumulation of pores in the ‘recovering’/’unavailable’ state. In the figure below, we have demonstrated pores can be reverted to the 'active'/'single pore' state by pausing sequencing and washing the flow cell with DNase I in the Flow Cell Wash Kit (EXP-WSH004). Below, in figure a), the astrisks indicate where sequencing has been paused and the flow cell washed. Note: If the sequencing run is paused in MinKNOW for the flow cell wash, you will only see the restoration of sequencing pores after a new pore scan has been performed.
The wash step is only recommended where sequencing channels are lost to the 'recovering/unavailable' state. Where pores are in the ‘saturated’ state, the wash step will not be able to revert to the ‘active’ pore state.
![Prom Min flow cell wash image]
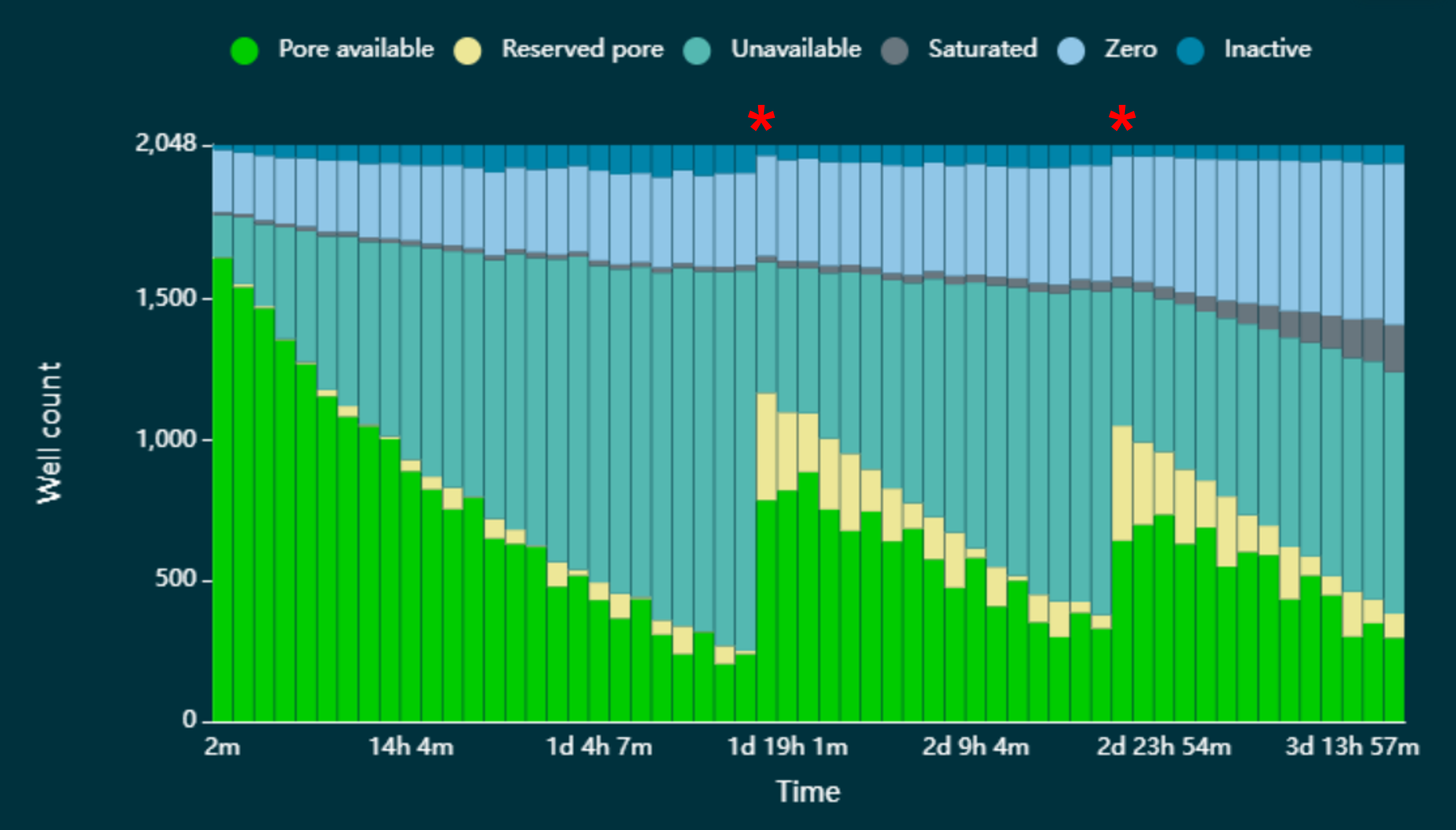
Pore states observed on a flow cell before and after wash steps are performed. A flow cell has been loaded with a sequencing library that has resulted in an accumulation of pores in the “unavailable” state, leading to a decrease in the rate of data acquisition. The red asterisks indicate when a wash step has been performed. A significant number of the pores that had been lost to the “unavailable” state have reverted to the “Pore available” state and are available for sequencing once again.The flow cell wash kit allows multiple uses of a flow cell with different samples and the recovery of unavailable channels during an experiment. Samples can be multiplexed to reduce costs and sequence many samples simultaneously on a single flow cell, for example using one of the Native Barcoding kits or the PCR Barcoding Expansions.
