-
Introduction to barcoding kits
The barcoding kits are designed to allow the pooling and running of multiple libraries on our flow cell by attaching a known DNA sequence (barcode) to your samples within a pooled library. The pooled library is then sequenced in a single run and MinKNOW uses the barcodes to demultiplex the library during a sequencing run. The barcodes have been carefully designed and extensively purified to minimise cross-talk.
We recommend using our barcoding kits:
- To reduce cost per sample.
- For more efficient use of flow cells when less data per sample is required than the total amount of data that can be generated from a single flow cell.
- For optimal re-use of a flow cell when used with the Flow Cell Wash kit as any barcoded residual reads on a flow cell will be identified between barcoded samples.
Note: As more samples are multiplexed, less data per sample is generated.
Multiplexing samples onto one flow cell can reduce the cost per sample for a user. In the table below, we illustrate how barcoding 1 to 96 samples onto a single flow cell can alter the cost per sample.
Note: These costs are an example and do not reflect our current store prices.Barcodes 0 6 12 24 48 96 Flow cell price $500 $500 $500 $500 $500 $500 Library price $99 $99 $99 $99 $99 $99 Barcode price - $18.75 $37.50 $75 $150 $300 Price per sample $599 $102.95 $53.04 $28.08 $15.60 $9.36 There are four types of barcoding kits:
- Ligation-based barcoding kits
- Ligation-based PCR barcoding kits
- Rapid chemistry-based barcoding kits
- Rapid chemistry-based PCR barcoding kits
The recently released barcoding kits have been updated to combine all the reagents required for library preparation, barcoding and flow cell priming into a kit.
All our kits with 96 barcodes have been developed to allow flexibility in the number of samples that can be multiplexed. Users can prepare multiplexed libraries using various options without extra expansion packs:
- 12 libraries containing 24 barcodes
- 6 libraries containing 48 barcodes
- 3 libraries containing 96 barcodes
Expansion packs for library preparation and flow cell priming reagents are available to make use of any remaining barcodes in a kit.
The barcode sequences are available at the back of the document in the appendix.
-
Ligation-based barcoding kits
Native Barcoding Kit 24 and 96 V14
There are two standalone native barcoding kits and one expansion:
- Native Barcoding Kit 24 V14 (SQK-NBD114.24)
- Native Barcoding Kit 96 V14 (SQK-NBD114.96)
- Native Barcoding Auxiliary Kit V14 (EXP-NBA114)
The Native Barcoding Kits 24 and 96 V14 are the newest native barcoding sequencing kit and contain all required reagents for barcoding 24 or 96 samples, preparing libraries for sequencing and flow cell priming. The kits are optimised to achieve sequencing accuracies of over 99% (Q20+), with high output. The native barcodes in these kits are shipped at 640 nM.
The Native Barcoding Kits are recommended for users who want to multiplex their samples with a PCR-free method to preserve base modifications. There are up to 24 or 96 unique barcodes which can be used for gDNA or amplicons. The barcoding kits are optimised to generate maximum output without the need for PCR.
These kits require double-stranded DNA as input with the following recommendations below. Due to the higher capture of the adapter, it is important to follow the flow cell loading recommendations in the protocols.
Kit Input requirements Native Barcoding Kit 24 V14 (SQK-NBD114.24)
for barcoding up to 24 samples• <4 barcodes, 1000 ng of gDNA per sample
• >4 barcodes, 400 ng of gDNA per sample
• 200 fmol (130 ng for 1 kb amplicons) DNA per sampleNative Barcoding Kit 96 V14 (SQK-NBD114.96)
for barcoding 96 samples• 400 ng of gDNA per sample
• 200 fmol (130 ng for 1 kb amplicons) DNA per sampleThe Native Barcoding Auxiliary Kit V14 (EXP-NBA114) contains additional library preparation reagents for use with the Native Barcoding Kits for another 12 reactions. For further information on our available expansions, please see the Expansion packs section.
Workflow
The DNA is repaired and the ends dA-tailed in preparation for the dT-tailed native barcode ligation. After barcoding, the samples are pooled together and sequencing adapters are ligated to the barcode ends for sequencing.
Multiplex Ligation Barcoding Kit V14 96 XL
This kit is our recently updated kit for the high output and PCR-free, low-plex sequencing of up to three samples of double-stranded DNA. This kit is recommended to perform low-plex whole genome sequencing. All required reagents for barcoding up to 96 samples, preparing libraries for sequencing and flow cell loading are included, using our Kit 14 chemistry.
We recommend using 1 µg gDNA per sample to sequence two samples per flow cell. This results in the sequencing of up to 96 samples across 48 flow cells. Three samples across two flow cells can also be used to sequence up to 96 samples across 64 flow cells to maximise sequencing output.