Note: The wash kit should remove 99% of the library. However, some residual DNA may remain on the flow cell. Therefore, we recommend users barcode their libraries when used in conjunction with the Flow Cell Wash Kit, to ensure reads from different libraries can be separated from each other. RNA is also efficiently flushed out of the flow cell but RNA is not digested. Successful deconvolution of DNA reads has been demonstrated in Oxford Nanopore’s internal development.
a) Sequential loading of barcoded libraries without washing. b) Sequential loading of barcoded libraries with washing. Only residual library remained on the flow cell with washing and barcoding samples that are to be sequentially run can help avoid cross-contaimination.The Flow Cell Wash Kit (EXP-WSH004) presents two options:
- Wash and store the flow cell
- Allows for flow cells to be used for multiple experiments but not to be reused immediately after one experiment is complete
- Wash and reuse the flow cell straight away
- We recommend using the pause option in the MinKNOW UI to pause the experiment and to restart as soon as the wash is complete and library loaded.
- This is recommended when there is a build-up of ‘unavailable’ pores.
- Note: When the Pause option is used, all data produced pre- and post-wash is stored in the same data file. Therefore, if changing the sample, be sure to use the Stop function rather than pausing to ensure data is stored in a new file when sequencing is restarted.
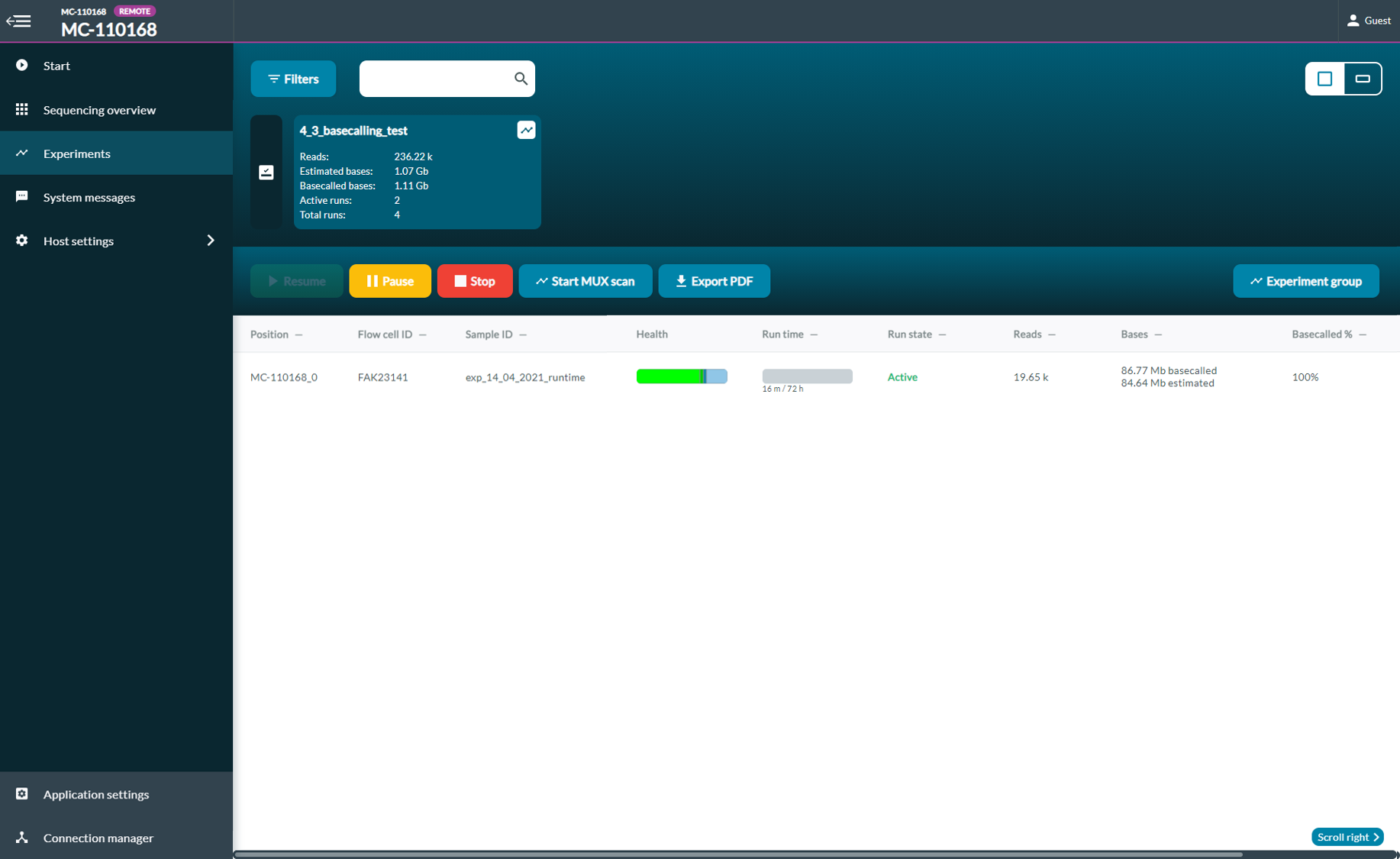
Navigate to the experiments page and select a flow cell to either Pause or Stop a sequencing run to wash a flow cell. - Wash and store the flow cell
-
Pore occupancy
The pore occupancy plot is a summary of time spent in each of the channel states during the run. The channel panel illustrates the real-time representation of each of the channels. Pore occupancy is driven by the number of sequenceable ends (sample molecules with sequencing adapters) loaded onto the flow cell. We recommend monitoring ‘inactive’ and ‘recovering’ channels to keep track of saturated and blocked pores which may reduce output of a run if not dealt with appropriately.
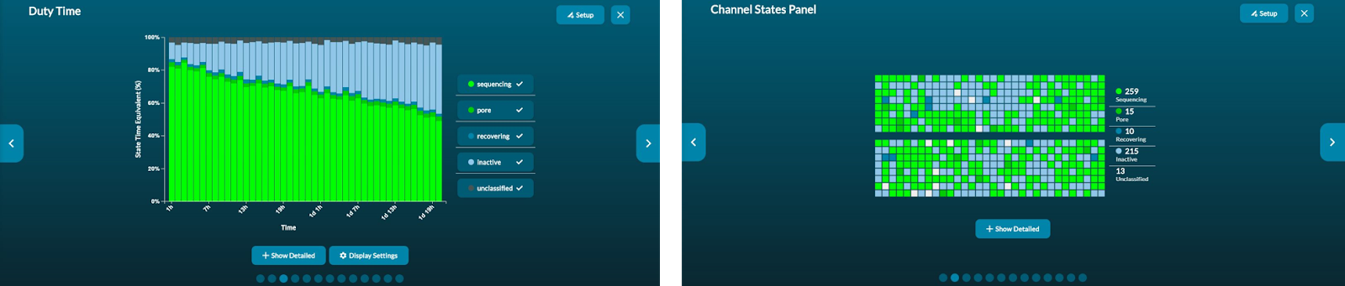
Left: Pore occupancy plot illustrates all the channel states during the entire sequencing run. Right: Channel states panel illustrates the current channel states. -
Pore scan
The pore scan analyses pore status and selects the best performing pores for sequencing. We recommend triggering a pore scan after pausing or stopping an experiment, whether it is for refueling, reloading or washing a flow cell. This is to ensure all the nanopore channels are assessed most recently with the best selection of pores for sequencing.
The pore scan results can be used to assess sequencing efficiency with the duty time plot. If there is a high level of ‘unavailable’ pores, we recommend washing the flow cell as there is blocking occuring which will reduce sequencing and data output. The rate of blocking is mainly due to the purity of the sample; low quality DNA/RNA or contamination from either extraction, sample prep or secondary structure can be some of the causes of an increase of pores in the unavailable state.
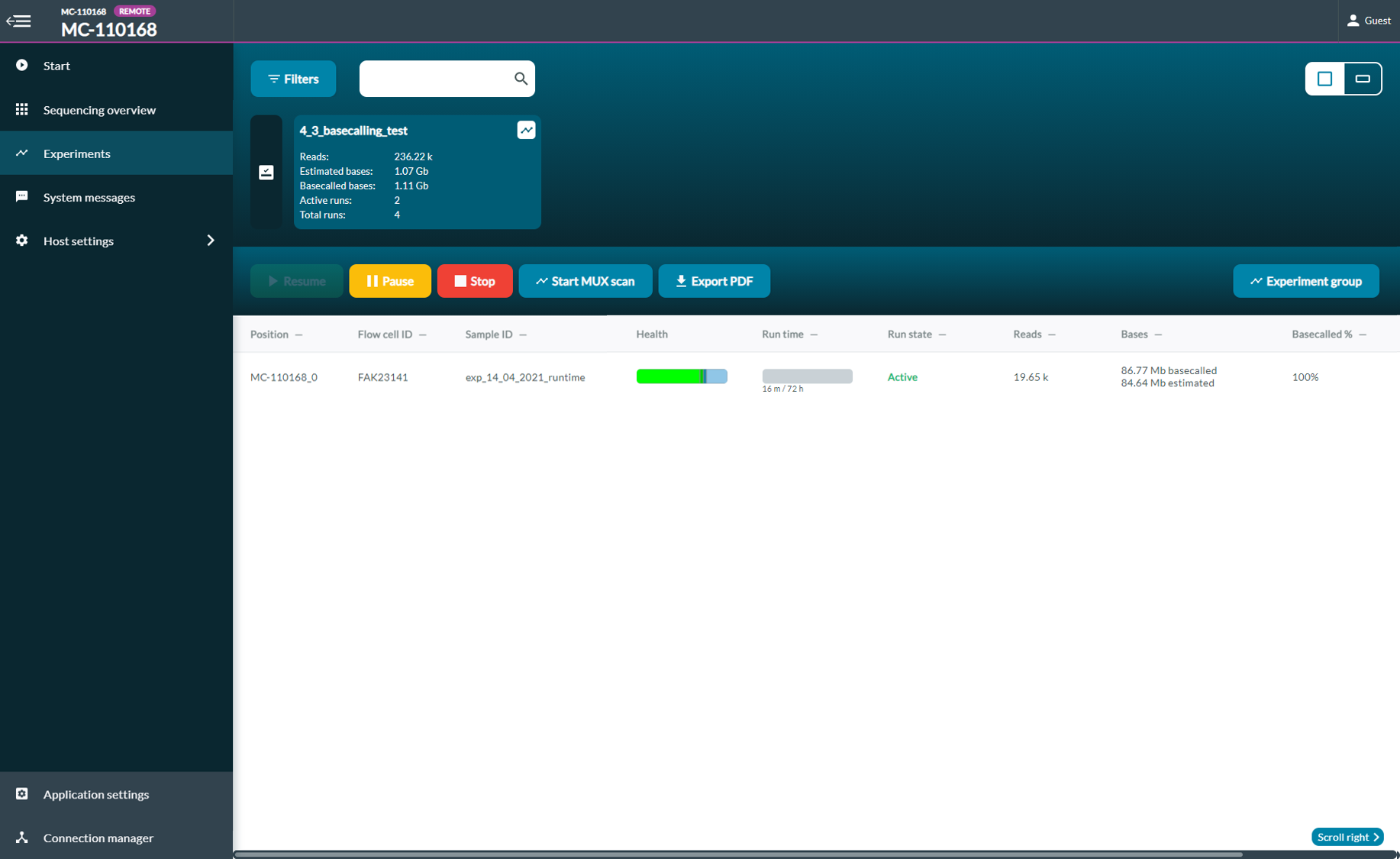
To start a pore scan, navigate to the experiments page and click Start pore scan and select the flow cell to scan in the dialogue box that has opened. This will start a pore scan on the selected flow cell(s) and select a new set of channels for sequencing. -
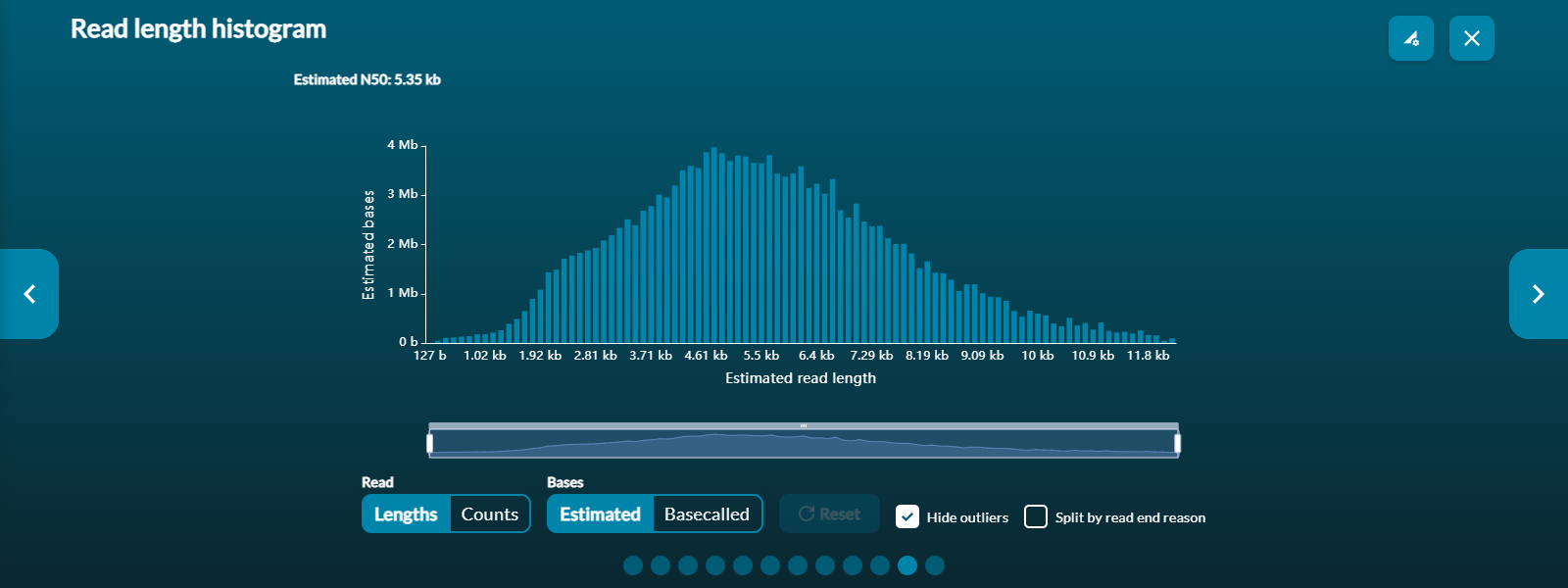
Read length histogram
This visualises the read lengths of the sequencing libraries in real-time and changes throughout the experiment. This can be used to assess if the sequencing library contains an expected sample if the read length distribution roughly matches the input library fragment sizes. If read length does not reflect the input library fragment sizes, the experiment can be stopped and the flow cell washed and stored whilst the library preparation can be optimised, preventing the waste of a flow cell. Note: This graph will take slightly longer to populate for longer reads.
-
Disk space plot
This allows the user to monitor the amount of data to be written on the device memory during sequencing. If all the disk space will be used up, the user can pause the experiment whilst making more disk space and preserve the flow cell.