This kit uses our native barcodes and is compatible with the Native Barcoding Auxiliary Kit V14 (EXP-NBA114) for additional library preparation reagents.
Workflow
The library preparation method is similar to the Ligation Sequencing Kit protocol; DNA ends are repaired and dA-tailed in preparation for the dT-tailed native barcode ligation. After barcoding, the samples are pooled together and the sequencing adapters are ligated to the barcode ends for sequencing.
-
Ligation-based PCR barcoding kits
PCR Barcoding Expansion 1-12 and 1-96
For low starting inputs, there are two PCR barcoding expansion packs with 12 or 96 barcodes; PCR Barcoding Expansion 1-12 (EXP-PBC001) and PCR Barcoding Expansion 1-96 (EXP-PBC096). These barcoding expansions are available to use in combination with our Ligation Sequencing Kit, to enable the pooling and running of multiple PCR amplified sequencing libraries for low inputs of gDNA, amplicon, and cDNA. The kits include adapters and up to 96 forward and reverse PCR primers with the same barcode in both forward and reverse primers.
An input of <100-200 fmol per DNA sample is required or <100-200 fmol of first-round PCR product with tailed primers per sample.
Note: cDNA and amplicons have to be prepared in advance for barcoding, as reverse transcription or the first round of amplification are not included in the library preparation protocol. Details of recommendations are outlined in the protocol.
Workflow
Depending on your input, the sample preparation step will very before the library preparation step.
For low gDNA inputs, you will need to prepare your DNA ends for PCR adapter ligation and perform PCR amplification with barcoded primers before pooling your samples prior to library preparation.
For low cDNA and amplicon inputs, you will need to perform a round of PCR to incorporate tailed primers. A second round of PCR amplification step with barcoded primers will be performed and the samples are pooled prior to library preparation.
The library preparation step is the same across all sample inputs and the pooled barcoded samples are end-prepped for the sequencing adapters to ligate to the barcodes before sequencing.
Dual barcoding protocol
To perform massively parallel sequencing of up to 2,304 samples of gDNA or amplicons on a single flow cell, the PCR Barcoding Expansion 1-96 (EXP-PBC096) can be combined with the Native Barcoding Kit V14 24 (SQK-NBD114.24). The dual barcoding protocol combines both kits to dual barcode the libraries for pooling and sequencing on a single flow cell.
We recommend dual barcoding for users wishing to sequence more than 96 samples or want to sort libraries into multiple categories as each barcode can be used to denote a different sample feature. The inner barcode is a PCR barcode and the outer barcode is a native barcode.
An input of <100-200 fmol per DNA sample is required or <100-200 fmol of first-round PCR product with tailed primers per sample.
Note: cDNA and amplicons have to be prepared in advance for barcoding, as reverse transcription or the first round of amplification are not included in the library preparation protocol. Details of recommendations are outlined in the protocol.
Workflow
The ends of the DNA input are first prepared for PCR adapter ligation and PCR is performed with barcoded primers to barcode up to 96 samples. The 96 samples are pooled into a single library and this is repeated to create 24 pools of 96 barcoded samples. Next, prepare the DNA ends for native barcode ligation for each pool. After barcoding, pool the dual barcoded libraries into a single pool and ligate the sequencing adapters to the native barcodes before sequencing on a single flow cell.
-
Rapid chemistry-based barcoding kits
Rapid Barcoding Kit 24 and 96
There are two standalone rapid barcoding kits and one expansion using Kit 14 chemistry:
- Rapid Barcoding Kit 24 V14 (SQK-RBK114.24)
- Rapid Barcoding Kit 96 V14 (SQK-RBK114.96)
- Rapid Adapter Auxiliary V14 (EXP-RAA114)
The Rapid Barcoding Kit 24 V14 (SQK-RBK114.24) and Rapid Barcoding Kit 96 V14 (SQK-RBK114.96) are both standalone kits for barcoding and sequencing up to 24 or 96 samples, respectively, using a library preparation optimised for speed and simplicity, requiring minimal laboratory equipment. For users requiring high accuracy and output, we recommend the Native Barcoding Kit 24 or 96 V14 kits (SQK-NBD114.24 and SQK-NBD114.96).
This kit has been upgraded to use Kit 14 chemistry with our updated sequencing adapter to improve sequencing accuracies and higher outputs. The barcodes in our rapid-based kits are shipped at 10 µM.
- A sample input of 200 ng of gDNA is required for each sample to be barcoded and sequenced.
Note: We also have our Rapid Barcoding Kit 96 (SQK-RBK110.96) that uses our previous Kit 10 chemistry. The library preparation workflow and input are the same but uses our previous sequencing adapter and flow cell type.
Workflow
The DNA input is undergoes a transposase step to fragment the samples whilst attaching rapid barcodes to the ends simultaneously. The samples can be pooled and a bead clean-up performed to remove any excess transposases in the reaction. The sequencing adapters are then attached to the barcodes and the library can be sequenced.
Midnight RT PCR Expansion
The Midnight RT PCR Expansion (EXP-MRT001) has been developed to work in conjunction with the Rapid Barcoding Kit (SQK-RBK110.96) or the Rapid Barcoding Kit 96 V14 (SQK-RBK114.96) to enable PCR tiling of overlapping 1.2 kb amplicons across the SARS-CoV-2 genome.
The primers provided in this kits were initially developed by Freed et al., 2020, using the Primal Scheme. These primers are updated as new mutations are identified in the SARS-CoV-2 lineages and we will work closely with the community (including the entire ARTIC Nextwork, Nikki Freed, Olin Silander, Josh Quick, John Tyson, Nick Loman and many more) to rapidly respond and ensure performance of Oxford Nanopore Technologies' protocols are always kept up-to-date, providing the best performance for robust SARS-CoV-2 whole genome sequencing.
For this kit, an input of extracted RNA in 10 mM of Tris-HCl, pH 8.0 is required.
Workflow
The extracted RNA is reverse transcribed with random hexamers and the samples are split into pools A and B for tiled PCR. The primer pools are combined with their associated sample, making sure to not combine different samples together. The DNA samples are then fragmented using a transposase and the rapid barcodes are attached to the fragment ends simultaneously. After barcoding, the samples can be combined into a single pool and a bead clean-up performed to remove any excess barcodes and transposases before sequencing on a flow cell.
16S Barcoding Kit 24 V14
This is our updated 16S kit using our latest Kit 14 chemistry to amplify the entire ~1,500 bp 16S rRNA gene from extracted gDNA. With our chemistry upgrade, sequencing accuracies and output are improved and a lower flow cell loading amount is required. The number of barcodes have also been increased to enable barcoding of up to 24 samples into a single library. The barcodes in our 16S rapid-based kit are shipped at 1 µM.
We recommend this kit for rapid 16S sequencing for bacterial identification as the highly conserved and variant regions across different bacterial species are sequenced. A specific region of interest can also be selected for before preparing the library with this kit to enable the sequencing of the organisms in a sample without unnecessary regions of the genome to improve the efficiency of identification by reducing speed and costs.
An input of 10 ng of gDNA per sample is required.
Workflow
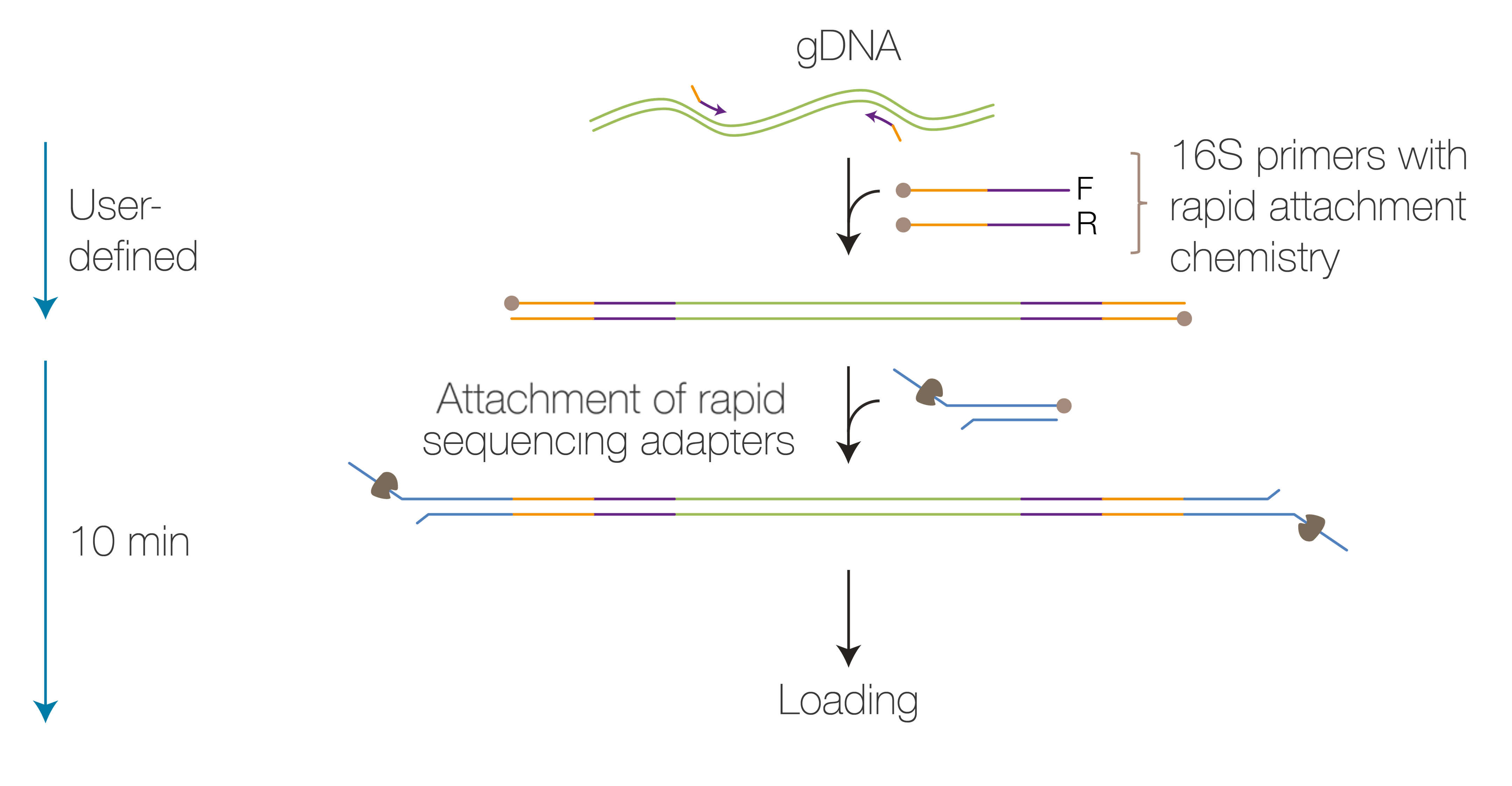
The gDNA undergoes PCR amplification with 16S primers (27F and 1492R) to amplify the 16S gene present in the sample whilst simultaneously attaching rapid adapters to the DNA ends. The rapid barcodes are then attached to the rapid adapters and the samples pooled before a bead clean-up. The rapid sequencing adapters are then attached to the rapid barcodes before sequencing.
-
Rapid chemistry-based PCR barcoding kits
Rapid PCR Barcoding Kit 24 V14
The Rapid PCR Barcoding Kit 24 V14 (SQK-RPB114.24) is the fastest and simplest method of preparing barcoded libraries starting with low inputs of gDNA (1‒5 ng). PCR amplification is performed to increase the number of templates for sequencing. This kit has been upgraded to use Kit 14 chemistry and the number of barcodes available in the kit have been increased to 24. The Kit 14 upgrade includes improved raw read sequencing accuracies and higher output. The barcodes in our rapid-based kits are shipped at 10 µM.
A low starting input of 1-5 ng of gDNA per sample is required.
Note: The DNA must be at least 4 kb in length to ensure correct tagmentation and PCR amplification.
Workflow
The gDNA is prepared for sequencing by undergoing tagmentation to fragment the DNA and to simultaneously attach PCR primers to the fragment ends. PCR is performed and rapid barcode primers are attached to the DNA ends in the same step. After barcoding, the samples are pooled and the rapid sequencing adapter is attached to the ends of the DNA samples for sequencing.
-
cDNA barcoding kit
cDNA-PCR Barcoding Kit V14
This our recently updated cDNA barcoding kit to barcode up to 24 RNA samples for sequencing of the cDNA. Identification and qualtification of full-length transcripts can be performed with this kit and isoforms, splice variants, and fusion transcripts can be sequenced. The barcodes in our rapid-based kits are shipped at 10 µM.
The Kit 14 chemistry upgrade in this kit reduces the level of free sequencing adapter and improves sequencing accuracies and output. Other updates include the addition of a cDNA RT adapter and RT primer to reduce transcript overlaps during the reverse transcription step to enable the measurement of the poly(A) tail length and a unique molecular identifier (UMI) for the identification of splice variants.
A low input of 10 ng of enriched RNA per sample is required which can be poly(A)-tailed or ribodepleted RNA. Total RNA can also be used but requires 500 ng per sample.
Workflow
The RNA undergoes a reverse transcription step to prepare full-length cDNA from the input RNA and incorporates the UMI. During reverse transcription, the poly(dT) reverse transcription adapter is ligated to the 3’ terminal poly(A) tail of the template molecule. The bottom strand of the adapter is removed and a reverse transcription primer is annealed, anchoring the start of transcription to include the entire 3’ terminal poly(A) tail. Then a strand-switching primer, containing a UMI, is added during reverse transcription, allowing strand switching to occur and generate a full-length cDNA strand. This is tagged with universal sequences on both ends.
Note: The reverse transcriptase inhibits downstream PCR and the enzymes must be heat-inactivated and the reverse transcribed sample to be split across four PCR reactions to dilute the inhibitors. This is to allow the amplification of cDNA with maximum efficiency, without losing sensitivity in the next step.
Once the full-length cDNA is prepared, PCR amplification is performed and rapid barcode primers are attached to the cDNA samples simultaneously. After barcoding, the samples are pooled into a single library and the rapid sequencing adapters attached are to the rapid barcode primers for sequencing.
